5FEP
 
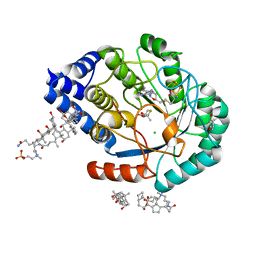 | | HydE from T. maritima in complex with (2R,4R)-MeTDA | | Descriptor: | (2R,4R)-2-methyl-1,3-thiazolidine-2,4-dicarboxylic acid, 3-[(3-CHOLAMIDOPROPYL)DIMETHYLAMMONIO]-1-PROPANESULFONATE, CHLORIDE ION, ... | | Authors: | Rohac, R, Amara, P, Benjdia, A, Martin, L, Ruffie, P, Favier, A, Berteau, O, Mouesca, J.M, Fontecilla-Camps, J.C, Nicolet, Y. | | Deposit date: | 2015-12-17 | | Release date: | 2016-04-06 | | Last modified: | 2024-01-10 | | Method: | X-RAY DIFFRACTION (1.45 Å) | | Cite: | Carbon-sulfur bond-forming reaction catalysed by the radical SAM enzyme HydE.
Nat.Chem., 8, 2016
|
|
5FF0
 
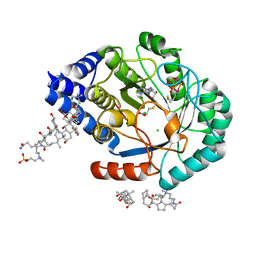 | | HydE from T. maritima in complex with S-adenosyl-L-cysteine and methionine | | Descriptor: | 3-[(3-CHOLAMIDOPROPYL)DIMETHYLAMMONIO]-1-PROPANESULFONATE, CHLORIDE ION, Fe4-Se4 cluster, ... | | Authors: | Rohac, R, Amara, P, Benjdia, A, Martin, L, Ruffie, P, Favier, A, Berteau, O, Mouesca, J.M, Fontecilla-Camps, J.C, Nicolet, Y. | | Deposit date: | 2015-12-17 | | Release date: | 2016-04-06 | | Last modified: | 2024-01-10 | | Method: | X-RAY DIFFRACTION (1.49 Å) | | Cite: | Carbon-sulfur bond-forming reaction catalysed by the radical SAM enzyme HydE.
Nat.Chem., 8, 2016
|
|
4J9J
 
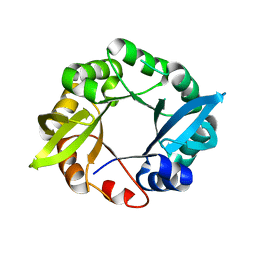 | | Structure of designed HisF | | Descriptor: | Imidazole glycerol phosphate synthase subunit HisF | | Authors: | Sterner, R, Rajendran, C, Sperl, J. | | Deposit date: | 2013-02-16 | | Release date: | 2013-07-17 | | Last modified: | 2023-09-20 | | Method: | X-RAY DIFFRACTION (2.3 Å) | | Cite: | Establishing catalytic activity on an artificial ( beta alpha )8-barrel protein designed from identical half-barrels.
Febs Lett., 587, 2013
|
|
5FEW
 
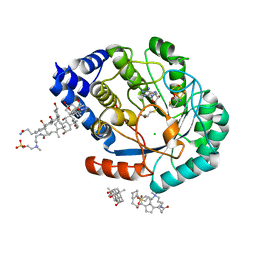 | | HydE from T. maritima in complex with S-adenosyl-L-cysteine (final product) | | Descriptor: | (2R,4R)-2-methyl-1,3-thiazolidine-2,4-dicarboxylic acid, 3-[(3-CHOLAMIDOPROPYL)DIMETHYLAMMONIO]-1-PROPANESULFONATE, CHLORIDE ION, ... | | Authors: | Rohac, R, Amara, P, Benjdia, A, Martin, L, Ruffie, P, Favier, A, Berteau, O, Mouesca, J.M, Fontecilla-Camps, J.C, Nicolet, Y. | | Deposit date: | 2015-12-17 | | Release date: | 2016-04-06 | | Last modified: | 2024-01-10 | | Method: | X-RAY DIFFRACTION (1.17 Å) | | Cite: | Carbon-sulfur bond-forming reaction catalysed by the radical SAM enzyme HydE.
Nat.Chem., 8, 2016
|
|
3PIH
 
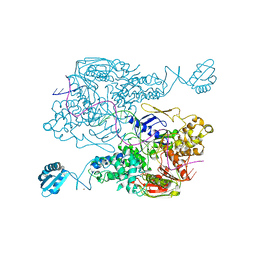 | | T. maritima UvrA in complex with fluorescein-modified DNA | | Descriptor: | DNA (32-MER), PYROPHOSPHATE, UvrABC system protein A, ... | | Authors: | Jaciuk, M, Nowak, E, Nowotny, M. | | Deposit date: | 2010-11-06 | | Release date: | 2011-01-19 | | Last modified: | 2023-09-06 | | Method: | X-RAY DIFFRACTION (2.9 Å) | | Cite: | Structure of UvrA nucleotide excision repair protein in complex with modified DNA.
Nat.Struct.Mol.Biol., 18, 2011
|
|
4I6Z
 
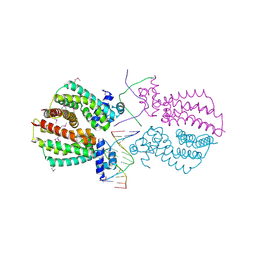 | | Crystal structure of the transcriptional regulator TM1030 with 24bp DNA oligonucleotide | | Descriptor: | DNA OLIGONUCLEOTIDE, Transcriptional regulator, TetR family | | Authors: | Koclega, K.D, Chruszcz, M, Cooper, D.R, Petkowski, J.J, Tkaczuk, K.L, Joachimiak, A, Minor, W, Midwest Center for Structural Genomics (MCSG) | | Deposit date: | 2012-11-30 | | Release date: | 2013-01-02 | | Last modified: | 2023-12-06 | | Method: | X-RAY DIFFRACTION (3.2 Å) | | Cite: | Crystal structure of the transcriptional regulator TM1030 with 24bp DNA oligonucleotide
To be Published
|
|
4KAR
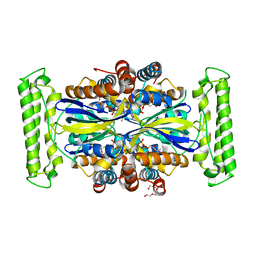 
 | | Crystal structure of FDTS (TM0449) mutant (H53D) with FAD | | Descriptor: | FLAVIN-ADENINE DINUCLEOTIDE, NONAETHYLENE GLYCOL, Thymidylate synthase | | Authors: | Mathews, I.I. | | Deposit date: | 2013-04-22 | | Release date: | 2014-03-26 | | Last modified: | 2023-09-20 | | Method: | X-RAY DIFFRACTION (2.03 Å) | | Cite: | Flavin-Dependent Thymidylate Synthase as a Drug Target for Deadly Microbes: Mutational Study and a Strategy for Inhibitor Design.
J Bioterror Biodef, Suppl 12, 2013
|
|
7OG9
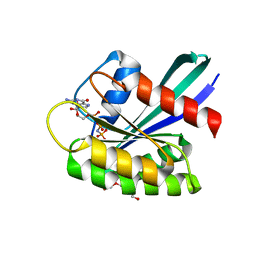 
 | | GTPase HRAS under ambient pressure | | Descriptor: | GTPase HRas, MAGNESIUM ION, PHOSPHOAMINOPHOSPHONIC ACID-GUANYLATE ESTER, ... | | Authors: | Colloc'h, N, Girard, E, Prange, T, Kalbitzer, H.R. | | Deposit date: | 2021-05-06 | | Release date: | 2022-02-09 | | Last modified: | 2024-01-31 | | Method: | X-RAY DIFFRACTION (1.75 Å) | | Cite: | Equilibria between conformational states of the Ras oncogene protein revealed by high pressure crystallography.
Chem Sci, 13, 2022
|
|
7P4M
 
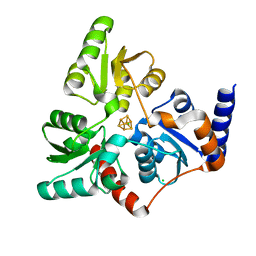 | |
7P4Q
 
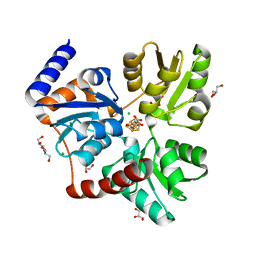 | |
7P4P
 
 | |
2NB6
 | |
4EV2
 
 | |
4EV5
 | |
4EWN
 
 | | Structure of HisF-D130V+D176V with bound rCdRP | | Descriptor: | 1-(O-carboxy-phenylamino)-1-deoxy-D-ribulose-5-phosphate, Imidazole glycerol phosphate synthase subunit HisF | | Authors: | Reisinger, B, Bocola, M, Rajendran, C, List, F, Sterner, R. | | Deposit date: | 2012-04-27 | | Release date: | 2012-12-05 | | Last modified: | 2024-02-28 | | Method: | X-RAY DIFFRACTION (1.903 Å) | | Cite: | A sugar isomerization reaction established on various (beta-alpha)8-barrel scaffolds is based on substrate-assisted catalysis
Protein Eng.Des.Sel., 25, 2012
|
|
5TQL
 | | Crystal structure of TIM-Barrel protein HisF-C9S | | Descriptor: | Imidazole glycerol phosphate synthase subunit HisF, PHOSPHATE ION | | Authors: | Bender, B.J, Meiler, J. | | Deposit date: | 2016-10-24 | | Release date: | 2016-11-16 | | Last modified: | 2023-10-04 | | Method: | X-RAY DIFFRACTION (1.9 Å) | | Cite: | Experimental and Computational Identification of Naive Binders to a TIM-Barrel Protein Scaffold
To Be Published
|
|
5TWI
 
 | |
4FHR
 
 | | Crystal structure of the complex between the flagellar motor proteins FliG and FliM. | | Descriptor: | Flagellar motor switch protein FliG, Flagellar motor switch protein FliM | | Authors: | Paz, A, Vartanian, A.S, Fortgang, E.A, Abramson, J, Dahlquist, F.W. | | Deposit date: | 2012-06-06 | | Release date: | 2012-08-22 | | Last modified: | 2023-09-13 | | Method: | X-RAY DIFFRACTION (1.931 Å) | | Cite: | Structure of flagellar motor proteins in complex allows for insights into motor structure and switching.
J.Biol.Chem., 287, 2012
|
|
8OYQ
 
 | |
8OYZ
 | |
8OZ9
 
 | |
8P59
 | |
8P3R
 
 | |
8PF2
 
 | |
8QMN
 
 | | [FeFe]-hydrogenase maturase HydE from T. maritima - dialysis experiment - empty structure | | Descriptor: | 3-[(3-CHOLAMIDOPROPYL)DIMETHYLAMMONIO]-1-PROPANESULFONATE, CHLORIDE ION, IRON/SULFUR CLUSTER, ... | | Authors: | Omeiri, J, Martin, L, Usclat, A, Cherrier, M.V, Nicolet, Y. | | Deposit date: | 2023-09-23 | | Release date: | 2023-11-22 | | Last modified: | 2023-12-27 | | Method: | X-RAY DIFFRACTION (1.48 Å) | | Cite: | Maturation of the [FeFe]-Hydrogenase: Direct Transfer of the ( kappa 3 -cysteinate)Fe II (CN)(CO) 2 Complex B from HydG to HydE.
Angew.Chem.Int.Ed.Engl., 62, 2023
|
|
