1KDW
 
 | |
1KDS
 
 | |
1X98
 
 | | Crystal structure of Aldose Reductase complexed with 2S4R (Stereoisomer of Fidarestat, 2S4S) | | Descriptor: | (2S,4R)-2-AMINOFORMYL-6-FLUORO-SPIRO[CHROMAN-4,4'-IMIDAZOLIDINE]-2',5'-DIONE, Aldose Reductase, CITRIC ACID, ... | | Authors: | El-Kabbani, O, Darmanin, C, Oka, M, Schulze-Briese, C, Tomizaki, T, Hazemann, I, Mitschler, A, Podjarny, A. | | Deposit date: | 2004-08-19 | | Release date: | 2004-09-07 | | Last modified: | 2023-10-25 | | Method: | X-RAY DIFFRACTION (1.3 Å) | | Cite: | High-Resolution Structures of Human Aldose Reductase Holoenzyme in Complex with Stereoisomers of the Potent Inhibitor Fidarestat: Stereospecific Interaction between the Enzyme and a Cyclic Imide Type Inhibitor
J.Med.Chem., 47, 2004
|
|
2CGU
 
 | | Identification of chemically diverse Chk1 inhibitors by receptor- based virtual screening | | Descriptor: | 2,2'-{[9-(HYDROXYIMINO)-9H-FLUORENE-2,7-DIYL]BIS(OXY)}DIACETIC ACID, SERINE/THREONINE-PROTEIN KINASE CHK1 | | Authors: | Foloppe, N, Fisher, L.M, Howes, R, Potter, A, Robertson, A.G.S, Surgenor, A.E. | | Deposit date: | 2006-03-09 | | Release date: | 2006-04-05 | | Last modified: | 2023-12-13 | | Method: | X-RAY DIFFRACTION (2.5 Å) | | Cite: | Identification of Chemically Diverse Chk1 Inhibitors by Receptor-Based Virtual Screening.
Bioorg.Med.Chem., 14, 2006
|
|
2CGV
 
 | | Identification of chemically diverse Chk1 inhibitors by receptor- based virtual screening | | Descriptor: | (2S)-1-AMINO-3-[(5-NITROQUINOLIN-8-YL)AMINO]PROPAN-2-OL, SERINE/THREONINE-PROTEIN KINASE CHK1 | | Authors: | Foloppe, N, Fisher, L.M, Howes, R, Potter, A, Robertson, A.G.S, Surgenor, A.E. | | Deposit date: | 2006-03-09 | | Release date: | 2006-04-05 | | Last modified: | 2023-12-13 | | Method: | X-RAY DIFFRACTION (2.6 Å) | | Cite: | Identification of Chemically Diverse Chk1 Inhibitors by Receptor-Based Virtual Screening.
Bioorg.Med.Chem., 14, 2006
|
|
2CGW
 
 | | Identification of chemically diverse Chk1 inhibitors by receptor- based virtual screening | | Descriptor: | 4,4'-(1-PROPYL-1H-1,2,4-TRIAZOLE-3,5-DIYL)BIS(2,5-DIHYDRO-1,2,5-OXADIAZOL-3-AMINE), SERINE/THREONINE-PROTEIN KINASE CHK1 | | Authors: | Foloppe, N, Fisher, L.M, Howes, R, Potter, A, Robertson, A.G.S, Surgenor, A.E. | | Deposit date: | 2006-03-09 | | Release date: | 2006-04-05 | | Last modified: | 2023-12-13 | | Method: | X-RAY DIFFRACTION (2.2 Å) | | Cite: | Identification of Chemically Diverse Chk1 Inhibitors by Receptor-Based Virtual Screening.
Bioorg.Med.Chem., 14, 2006
|
|
5D1U
 
 | | Anthrax toxin lethal factor with hydroxamic acid inhibitor | | Descriptor: | Lethal factor, N~2~-(6-aminohexyl)-N~2~-[(4-fluoro-3-methylphenyl)sulfonyl]-N-hydroxy-D-alaninamide, ZINC ION | | Authors: | Maize, K.M, Finzel, B.C. | | Deposit date: | 2015-08-04 | | Release date: | 2015-11-11 | | Last modified: | 2023-09-27 | | Method: | X-RAY DIFFRACTION (2.8503 Å) | | Cite: | Probing the S2' Subsite of the Anthrax Toxin Lethal Factor Using Novel N-Alkylated Hydroxamates.
J.Med.Chem., 58, 2015
|
|
7TRB
 
 | | CRYSTAL STRUCTURE OF FARNESOID X-ACTIVATED RECEPTOR COMPLEXED WITH COMPOUND-32 AKA (1S,3S)-N-({4-[5-(2-FLUOROPR OPAN-2-YL)-1,2,4-OXADIAZOL-3-YL]BICYCLO[2.2.2]OCTAN-1-YL}M ETHYL)-3-HYDROXY-N-[4'-(2-HYDROXYPROPAN-2-YL)-[1,1'-BIPHEN YL]-3-YL]-3-(TRIFLUOROMETHYL)CYCLOBUTANE-1-CARBOXAMIDE | | Descriptor: | (1s,3s)-N-({4-[5-(2-fluoropropan-2-yl)-1,2,4-oxadiazol-3-yl]bicyclo[2.2.2]octan-1-yl}methyl)-3-hydroxy-N-[4'-(2-hydroxypropan-2-yl)[1,1'-biphenyl]-3-yl]-3-(trifluoromethyl)cyclobutane-1-carboxamide, Bile acid receptor, co-activator | | Authors: | Khan, J.A, Ruzanov, M. | | Deposit date: | 2022-01-28 | | Release date: | 2022-06-29 | | Last modified: | 2023-10-18 | | Method: | X-RAY DIFFRACTION (2.15 Å) | | Cite: | Discovery of BMS-986339, a Pharmacologically Differentiated Farnesoid X Receptor Agonist for the Treatment of Nonalcoholic Steatohepatitis.
J.Med.Chem., 65, 2022
|
|
6M5D
 
 | | Human serum albumin (apo form) | | Descriptor: | PHOSPHATE ION, Serum albumin | | Authors: | Ito, S, Senoo, A, Nagatoishi, S, Yamamoto, M, Tsumoto, K, Wakui, N. | | Deposit date: | 2020-03-10 | | Release date: | 2020-11-18 | | Last modified: | 2023-11-29 | | Method: | X-RAY DIFFRACTION (2.6 Å) | | Cite: | Structural Basis for the Binding Mechanism of Human Serum Albumin Complexed with Cyclic Peptide Dalbavancin.
J.Med.Chem., 63, 2020
|
|
1KE4
 
 | |
7T79
 
 | |
7T78
 
 | |
7LS3
 
 | | Co-complex CYP46A1 with 8114 (3f) | | Descriptor: | (5-methyl-2-pyridin-4-yl-phenyl)-[4-oxidanyl-4-(phenylmethyl)piperidin-1-yl]methanone, Cholesterol 24-hydroxylase, PROTOPORPHYRIN IX CONTAINING FE | | Authors: | Lane, W, Yano, J. | | Deposit date: | 2021-02-17 | | Release date: | 2021-08-25 | | Last modified: | 2023-10-18 | | Method: | X-RAY DIFFRACTION (2.15 Å) | | Cite: | Discovery of Soticlestat, a Potent and Selective Inhibitor for Cholesterol 24-Hydroxylase (CH24H).
J.Med.Chem., 64, 2021
|
|
5D1T
 
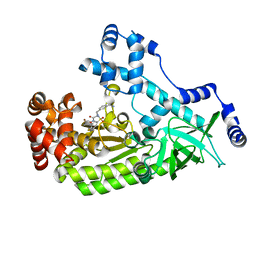 | | Anthrax toxin lethal factor with hydroxamic acid inhibitor | | Descriptor: | Lethal factor, N~2~-[3-(aminomethyl)benzyl]-N~2~-[(4-fluoro-3-methylphenyl)sulfonyl]-N-hydroxy-D-alaninamide, ZINC ION | | Authors: | Maize, K.M, Finzel, B.C. | | Deposit date: | 2015-08-04 | | Release date: | 2015-11-11 | | Last modified: | 2023-09-27 | | Method: | X-RAY DIFFRACTION (2.2 Å) | | Cite: | Probing the S2' Subsite of the Anthrax Toxin Lethal Factor Using Novel N-Alkylated Hydroxamates.
J.Med.Chem., 58, 2015
|
|
1UML
 
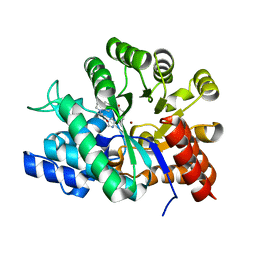 | | Crystal structure of adenosine deaminase complexed with a potent inhibitor FR233624 | | Descriptor: | 1-((1R)-1-(HYDROXYMETHYL)-3-{6-[(3-PHENYLPROPANOYL)AMINO]-1H-INDOL-1-YL}PROPYL)-1H-IMIDAZOLE-4-CARBOXAMIDE, Adenosine deaminase, ZINC ION | | Authors: | Kinoshita, T. | | Deposit date: | 2003-10-03 | | Release date: | 2004-09-21 | | Last modified: | 2023-12-27 | | Method: | X-RAY DIFFRACTION (2.5 Å) | | Cite: | Structure-based design, synthesis, and structure-activity relationship studies of novel non-nucleoside adenosine deaminase inhibitors
J.Med.Chem., 47, 2004
|
|
1O5M
 
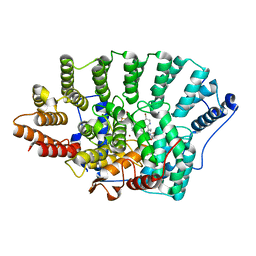 | | Structure of FPT bound to the inhibitor SCH66336 | | Descriptor: | 4-{2-[4-(3,10-DIBROMO-8-CHLORO-6,11-DIHYDRO-5H-BENZO[5,6]CYCLOHEPTA[1,2-B]PYRIDIN-11-YL)PIPERIDIN-1-YL]-2-OXOETHYL}PIPERIDINE-1-CARBOXAMIDE, FARNESYL DIPHOSPHATE, Protein farnesyltransferase alpha subunit, ... | | Authors: | Strickland, C.L, Weber, P.C, Ganguly, A.K. | | Deposit date: | 2003-09-26 | | Release date: | 2003-10-21 | | Last modified: | 2023-12-27 | | Method: | X-RAY DIFFRACTION (2.3 Å) | | Cite: | Tricyclic Farnesyl Protein Transferase Inhibitors: Crystallographic and Calorimetric Studies of Structure-Activity Relationships
J.Med.Chem., 42, 1999
|
|
7LTY
 
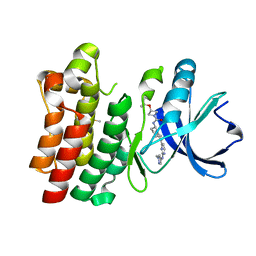 | | Bruton's tyrosine kinase in complex with compound 23 | | Descriptor: | DIMETHYL SULFOXIDE, Isoform BTK-C of Tyrosine-protein kinase BTK, ~{N}-[(5~{R})-2-[2-[(1-methylpyrazol-4-yl)amino]pyrimidin-4-yl]-6,7,8,9-tetrahydro-5~{H}-benzo[7]annulen-5-yl]-3-propan-2-yloxy-azetidine-1-carboxamide | | Authors: | Metrick, C.M, Marcotte, D.J. | | Deposit date: | 2021-02-20 | | Release date: | 2022-01-12 | | Last modified: | 2023-10-18 | | Method: | X-RAY DIFFRACTION (1.69 Å) | | Cite: | Discovery and Preclinical Characterization of BIIB091, a Reversible, Selective BTK Inhibitor for the Treatment of Multiple Sclerosis.
J.Med.Chem., 65, 2022
|
|
7LTZ
 
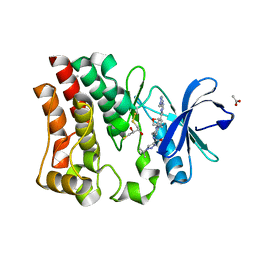 | | Bruton's tyrosine kinase in complex with compound 51 | | Descriptor: | 1-~{tert}-butyl-~{N}-[(5~{R})-8-[2-[(1-methylpyrazol-4-yl)amino]pyrimidin-4-yl]-2-(oxetan-3-yl)-1,3,4,5-tetrahydro-2-benzazepin-5-yl]-1,2,3-triazole-4-carboxamide, 2,3-DIHYDROXY-1,4-DITHIOBUTANE, ACETATE ION, ... | | Authors: | Metrick, C.M, Marcotte, D.J. | | Deposit date: | 2021-02-20 | | Release date: | 2022-01-12 | | Last modified: | 2023-10-18 | | Method: | X-RAY DIFFRACTION (1.53 Å) | | Cite: | Discovery and Preclinical Characterization of BIIB091, a Reversible, Selective BTK Inhibitor for the Treatment of Multiple Sclerosis.
J.Med.Chem., 65, 2022
|
|
7R3O
 
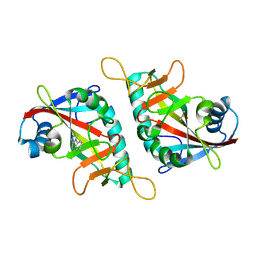 | | PARP15 catalytic domain in complex with OUL40 | | Descriptor: | 6-methyl-[1,2,4]triazolo[3,4-b][1,3]benzothiazole, DIMETHYL SULFOXIDE, Protein mono-ADP-ribosyltransferase PARP15 | | Authors: | Maksimainen, M.M, Murthy, S, Lehtio, L. | | Deposit date: | 2022-02-07 | | Release date: | 2023-01-25 | | Last modified: | 2024-02-07 | | Method: | X-RAY DIFFRACTION (2.2 Å) | | Cite: | [1,2,4]Triazolo[3,4- b ]benzothiazole Scaffold as Versatile Nicotinamide Mimic Allowing Nanomolar Inhibition of Different PARP Enzymes.
J.Med.Chem., 66, 2023
|
|
6RJZ
 
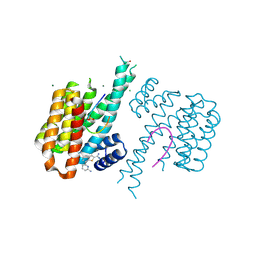 | | Fragment AZ-015 binding at the p53pT387/14-3-3 sigma interface | | Descriptor: | 14-3-3 protein sigma, 7-(6-azanylpyridin-2-yl)-1-benzothiophene-2-carboximidamide, CHLORIDE ION, ... | | Authors: | Genet, S, Wolter, M, Guillory, X, Somsen, B, Leysen, S, Castaldi, P, Ottmann, C. | | Deposit date: | 2019-04-29 | | Release date: | 2020-06-17 | | Last modified: | 2024-01-24 | | Method: | X-RAY DIFFRACTION (1.58 Å) | | Cite: | Fragment-based Differential Targeting of PPI Stabilizer Interfaces.
J.Med.Chem., 63, 2020
|
|
6RWI
 
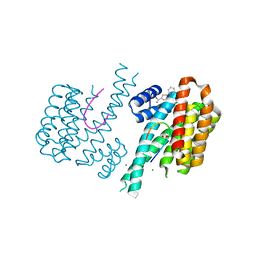 | | Fragment AZ-002 binding at the p53pT387/14-3-3 sigma interface | | Descriptor: | 14-3-3 protein sigma, 5-phenylthiophene-2-carboximidamide, CALCIUM ION, ... | | Authors: | Leysen, S, Guillory, X, Wolter, M, Genet, S, Somsen, B, Patel, J, Castaldi, P, Ottmann, C. | | Deposit date: | 2019-06-05 | | Release date: | 2020-06-17 | | Last modified: | 2024-01-24 | | Method: | X-RAY DIFFRACTION (1.65 Å) | | Cite: | Fragment-based Differential Targeting of PPI Stabilizer Interfaces.
J.Med.Chem., 63, 2020
|
|
6RWS
 
 | | Fragment AZ-009 binding at the p53pT387/14-3-3 sigma interface | | Descriptor: | 14-3-3 protein sigma, 4-chloranyl-6-(sulfanylmethyl)-1-benzothiophene-2-carboximidamide, Cellular tumor antigen p53, ... | | Authors: | Leysen, S, Guillory, X, Wolter, M, Genet, S, Somsen, B, Patel, J, Castaldi, P, Ottmann, C. | | Deposit date: | 2019-06-06 | | Release date: | 2020-06-17 | | Last modified: | 2024-01-24 | | Method: | X-RAY DIFFRACTION (1.53 Å) | | Cite: | Fragment-based Differential Targeting of PPI Stabilizer Interfaces.
J.Med.Chem., 63, 2020
|
|
6RKI
 
 | | Fragment AZ-023 binding at the p53pT387/14-3-3 sigma interface | | Descriptor: | 14-3-3 protein sigma, 5-[(3-aminophenyl)amino]-4-phenyl-thiophene-2-carboximidamide, CHLORIDE ION, ... | | Authors: | Genet, S, Wolter, M, Guillory, X, Somsen, B, Leysen, S, Castaldi, P, Ottmann, C. | | Deposit date: | 2019-04-30 | | Release date: | 2020-06-17 | | Last modified: | 2024-01-24 | | Method: | X-RAY DIFFRACTION (1.88 Å) | | Cite: | Fragment-based Differential Targeting of PPI Stabilizer Interfaces.
J.Med.Chem., 63, 2020
|
|
6RKM
 
 | | Fragment AZ-022 binding at the p53pT387/14-3-3 sigma interface | | Descriptor: | 14-3-3 protein sigma, 4-phenyl-5-phenylazanyl-thiophene-2-carboximidamide, CHLORIDE ION, ... | | Authors: | Genet, S, Wolter, M, Guillory, X, Somsen, B, Leysen, S, Patel, J, Castaldi, P, Ottmann, C. | | Deposit date: | 2019-04-30 | | Release date: | 2020-06-17 | | Last modified: | 2024-01-24 | | Method: | X-RAY DIFFRACTION (1.88 Å) | | Cite: | Fragment-based Differential Targeting of PPI Stabilizer Interfaces.
J.Med.Chem., 63, 2020
|
|
6RKQ
 
 | | Crystal Structure of TGT in complex with N2-methyl-8-(prop-1-yn-1-yl)-3H,7H,8H-imidazo[4,5-g]quinazoline-2,6-diamine | | Descriptor: | (8~{R})-~{N}2-methyl-8-prop-1-ynyl-7,8-dihydro-3~{H}-imidazo[4,5-g]quinazoline-2,6-diamine, DIMETHYL SULFOXIDE, GLYCEROL, ... | | Authors: | Hassaan, E, Heine, A, Klebe, G. | | Deposit date: | 2019-04-30 | | Release date: | 2020-06-03 | | Last modified: | 2024-01-24 | | Method: | X-RAY DIFFRACTION (1.665 Å) | | Cite: | Fragment Screening Hit Draws Attention to a Novel Transient Pocket Adjacent to the Recognition Site of the tRNA-Modifying Enzyme TGT.
J.Med.Chem., 63, 2020
|
|
