7P4Y
 
 | |
7P4V
 
 | |
7P50
 | |
7P52
 
 | |
7PJX
 
 | | Structure of the 70S-EF-G-GDP ribosome complex with tRNAs in hybrid state 1 (H1-EF-G-GDP) | | Descriptor: | 16S ribosomal RNA, 23S ribosomal RNA, 30S ribosomal protein S10, ... | | Authors: | Petrychenko, V, Peng, B.Z, Schwarzer, A.C, Peske, F, Rodnina, M.V, Fischer, N. | | Deposit date: | 2021-08-24 | | Release date: | 2021-10-20 | | Last modified: | 2025-03-19 | | Method: | ELECTRON MICROSCOPY (6.5 Å) | | Cite: | Structural mechanism of GTPase-powered ribosome-tRNA movement.
Nat Commun, 12, 2021
|
|
7PJW
 | | Structure of the 70S-EF-G-GDP-Pi ribosome complex with tRNAs in hybrid state 2 (H2-EF-G-GDP-Pi) | | Descriptor: | 16S ribosomal RNA, 23S ribosomal RNA, 30S ribosomal protein S10, ... | | Authors: | Petrychenko, V, Peng, B.Z, Schwarzer, A.C, Peske, F, Rodnina, M.V, Fischer, N. | | Deposit date: | 2021-08-24 | | Release date: | 2021-10-20 | | Last modified: | 2025-03-12 | | Method: | ELECTRON MICROSCOPY (4 Å) | | Cite: | Structural mechanism of GTPase-powered ribosome-tRNA movement.
Nat Commun, 12, 2021
|
|
7PJY
 
 | | Structure of the 70S-EF-G-GDP ribosome complex with tRNAs in chimeric state 1 (CHI1-EF-G-GDP) | | Descriptor: | 16S ribosomal RNA, 23S ribosomal RNA, 30S ribosomal protein S10, ... | | Authors: | Petrychenko, V, Peng, B.Z, Schwarzer, A.C, Peske, F, Rodnina, M.V, Fischer, N. | | Deposit date: | 2021-08-24 | | Release date: | 2021-10-20 | | Last modified: | 2025-03-12 | | Method: | ELECTRON MICROSCOPY (3.1 Å) | | Cite: | Structural mechanism of GTPase-powered ribosome-tRNA movement.
Nat Commun, 12, 2021
|
|
7PJZ
 
 | | Structure of the 70S-EF-G-GDP ribosome complex with tRNAs in chimeric state 2 (CHI2-EF-G-GDP) | | Descriptor: | 16S ribosomal RNA, 23S ribosomal RNA, 30S ribosomal protein S10, ... | | Authors: | Petrychenko, V, Peng, B.Z, Schwarzer, A.C, Peske, F, Rodnina, M.V, Fischer, N. | | Deposit date: | 2021-08-24 | | Release date: | 2021-10-20 | | Last modified: | 2025-03-12 | | Method: | ELECTRON MICROSCOPY (6 Å) | | Cite: | Structural mechanism of GTPase-powered ribosome-tRNA movement.
Nat Commun, 12, 2021
|
|
7PJS
 
 | | Structure of the 70S ribosome with tRNAs in the classical pre-translocation state and apramycin (C) | | Descriptor: | 16S ribosomal RNA, 23S ribosomal RNA, 30S ribosomal protein S10, ... | | Authors: | Petrychenko, V, Peng, B.Z, Schwarzer, A.C, Peske, F, Rodnina, M.V, Fischer, N. | | Deposit date: | 2021-08-24 | | Release date: | 2021-10-20 | | Last modified: | 2025-03-12 | | Method: | ELECTRON MICROSCOPY (2.35 Å) | | Cite: | Structural mechanism of GTPase-powered ribosome-tRNA movement
Nat Commun, 12, 2021
|
|
7PJU
 
 | | Structure of the 70S ribosome with tRNAs in hybrid state 2 (H2) | | Descriptor: | 16S ribosomal RNA, 23S ribosomal RNA, 30S ribosomal protein S10, ... | | Authors: | Petrychenko, V, Peng, B.Z, Schwarzer, A.C, Peske, F, Rodnina, M.V, Fischer, N. | | Deposit date: | 2021-08-24 | | Release date: | 2021-11-17 | | Last modified: | 2025-03-19 | | Method: | ELECTRON MICROSCOPY (9.5 Å) | | Cite: | Structural mechanism of GTPase-powered ribosome-tRNA movement.
Nat Commun, 12, 2021
|
|
7P1G
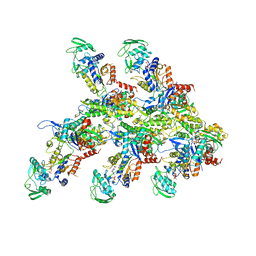 
 | | Structure of the P. aeruginosa ExoY-F-actin complex | | Descriptor: | 3'-DEOXY-GUANOSINE-5'-TRIPHOSPHATE, ADENOSINE-5'-DIPHOSPHATE, Actin, ... | | Authors: | Belyy, A, Merino, F, Raunser, S. | | Deposit date: | 2021-07-01 | | Release date: | 2021-11-17 | | Last modified: | 2021-12-01 | | Method: | ELECTRON MICROSCOPY (3.2 Å) | | Cite: | Mechanism of actin-dependent activation of nucleotidyl cyclase toxins from bacterial human pathogens.
Nat Commun, 12, 2021
|
|
7PJV
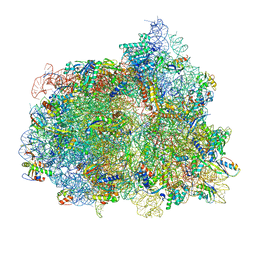 
 | | Structure of the 70S-EF-G-GDP-Pi ribosome complex with tRNAs in hybrid state 1 (H1-EF-G-GDP-Pi) | | Descriptor: | 16S ribosomal RNA, 23S ribosomal RNA, 30S ribosomal protein S10, ... | | Authors: | Petrychenko, V, Peng, B.Z, Schwarzer, A.C, Peske, F, Rodnina, M.V, Fischer, N. | | Deposit date: | 2021-08-24 | | Release date: | 2021-10-20 | | Last modified: | 2025-03-12 | | Method: | ELECTRON MICROSCOPY (3.1 Å) | | Cite: | Structural mechanism of GTPase-powered ribosome-tRNA movement.
Nat Commun, 12, 2021
|
|
7P1H
 
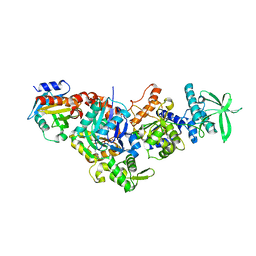 | | Structure of the V. vulnificus ExoY-G-actin-profilin complex | | Descriptor: | ADENOSINE-5'-TRIPHOSPHATE, Actin, cytoplasmic 1, ... | | Authors: | Belyy, A, Merino, F, Raunser, S. | | Deposit date: | 2021-07-01 | | Release date: | 2021-11-17 | | Last modified: | 2021-12-01 | | Method: | ELECTRON MICROSCOPY (3.9 Å) | | Cite: | Mechanism of actin-dependent activation of nucleotidyl cyclase toxins from bacterial human pathogens.
Nat Commun, 12, 2021
|
|
3NZ8
 
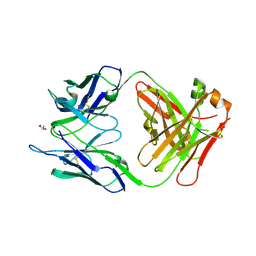 | | Crystal structure of the HIV-2 neutralizing Fab fragment 7C8 | | Descriptor: | GLYCEROL, Mouse anti V3 antibody 7C8 Fab, heavy chain, ... | | Authors: | Uchtenhagen, H, Friemann, R, Raszewski, G, Spetz, A.-L, Nilsson, L, Achour, A. | | Deposit date: | 2010-07-16 | | Release date: | 2011-05-18 | | Last modified: | 2024-11-06 | | Method: | X-RAY DIFFRACTION (2.7 Å) | | Cite: | Crystal Structure of the HIV-2 Neutralizing Fab Fragment 7C8 with High Specificity to the V3 Region of gp125.
Plos One, 6, 2011
|
|
3QUK
 
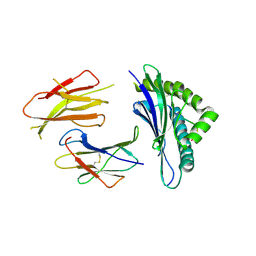 | | Crystal structures of the murine class I major histocompatibility complex H-2Db in complex with LCMV-derived gp33 altered peptide ligand (Y4A) | | Descriptor: | Beta-2-microglobulin, H-2 class I histocompatibility antigen, D-B alpha chain, ... | | Authors: | Allerbring, E, Duru, A.D, Uchtenhagen, H, Madhurantakam, C, Grimm, S, Tomek, M.B, Mazumdar, P.A, Spetz, A, Friemann, R, Sandalova, T, Uhlin, M, Nygren, P, Achour, A. | | Deposit date: | 2011-02-24 | | Release date: | 2012-03-21 | | Last modified: | 2024-10-16 | | Method: | X-RAY DIFFRACTION (2.41 Å) | | Cite: | Unexpected T-cell recognition of an altered peptide ligand is driven by reversed thermodynamics.
Eur.J.Immunol., 42, 2012
|
|
3RRW
 
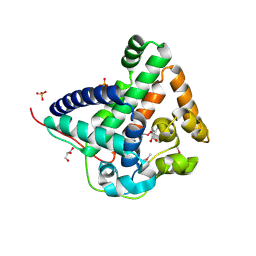 | | Crystal structure of the TL29 protein from Arabidopsis thaliana | | Descriptor: | CALCIUM ION, GLYCEROL, PHOSPHATE ION, ... | | Authors: | Lundberg, E, Storm, P, Schroder, W.P, Funk, C. | | Deposit date: | 2011-05-01 | | Release date: | 2011-08-03 | | Last modified: | 2024-10-16 | | Method: | X-RAY DIFFRACTION (2.5 Å) | | Cite: | Crystal structure of the TL29 protein from Arabidopsis thaliana: An APX homolog without peroxidase activity.
J.Struct.Biol., 176, 2011
|
|
3R8R
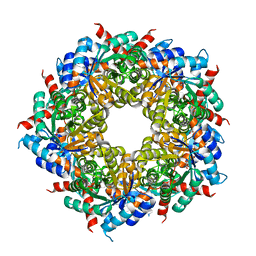 
 | |
4YTE
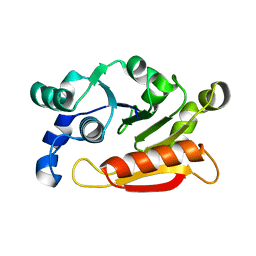 
 | |
3QGZ
 
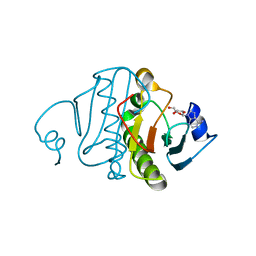 | | Re-investigated high resolution crystal structure of histidine triad nucleotide-binding protein 1 (HINT1) from rabbit complexed with adenosine | | Descriptor: | ADENOSINE, Histidine triad nucleotide-binding protein 1 | | Authors: | Dolot, R.M, Ozga, M, Krakowiak, A, Nawrot, B, Stec, W.J. | | Deposit date: | 2011-01-25 | | Release date: | 2011-02-16 | | Last modified: | 2023-09-13 | | Method: | X-RAY DIFFRACTION (1.1 Å) | | Cite: | High-resolution X-ray crystal structure of rabbit histidine triad nucleotide-binding protein 1 (rHINT1) - adenosine complex at 1.10A resolution
Acta Crystallogr.,Sect.D, 67, 2011
|
|
4YT2
 
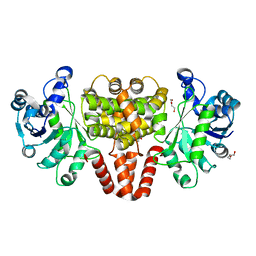 | | Hmd II from Methanocaldococcus jannaschii | | Descriptor: | DI(HYDROXYETHYL)ETHER, H(2)-forming methylenetetrahydromethanopterin dehydrogenase-related protein MJ1338, TRIETHYLENE GLYCOL | | Authors: | Fujishiro, T, Ermler, U, Shima, S. | | Deposit date: | 2015-03-17 | | Release date: | 2015-07-01 | | Last modified: | 2024-05-01 | | Method: | X-RAY DIFFRACTION (1.65 Å) | | Cite: | Towards a functional identification of catalytically inactive [Fe]-hydrogenase paralogs.
Febs J., 282, 2015
|
|
3QUL
 
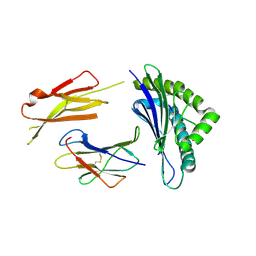 | | Crystal structures of the murine class I major histocompatibility complex H-2Db in complex with LCMV-derived gp33 altered peptide ligand (Y4S) | | Descriptor: | Beta-2-microglobulin, H-2 class I histocompatibility antigen, D-B alpha chain, ... | | Authors: | Allerbring, E, Duru, A.D, Uchtenhagen, H, Madhurantakam, C, Grimm, S, Tomek, M.B, Mazumdar, P.A, Spetz, A, Friemann, R, Sandalova, T, Uhlin, M, Nygren, P, Achour, A. | | Deposit date: | 2011-02-24 | | Release date: | 2012-03-21 | | Last modified: | 2024-11-06 | | Method: | X-RAY DIFFRACTION (2 Å) | | Cite: | Unexpected T-cell recognition of an altered peptide ligand is driven by reversed thermodynamics.
Eur.J.Immunol., 42, 2012
|
|
3RIE
 
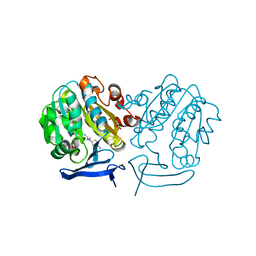 | | The structure of Plasmodium falciparum spermidine synthase in complex with 5'-methylthioadenosine and N-(3-aminopropyl)-trans-cyclohexane-1,4-diamine | | Descriptor: | 2-(2-{2-[2-(2-METHOXY-ETHOXY)-ETHOXY]-ETHOXY}-ETHOXY)-ETHANOL, 5'-DEOXY-5'-METHYLTHIOADENOSINE, GLYCEROL, ... | | Authors: | Williams, M, Sprenger, J, Persson, L, Louw, A.I, Birkholtz, L, Al-Karadaghi, S. | | Deposit date: | 2011-04-13 | | Release date: | 2012-04-25 | | Last modified: | 2023-09-13 | | Method: | X-RAY DIFFRACTION (1.9 Å) | | Cite: | A novel inhibitor of Plasmodium falciparum spermidine synthase: a twist in the tail.
Malar J, 14, 2015
|
|
3OER
 
 | | Crystal structure of trimeric frataxin from the yeast saccharomyces cerevisiae, complexed with cobalt | | Descriptor: | COBALT (II) ION, Frataxin homolog, mitochondrial | | Authors: | Soderberg, C.A.G, Rajan, S, Gakh, O, Ta, C, Isaya, G, Al-Karadaghi, S. | | Deposit date: | 2010-08-13 | | Release date: | 2011-08-24 | | Last modified: | 2023-09-06 | | Method: | X-RAY DIFFRACTION (3.2 Å) | | Cite: | Oligomerization Propensity and Flexibility of Yeast Frataxin Studied by X-ray Crystallography and Small-Angle X-ray Scattering.
J.Mol.Biol., 414, 2011
|
|
3OEQ
 
 | | Crystal structure of trimeric frataxin from the yeast Saccharomyces cerevisiae, with full length n-terminus | | Descriptor: | Frataxin homolog, mitochondrial | | Authors: | Soderberg, C.A.G, Rajan, S, Gakh, O, Ta, C, Isaya, G, Al-Karadaghi, S. | | Deposit date: | 2010-08-13 | | Release date: | 2011-08-24 | | Last modified: | 2023-09-06 | | Method: | X-RAY DIFFRACTION (2.96 Å) | | Cite: | Oligomerization Propensity and Flexibility of Yeast Frataxin Studied by X-ray Crystallography and Small-Angle X-ray Scattering.
J.Mol.Biol., 414, 2011
|
|
3R5A
 
 | |
