8J6V
 
 | |
2ZKO
 
 | |
3ADI
 
 | |
3ADL
 
 | |
2N3R
 
 | | NMR structure of the II-III-VI three-way junction from the VS ribozyme and identification of magnesium-binding sites using paramagnetic relaxation enhancement | | Descriptor: | MAGNESIUM ION, RNA (62-MER) | | Authors: | Bonneau, E, Girard, N, Lemieux, S, Legault, P. | | Deposit date: | 2015-06-09 | | Release date: | 2015-07-15 | | Last modified: | 2024-05-01 | | Method: | SOLUTION NMR | | Cite: | The NMR structure of the II-III-VI three-way junction from the Neurospora VS ribozyme reveals a critical tertiary interaction and provides new insights into the global ribozyme structure.
Rna, 21, 2015
|
|
4CSF
 
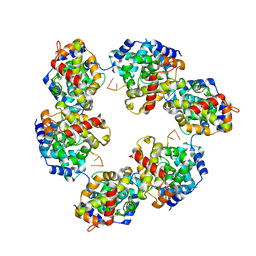 | | Structural insights into Toscana virus RNA encapsidation | | Descriptor: | NUCLEOPROTEIN, RNA (5'-R(*UP*GP*UP*GP*UP*UP*UP*CP*UP)-3') | | Authors: | Olal, D, Daumke, O. | | Deposit date: | 2014-03-07 | | Release date: | 2014-04-09 | | Last modified: | 2024-05-08 | | Method: | X-RAY DIFFRACTION (2.598 Å) | | Cite: | Structural Insights Into RNA Encapsidation and Helical Assembly of the Toscana Virus Nucleoprotein.
Nucleic Acids Res., 42, 2014
|
|
2ZI0
 
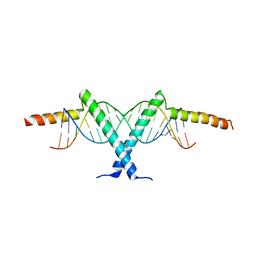 | | Crystal structure of Tav2b/siRNA complex | | Descriptor: | Protein 2b, RNA (5'-D(P*AP*GP*AP*CP*AP*GP*CP*AP*UP*UP*AP*UP*GP*CP*UP*GP*UP*CP*UP*UP*U)-3') | | Authors: | Yuan, Y.A, Chen, H.-Y. | | Deposit date: | 2008-02-12 | | Release date: | 2008-07-22 | | Last modified: | 2024-10-16 | | Method: | X-RAY DIFFRACTION (2.82 Å) | | Cite: | Structural basis for RNA-silencing suppression by Tomato aspermy virus protein 2b
Embo Rep., 9, 2008
|
|
6D12
 
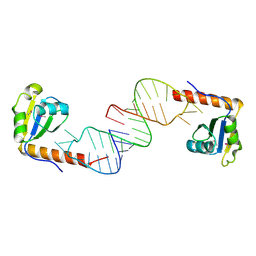 | | Crystal structure of C-terminal xRRM domain of human Larp7 bound to 7SK stem-loop 4 RNA | | Descriptor: | La-related protein 7, human 7SK RNA stem-loop 4 | | Authors: | Eichhorn, C.D, Cascio, D, Sawaya, M.R, Feigon, J. | | Deposit date: | 2018-04-11 | | Release date: | 2018-06-27 | | Last modified: | 2024-10-23 | | Method: | X-RAY DIFFRACTION (2.205 Å) | | Cite: | Structural basis for recognition of human 7SK long noncoding RNA by the La-related protein Larp7.
Proc. Natl. Acad. Sci. U.S.A., 115, 2018
|
|
1BNI
 
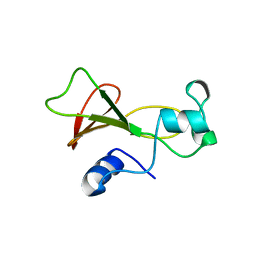 | | BARNASE WILDTYPE STRUCTURE AT PH 6.0 | | Descriptor: | BARNASE | | Authors: | Cameron, A, Henrick, K, Fersht, A.R, Dodson, G, Buckle, A.M. | | Deposit date: | 1995-05-17 | | Release date: | 1995-09-15 | | Last modified: | 2024-02-07 | | Method: | X-RAY DIFFRACTION (2.1 Å) | | Cite: | Crystal structural analysis of mutations in the hydrophobic cores of barnase.
J.Mol.Biol., 234, 1993
|
|
310D
 
 | | Crystal structure of 2'-O-Me(CGCGCG)2: an RNA duplex at 1.3 A resolution. Hydration pattern of 2'-O-methylated RNA | | Descriptor: | MAGNESIUM ION, RNA (5'-R(*(OMC)P*(OMG)P*(OMC)P*(OMG)P*(OMC)P*(OMG))-3') | | Authors: | Adamiak, D.A, Milecki, J, Adamiak, R.W, Dauter, Z, Rypniewski, W.R. | | Deposit date: | 1996-05-25 | | Release date: | 1997-06-24 | | Last modified: | 2024-04-03 | | Method: | X-RAY DIFFRACTION (1.3 Å) | | Cite: | Crystal structure of 2'-O-Me(CGCGCG)2, an RNA duplex at 1.30 A resolution. Hydration pattern of 2'-O-methylated RNA.
Nucleic Acids Res., 25, 1997
|
|
8Q4O
 
 | |
4C7O
 
 | | The structural basis of FtsY recruitment and GTPase activation by SRP RNA | | Descriptor: | GUANOSINE-5'-DIPHOSPHATE, MAGNESIUM ION, SIGNAL RECOGNITION PARTICLE PROTEIN, ... | | Authors: | Voigts-Hoffmann, F, Schmitz, N, Shen, K, Shan, S.O, Ataide, S.F, Ban, N. | | Deposit date: | 2013-09-23 | | Release date: | 2013-11-20 | | Last modified: | 2023-12-20 | | Method: | X-RAY DIFFRACTION (2.6 Å) | | Cite: | The Structural Basis of Ftsy Recruitment and Gtpase Activation by Srp RNA
Mol.Cell, 52, 2013
|
|
4N0I
 
 | | Crystal structure of S. cerevisiae mitochondrial GatFAB in complex with glutamine | | Descriptor: | GLUTAMINE, Glutamyl-tRNA(Gln) amidotransferase subunit A, mitochondrial, ... | | Authors: | Araiso, Y, Ishitani, R, Nureki, O. | | Deposit date: | 2013-10-02 | | Release date: | 2014-04-16 | | Last modified: | 2024-04-03 | | Method: | X-RAY DIFFRACTION (2.001 Å) | | Cite: | Crystal structure of Saccharomyces cerevisiae mitochondrial GatFAB reveals a novel subunit assembly in tRNA-dependent amidotransferases
Nucleic Acids Res., 42, 2014
|
|
8SH4
 
 | | Crystal structure of the tRNA (m1G37) methyltransferase apoenzyme from Anaplasma phagocytophilum | | Descriptor: | GLYCEROL, tRNA (guanine-N(1)-)-methyltransferase | | Authors: | Jannotta, C, Edele, D, Levanti, D, Carson, M, Prucha, G, Caesar, J, Picchiello, C, Collins, K, Garland, E, Handley-Pendleton, J, Hernandez, V, Leffler, S, Williams, D, Stojanoff, V, Perez, A, Halloran, J, Bolen, R. | | Deposit date: | 2023-04-13 | | Release date: | 2024-04-17 | | Method: | X-RAY DIFFRACTION (2.9 Å) | | Cite: | Crystal structure of the m1G37 tRNA methyltransferase apoenzyme from Anaplasma phagocytophilum
To Be Published
|
|
2M58
 
 | | Structure of 2'-5' AG1 lariat forming ribozyme in its inactive state | | Descriptor: | RNA (59-MER) | | Authors: | Carlomagno, T, Amata, I, Codutti, L, Falb, M, Fohrer, J, Simon, B. | | Deposit date: | 2013-02-18 | | Release date: | 2013-04-03 | | Last modified: | 2024-05-15 | | Method: | SOLUTION NMR | | Cite: | Structural principles of RNA catalysis in a 2'-5' lariat-forming ribozyme.
J.Am.Chem.Soc., 135, 2013
|
|
315D
 
 | |
4H5P
 
 | |
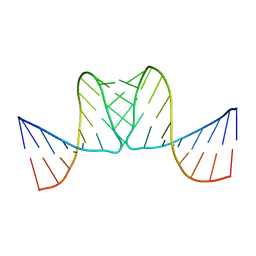
2BJ2
 | |
2M1V
 
 | |
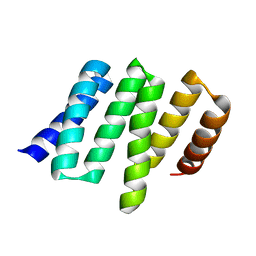
4YVO
 | |
1VOP
 
 | |
3WND
 
 | | Crystal structure of EF-Pyl | | Descriptor: | CITRIC ACID, Protein translation elongation factor 1A | | Authors: | Yanagisawa, T, Ishii, R, Fukunaga, R, Sengoku, T, Yokoyama, S, RIKEN Structural Genomics/Proteomics Initiative (RSGI) | | Deposit date: | 2013-12-08 | | Release date: | 2014-12-10 | | Last modified: | 2023-11-08 | | Method: | X-RAY DIFFRACTION (1.55 Å) | | Cite: | A SelB/EF-Tu/aIF2 gamma-like protein from Methanosarcina mazei in the GTP-bound form binds cysteinyl-tRNA(Cys.).
J. Struct. Funct. Genomics, 16, 2015
|
|
3HYF
 
 | | Crystal structure of HIV-1 RNase H p15 with engineered E. coli loop and active site inhibitor | | Descriptor: | 2-(3,4-dichlorobenzyl)-5,6-dihydroxypyrimidine-4-carboxylic acid, ACETATE ION, GLYCEROL, ... | | Authors: | Lansdon, E.B, Kirschberg, T.A. | | Deposit date: | 2009-06-22 | | Release date: | 2009-10-20 | | Last modified: | 2024-03-13 | | Method: | X-RAY DIFFRACTION (1.7 Å) | | Cite: | RNase H active site inhibitors of human immunodeficiency virus type 1 reverse transcriptase: design, biochemical activity, and structural information.
J.Med.Chem., 52, 2009
|
|
11AS
 
 | |
5V59
 
 | |
