2N3F
 
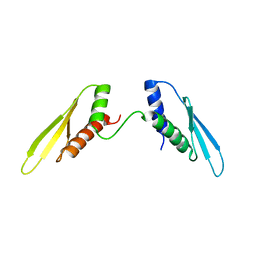 | |
1YRM
 
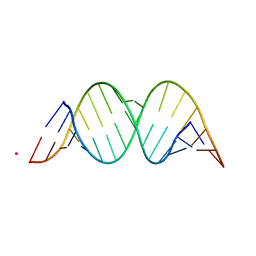 | |
1Z79
 
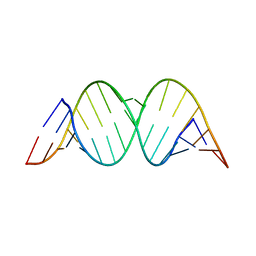 | |
2A64
 
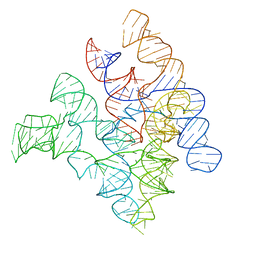 | | Crystal Structure of Bacterial Ribonuclease P RNA | | Descriptor: | ribonuclease P RNA | | Authors: | Kazantsev, A.V, Krivenko, A.A, Harrington, D.J, Holbrook, S.R, Adams, P.D, Pace, N.R. | | Deposit date: | 2005-07-01 | | Release date: | 2005-09-20 | | Last modified: | 2024-02-14 | | Method: | X-RAY DIFFRACTION (3.3 Å) | | Cite: | Crystal structure of a bacterial ribonuclease P RNA.
Proc.Natl.Acad.Sci.Usa, 102, 2005
|
|
1YZD
 
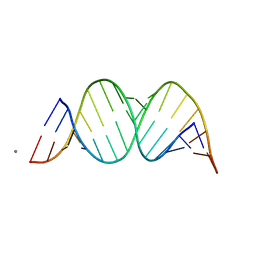 | |
1VFG
 
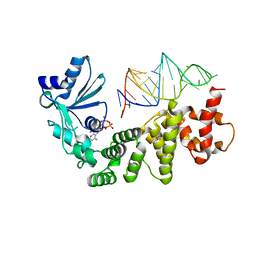 | | Crystal structure of tRNA nucleotidyltransferase complexed with a primer tRNA and an incoming ATP analog | | Descriptor: | DIPHOSPHOMETHYLPHOSPHONIC ACID ADENOSYL ESTER, RNA (75-MER), poly A polymerase | | Authors: | Tomita, K, Fukai, S, Ishitani, R, Ueda, T, Takeuchi, N, Vassylyev, D.G, Nureki, O, RIKEN Structural Genomics/Proteomics Initiative (RSGI) | | Deposit date: | 2004-04-13 | | Release date: | 2004-08-10 | | Last modified: | 2023-12-27 | | Method: | X-RAY DIFFRACTION (2.8 Å) | | Cite: | Structural basis for template-independent RNA polymerization.
Nature, 430, 2004
|
|
7EI5
 
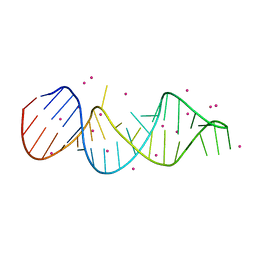 | |
7EI6
 
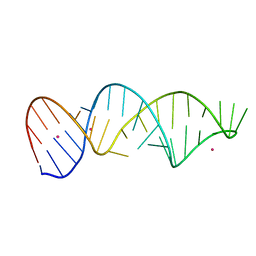 | |
1L1W
 
 | | NMR structure of a SRP19 binding domain in human SRP RNA | | Descriptor: | SRP19 binding domain of SRP RNA | | Authors: | Sakamoto, T, Morita, S, Tabata, K, Nakamura, K, Kawai, G. | | Deposit date: | 2002-02-20 | | Release date: | 2002-05-22 | | Last modified: | 2024-05-22 | | Method: | SOLUTION NMR | | Cite: | Solution structure of a SRP19 binding domain in human SRP RNA.
J.Biochem.(Tokyo), 132, 2002
|
|
2VXG
 
 | | Crystal structure of the conserved C-terminal region of Ge-1 | | Descriptor: | CG6181-PA, ISOFORM A | | Authors: | Jinek, M, Eulalio, A, Lingel, A, Helms, S, Conti, E, Izaurralde, E. | | Deposit date: | 2008-07-04 | | Release date: | 2008-09-09 | | Last modified: | 2024-05-08 | | Method: | X-RAY DIFFRACTION (1.9 Å) | | Cite: | The C-Terminal Region of Ge-1 Presents Conserved Structural Features Required for P-Body Localization.
RNA, 14, 2008
|
|
2QH4
 
 | |
7KGA
 
 | | Crystal structure of a mosquito-borne flavivirus dumbbell RNA | | Descriptor: | IRIDIUM HEXAMMINE ION, RNA (90-MER) | | Authors: | Akiyama, B.M, Graham, M.E, O'Donoghue, Z, Beckham, J.D, Kieft, J.S. | | Deposit date: | 2020-10-16 | | Release date: | 2021-06-23 | | Last modified: | 2024-03-06 | | Method: | X-RAY DIFFRACTION (2.1 Å) | | Cite: | Three-dimensional structure of a flavivirus dumbbell RNA reveals molecular details of an RNA regulator of replication.
Nucleic Acids Res., 49, 2021
|
|
3ZC4
 
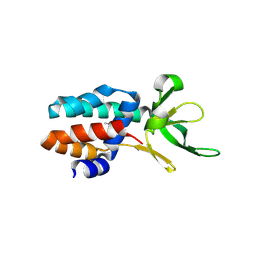 | | The structure of Csa5 from Sulfolobus solfataricus. | | Descriptor: | DI(HYDROXYETHYL)ETHER, SSO1398 | | Authors: | Reeks, J, Anderson, L, White, M.F, Naismith, J.H. | | Deposit date: | 2012-11-15 | | Release date: | 2013-02-20 | | Last modified: | 2013-07-31 | | Method: | X-RAY DIFFRACTION (2.72 Å) | | Cite: | Structure of the Archaeal Cascade Subunit Csa5: Relating the Small Subunits of Crispr Effector Complexes.
RNA Biol., 10, 2013
|
|
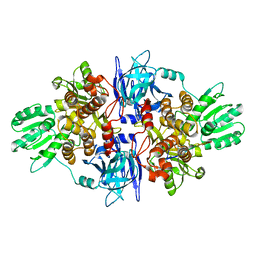
3O8C
 
 | |
1SI2
 
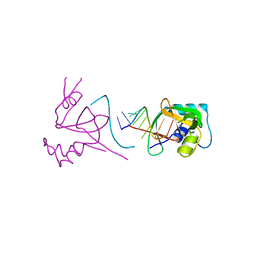 | |
8A0L
 
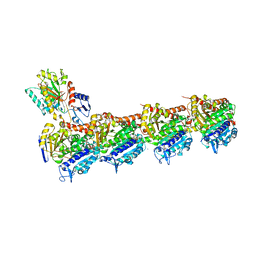 | | Tubulin-CW1-complex | | Descriptor: | (3~{S},4~{R},8~{S},10~{S},12~{S},14~{S})-14-[(~{Z},4~{R})-4-(hydroxymethyl)hex-2-en-2-yl]-4,12-dimethoxy-9,9-dimethyl-3,8,10-tris(oxidanyl)-1-oxacyclotetradecan-2-one, 2-(N-MORPHOLINO)-ETHANESULFONIC ACID, CALCIUM ION, ... | | Authors: | Prota, A.E, Diaz, J.F, Steinmetz, M.O, Oliva, M.A. | | Deposit date: | 2022-05-28 | | Release date: | 2022-12-14 | | Last modified: | 2024-01-31 | | Method: | X-RAY DIFFRACTION (1.9981 Å) | | Cite: | Chemical modulation of microtubule structure through the laulimalide/peloruside site.
Structure, 31, 2023
|
|
2B7G
 
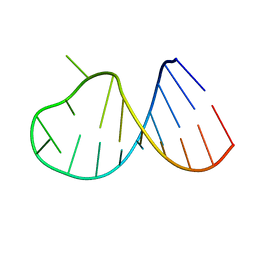 | |
3O8R
 
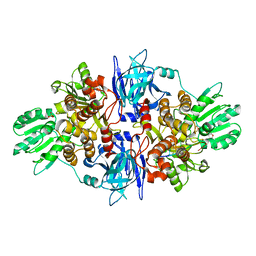 | |
1XSU
 
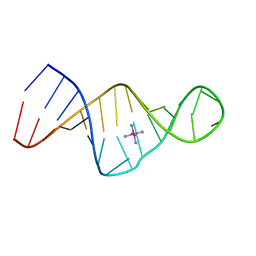 | |
3P59
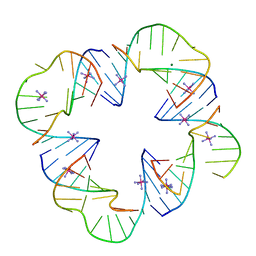 
 | | First Crystal Structure of a RNA Nanosquare | | Descriptor: | COBALT HEXAMMINE(III), MAGNESIUM ION, RNA (5'-R(*CP*CP*GP*GP*AP*GP*GP*AP*AP*CP*UP*AP*CP*(5BU)P*G)-3'), ... | | Authors: | Dibrov, S, Hermann, T, McLean, J, Parsons, J. | | Deposit date: | 2010-10-08 | | Release date: | 2011-04-06 | | Last modified: | 2024-02-21 | | Method: | X-RAY DIFFRACTION (2.1793 Å) | | Cite: | Self-assembling RNA square.
Proc.Natl.Acad.Sci.USA, 108, 2011
|
|
1SI3
 
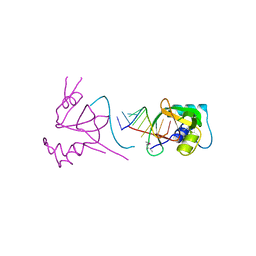 | |
2GRW
 
 | | Solution structure of the poliovirus 3'-UTR Y-stem | | Descriptor: | 5'-GGACCUCUCGAAAGAGAUGUCC-3' | | Authors: | Heus, H.A, Zoll, J, Tessari, M, van Kuppeveld, F.J.M, Melchers, W.J.G. | | Deposit date: | 2006-04-25 | | Release date: | 2007-03-06 | | Last modified: | 2024-05-29 | | Method: | SOLUTION NMR | | Cite: | Breaking pseudo-twofold symmetry in the poliovirus 3'-UTR Y-stem by restoring Watson-Crick base pairs.
Rna, 13, 2007
|
|
1XST
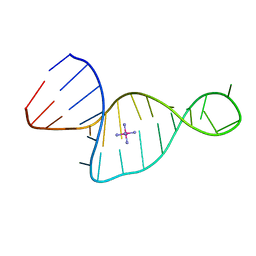 
 | |
1M5V
 
 | | Transition State Stabilization by a Catalytic RNA | | Descriptor: | (4S)-2-METHYL-2,4-PENTANEDIOL, CALCIUM ION, RNA HAIRPIN RIBOZYME, ... | | Authors: | Rupert, P.B, Massey, A.P, Sigurdsson, S.T, Ferre-D'Amare, A.R. | | Deposit date: | 2002-07-09 | | Release date: | 2002-10-12 | | Last modified: | 2024-02-14 | | Method: | X-RAY DIFFRACTION (2.4 Å) | | Cite: | Transition state stabilization by a catalytic RNA
Science, 298, 2002
|
|
1M5O
 
 | | Transition State Stabilization by a Catalytic RNA | | Descriptor: | CALCIUM ION, RNA SUBSTRATE, RNA HAIRPIN RIBOZYME, ... | | Authors: | Rupert, P.B, Massey, A.P, Sigurdsson, S.T, Ferre-D'Amare, A.R. | | Deposit date: | 2002-07-09 | | Release date: | 2002-10-12 | | Last modified: | 2024-02-14 | | Method: | X-RAY DIFFRACTION (2.2 Å) | | Cite: | Transition state stabilization by a catalytic RNA
Science, 298, 2002
|
|
