3OEI
 
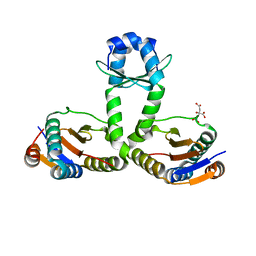 | | Crystal structure of Mycobacterium tuberculosis RelJK (Rv3357-Rv3358-RelBE3) | | Descriptor: | CITRATE ANION, RelJ (Antitoxin Rv3357), RelK (Toxin Rv3358) | | Authors: | Miallau, L, Cascio, D, Eisenberg, D, TB Structural Genomics Consortium (TBSGC) | | Deposit date: | 2010-08-12 | | Release date: | 2011-03-16 | | Last modified: | 2023-09-20 | | Method: | X-RAY DIFFRACTION (2.145 Å) | | Cite: | Comparative proteomics identifies the cell-associated lethality of M. tuberculosis RelBE-like toxin-antitoxin complexes.
Structure, 21, 2013
|
|
4MAX
 
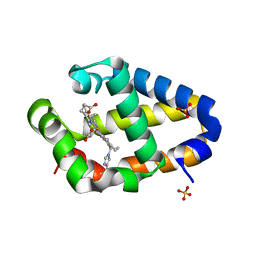 | | Crystal structure of Synechococcus sp. PCC 7002 globin at cryogenic temperature with heme modification | | Descriptor: | HEME B/C, SULFATE ION, cyanoglobin | | Authors: | Wenke, B.B, Schlessman, J.L, Heroux, A, Lecomte, J.T.J. | | Deposit date: | 2013-08-18 | | Release date: | 2013-08-28 | | Last modified: | 2024-10-30 | | Method: | X-RAY DIFFRACTION (1.444 Å) | | Cite: | The 2/2 hemoglobin from the cyanobacterium Synechococcus sp. PCC 7002 with covalently attached heme: Comparison of X-ray and NMR structures.
Proteins, 82, 2014
|
|
3MAX
 
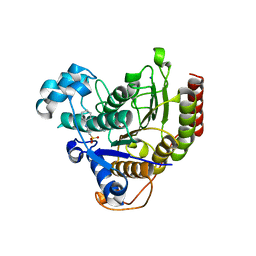 | |
6MAX
 
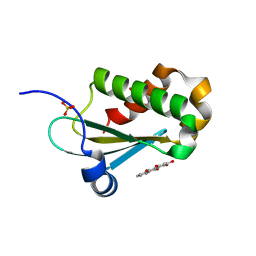 | |
1MAX
 
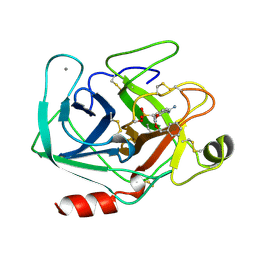 | | BETA-TRYPSIN PHOSPHONATE INHIBITED | | Descriptor: | BETA-TRYPSIN, CALCIUM ION, [N-(BENZYLOXYCARBONYL)AMINO](4-AMIDINOPHENYL)METHANE-PHOSPHONATE | | Authors: | Bertrand, J, Oleksyszyn, J, Kam, C, Boduszek, B, Presnell, S, Plaskon, R, Suddath, F, Powers, J, Williams, L. | | Deposit date: | 1996-02-06 | | Release date: | 1996-10-14 | | Last modified: | 2024-11-13 | | Method: | X-RAY DIFFRACTION (1.8 Å) | | Cite: | Inhibition of trypsin and thrombin by amino(4-amidinophenyl)methanephosphonate diphenyl ester derivatives: X-ray structures and molecular models.
Biochemistry, 35, 1996
|
|
2MAX
 
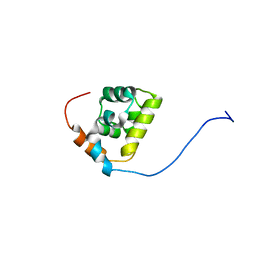 | |
7MAX
 
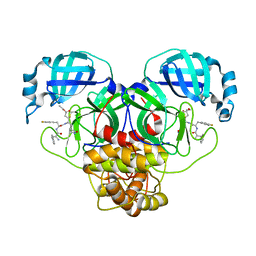 | |
4Y08
 
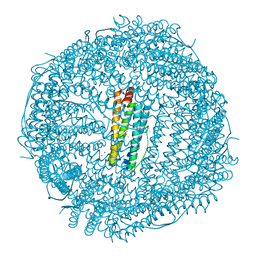 | | ONE MINUTE IRON LOADED HUMAN H FERRITIN | | Descriptor: | CHLORIDE ION, FE (II) ION, Ferritin heavy chain, ... | | Authors: | Pozzi, C, Di Pisa, F, Mangani, S. | | Deposit date: | 2015-02-05 | | Release date: | 2015-09-23 | | Last modified: | 2024-01-10 | | Method: | X-RAY DIFFRACTION (1.34 Å) | | Cite: | Iron binding to human heavy-chain ferritin.
Acta Crystallogr.,Sect.D, 71, 2015
|
|
4YKH
 
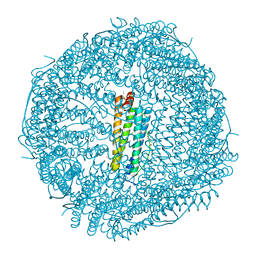 | | Thirty minutes iron loaded human H ferritin | | Descriptor: | BICINE, CHLORIDE ION, FE (II) ION, ... | | Authors: | Pozzi, C, Di Pisa, F, Mangani, S. | | Deposit date: | 2015-03-04 | | Release date: | 2015-09-09 | | Last modified: | 2024-01-10 | | Method: | X-RAY DIFFRACTION (1.52 Å) | | Cite: | Iron binding to human heavy-chain ferritin.
Acta Crystallogr.,Sect.D, 71, 2015
|
|
4ZJK
 
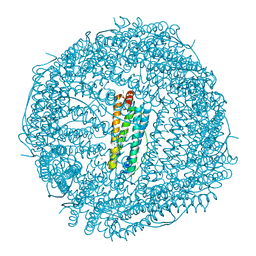 | | FIVE MINUTES IRON LOADED HUMAN H FERRITIN | | Descriptor: | CHLORIDE ION, FE (II) ION, Ferritin heavy chain, ... | | Authors: | Pozzi, C, Di Pisa, F, Mangani, S. | | Deposit date: | 2015-04-29 | | Release date: | 2015-09-09 | | Last modified: | 2024-01-10 | | Method: | X-RAY DIFFRACTION (1.56 Å) | | Cite: | Iron binding to human heavy-chain ferritin.
Acta Crystallogr.,Sect.D, 71, 2015
|
|
4OYN
 
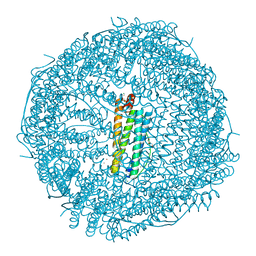 | | Fifteen minutes iron loaded human H ferritin | | Descriptor: | 2-AMINO-2-HYDROXYMETHYL-PROPANE-1,3-DIOL, BICINE, CHLORIDE ION, ... | | Authors: | Pozzi, C, Di Pisa, F, Mangani, S. | | Deposit date: | 2014-02-12 | | Release date: | 2015-09-09 | | Last modified: | 2023-09-27 | | Method: | X-RAY DIFFRACTION (1.43 Å) | | Cite: | Iron binding to human heavy-chain ferritin.
Acta Crystallogr.,Sect.D, 71, 2015
|
|
1OEY
 
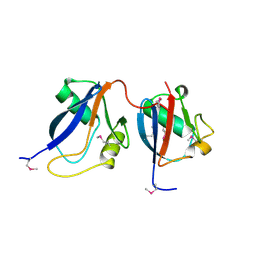 | | Heterodimer of p40phox and p67phox PB1 domains from human NADPH oxidase | | Descriptor: | NEUTROPHIL CYTOSOL FACTOR 2, NEUTROPHIL CYTOSOL FACTOR 4 | | Authors: | Wilson, M.I, Gill, D.J, Perisic, O, Quinn, M.T, Williams, R.L. | | Deposit date: | 2003-04-02 | | Release date: | 2003-07-29 | | Last modified: | 2024-10-16 | | Method: | X-RAY DIFFRACTION (2 Å) | | Cite: | Pb1 Domain-Mediated Heterodimerization in Nadph Oxidase and Signaling Complexes of Atypical Protein Kinase C with Par6 and P62
Mol.Cell, 12, 2003
|
|
7TIK
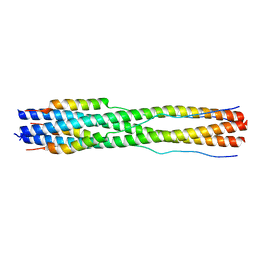 
 | |
5JQR
 
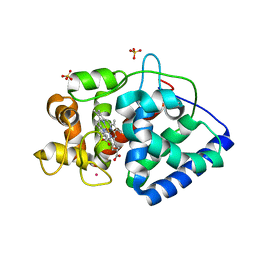 | | The Structure of Ascorbate Peroxidase Compound II formed by reaction with m-CPBA | | Descriptor: | Ascorbate peroxidase, POTASSIUM ION, PROTOPORPHYRIN IX CONTAINING FE, ... | | Authors: | Kwon, H, Raven, E.L, Moody, P.C.E. | | Deposit date: | 2016-05-05 | | Release date: | 2016-12-21 | | Last modified: | 2024-01-10 | | Method: | X-RAY DIFFRACTION (1.81 Å) | | Cite: | Direct visualization of a Fe(IV)-OH intermediate in a heme enzyme.
Nat Commun, 7, 2016
|
|
1UD1
 
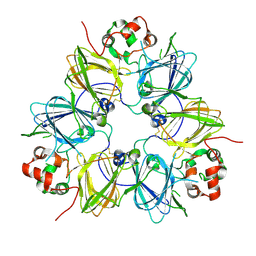 | | Crystal structure of proglycinin mutant C88S | | Descriptor: | Glycinin G1 | | Authors: | Utsumi, S, Adachi, M. | | Deposit date: | 2003-04-24 | | Release date: | 2004-02-03 | | Last modified: | 2024-10-30 | | Method: | X-RAY DIFFRACTION (3.1 Å) | | Cite: | Crystal Structures and Structural Stabilities of the Disulfide Bond-Deficient Soybean Proglycinin Mutants C12G and C88S.
J.Agric.Food Chem., 51, 2003
|
|
1U9Z
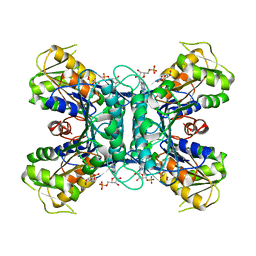 
 | | Crystal Structure of Phosphoribosyl Diphosphate Synthase Complexed with AMP and Ribose 5-Phosphate | | Descriptor: | ADENOSINE MONOPHOSPHATE, RIBOSE-5-PHOSPHATE, Ribose-phosphate pyrophosphokinase | | Authors: | Kadziola, A, Johansson, E, Jepsen, C.H, McGuire, J, Larsen, S, Hove-Jensen, B. | | Deposit date: | 2004-08-11 | | Release date: | 2005-08-23 | | Last modified: | 2023-08-23 | | Method: | X-RAY DIFFRACTION (2.8 Å) | | Cite: | Novel class III phosphoribosyl diphosphate synthase: structure and properties of the tetrameric, phosphate-activated, non-allosterically inhibited enzyme from Methanocaldococcus jannaschii
J.Mol.Biol., 354, 2005
|
|
1UCX
 
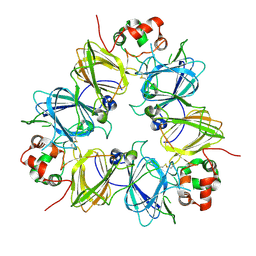 | | Crystal structure of proglycinin C12G mutant | | Descriptor: | Glycinin G1 | | Authors: | Utsumi, S, Adachi, M. | | Deposit date: | 2003-04-24 | | Release date: | 2004-02-03 | | Last modified: | 2024-10-30 | | Method: | X-RAY DIFFRACTION (3.2 Å) | | Cite: | Crystal Structures and Structural Stabilities of the Disulfide Bond-Deficient Soybean Proglycinin Mutants C12G and C88S.
J.Agric.Food Chem., 51, 2003
|
|
1FHF
 
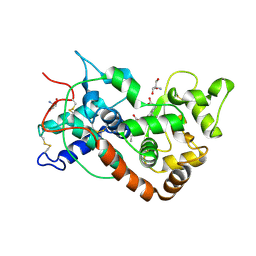 | | THE STRUCTURE OF SOYBEAN PEROXIDASE | | Descriptor: | 2-AMINO-2-HYDROXYMETHYL-PROPANE-1,3-DIOL, CALCIUM ION, PROTOPORPHYRIN IX CONTAINING FE, ... | | Authors: | Henriksen, A, Mirza, O, Indiana, C, Welinder, K, Teilum, K, Gajhede, M. | | Deposit date: | 2000-08-01 | | Release date: | 2001-02-14 | | Last modified: | 2024-10-30 | | Method: | X-RAY DIFFRACTION (2.8 Å) | | Cite: | Structure of soybean seed coat peroxidase: a plant peroxidase with unusual stability and haem-apoprotein interactions.
Protein Sci., 10, 2001
|
|
5JPR
 
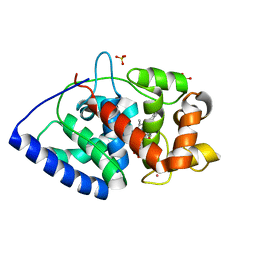 | | Neutron Structure of Compound II of Ascorbate Peroxidase | | Descriptor: | Ascorbate peroxidase, POTASSIUM ION, PROTOPORPHYRIN IX CONTAINING FE, ... | | Authors: | Kwon, H, Blakeley, M.P, Raven, E.L, Moody, P.C.E. | | Deposit date: | 2016-05-04 | | Release date: | 2016-12-21 | | Last modified: | 2024-05-01 | | Method: | NEUTRON DIFFRACTION (1.806 Å), X-RAY DIFFRACTION | | Cite: | Direct visualization of a Fe(IV)-OH intermediate in a heme enzyme.
Nat Commun, 7, 2016
|
|
8RP8
 
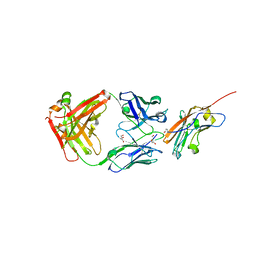 | | Structure of K2 Fab in complex with human CD47 ECD | | Descriptor: | 2-acetamido-2-deoxy-beta-D-glucopyranose, DI(HYDROXYETHYL)ETHER, GLYCEROL, ... | | Authors: | Laursen, M, Kelpsas, V, Rose, N. | | Deposit date: | 2024-01-12 | | Release date: | 2024-06-19 | | Last modified: | 2024-11-20 | | Method: | X-RAY DIFFRACTION (2 Å) | | Cite: | Structural analysis of light chain-driven bispecific antibodies targeting CD47 and PD-L1.
Mabs, 16, 2024
|
|
8RPB
 
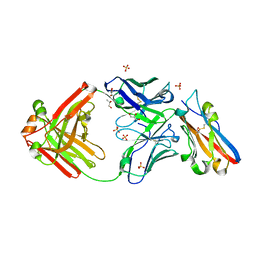 | | Structure of S79 Fab in complex with IgV domain of human PD-L1 | | Descriptor: | CHLORIDE ION, GLYCEROL, Programmed cell death 1 ligand 1, ... | | Authors: | Svensson, A, Kelpsas, V, Laursen, M, Rose, N. | | Deposit date: | 2024-01-15 | | Release date: | 2024-06-19 | | Last modified: | 2024-10-16 | | Method: | X-RAY DIFFRACTION (2.794 Å) | | Cite: | Structural analysis of light chain-driven bispecific antibodies targeting CD47 and PD-L1.
Mabs, 16, 2024
|
|
6TAE
 
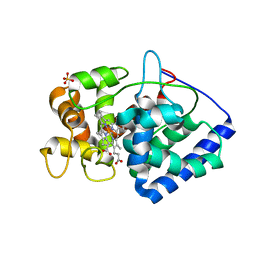 | | Neutron structure of ferric ascorbate peroxidase | | Descriptor: | Ascorbate peroxidase, PROTOPORPHYRIN IX CONTAINING FE, SULFATE ION | | Authors: | Kwon, H, Basran, J, Devos, J.M, Schrader, T.E, Ostermann, A, Blakeley, M.P, Raven, E.L, Moody, P.C.E. | | Deposit date: | 2019-10-29 | | Release date: | 2020-03-18 | | Last modified: | 2024-05-01 | | Method: | NEUTRON DIFFRACTION (1.9 Å), X-RAY DIFFRACTION | | Cite: | Visualizing the protons in a metalloenzyme electron proton transfer pathway.
Proc.Natl.Acad.Sci.USA, 117, 2020
|
|
2J4Q
 
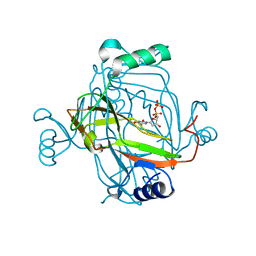 | | Crystal structure of a E138A Escherichia coli dCTP deaminase mutant enzyme in complex with dTTP | | Descriptor: | DEOXYCYTIDINE TRIPHOSPHATE DEAMINASE, MAGNESIUM ION, THYMIDINE-5'-DIPHOSPHATE, ... | | Authors: | Johansson, E, Thymark, M, Bynck, J.H, Fanoe, M, Larsen, S, Willemoes, M. | | Deposit date: | 2006-09-05 | | Release date: | 2007-08-07 | | Last modified: | 2023-12-13 | | Method: | X-RAY DIFFRACTION (2.6 Å) | | Cite: | Regulation of Dctp Deaminase from Escherichia Coli by Nonallosteric Dttp Binding to an Inactive Form of the Enzyme
FEBS J., 274, 2007
|
|
6O09
 
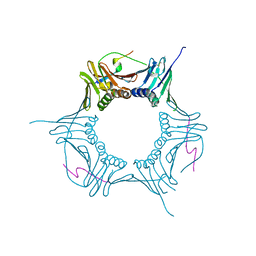 | |
2VCS
 
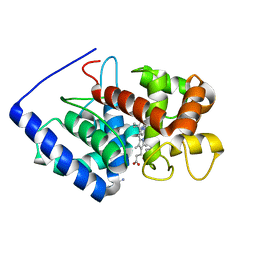 | | Structure of isoniazid (INH) bound to cytosolic soybean ascorbate peroxidase mutant H42A | | Descriptor: | 4-(DIAZENYLCARBONYL)PYRIDINE, ASCORBATE PEROXIDASE, PROTOPORPHYRIN IX CONTAINING FE, ... | | Authors: | Metcalfe, C.L, Macdonald, I.K, Brown, K.A, Raven, E.L, Moody, P.C.E. | | Deposit date: | 2007-09-26 | | Release date: | 2007-12-04 | | Last modified: | 2023-12-13 | | Method: | X-RAY DIFFRACTION (1.68 Å) | | Cite: | The Tuberculosis Prodrug Isoniazid Bound to Activating Peroxidases.
J.Biol.Chem., 283, 2008
|
|
