2B4M
 
 | | Crystal structure of the binding protein OpuAC in complex with proline betaine | | Descriptor: | 1,1-DIMETHYL-PROLINIUM, Glycine betaine-binding protein | | Authors: | Horn, C, Sohn-Boesser, L, Breed, J, Welte, W, Schmitt, L, Bremer, E. | | Deposit date: | 2005-09-26 | | Release date: | 2006-03-21 | | Last modified: | 2023-08-23 | | Method: | X-RAY DIFFRACTION (2.8 Å) | | Cite: | Molecular Determinants for Substrate Specificity of the Ligand-binding Protein OpuAC from Bacillus subtilis for the Compatible Solutes Glycine Betaine and Proline Betaine.
J.Mol.Biol., 357, 2006
|
|
2B4N
 
 | |
2B4O
 
 | |
2B4P
 
 | |
2B4Q
 
 | | Pseudomonas aeruginosa RhlG/NADP active-site complex | | Descriptor: | NADP NICOTINAMIDE-ADENINE-DINUCLEOTIDE PHOSPHATE, Rhamnolipids biosynthesis 3-oxoacyl-[acyl-carrier-protein] reductase | | Authors: | Miller, D.J, White, S.W. | | Deposit date: | 2005-09-26 | | Release date: | 2006-05-02 | | Last modified: | 2024-02-14 | | Method: | X-RAY DIFFRACTION (2.3 Å) | | Cite: | Structure of RhlG, an Essential beta-Ketoacyl Reductase in the Rhamnolipid Biosynthetic Pathway of Pseudomonas aeruginosa.
J.Biol.Chem., 281, 2006
|
|
2B4R
 
 | | Crystal structure of glyceraldehyde-3-phosphate dehydrogenase from Plasmodium falciparum at 2.25 Angstrom Resolution reveals intriguing extra electron density in the active site | | Descriptor: | 4-(2-AMINOETHYL)BENZENESULFONYL FLUORIDE, GLYCEROL, NICOTINAMIDE-ADENINE-DINUCLEOTIDE, ... | | Authors: | Robien, M.A, Bosch, J, Hol, W.G.J, Structural Genomics of Pathogenic Protozoa Consortium (SGPP) | | Deposit date: | 2005-09-26 | | Release date: | 2005-10-04 | | Last modified: | 2023-08-23 | | Method: | X-RAY DIFFRACTION (2.25 Å) | | Cite: | Crystal structure of glyceraldehyde-3-phosphate dehydrogenase from Plasmodium falciparum at 2.25 A resolution reveals intriguing extra electron density in the active site
Proteins, 62, 2006
|
|
2B4S
 
 | | Crystal structure of a complex between PTP1B and the insulin receptor tyrosine kinase | | Descriptor: | Insulin receptor, SULFATE ION, Tyrosine-protein phosphatase, ... | | Authors: | Li, S, Depetris, R.S, Barford, D, Chernoff, J, Hubbard, S.R. | | Deposit date: | 2005-09-26 | | Release date: | 2005-11-15 | | Last modified: | 2023-11-15 | | Method: | X-RAY DIFFRACTION (2.3 Å) | | Cite: | Crystal Structure of a Complex between Protein Tyrosine Phosphatase 1B and the Insulin Receptor Tyrosine Kinase.
Structure, 13, 2005
|
|
2B4T
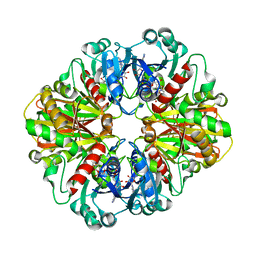 
 | | Crystal structure of glyceraldehyde-3-phosphate dehydrogenase from Plasmodium falciparum at 2.25 Angstrom resolution reveals intriguing extra electron density in the active site | | Descriptor: | 4-(2-AMINOETHYL)BENZENESULFONYL FLUORIDE, NICOTINAMIDE-ADENINE-DINUCLEOTIDE, glyceraldehyde-3-phosphate dehydrogenase | | Authors: | Robien, M.A, Bosch, J, Hol, W.G.J, Structural Genomics of Pathogenic Protozoa Consortium (SGPP) | | Deposit date: | 2005-09-26 | | Release date: | 2005-10-04 | | Last modified: | 2023-08-23 | | Method: | X-RAY DIFFRACTION (2.5 Å) | | Cite: | Crystal structure of glyceraldehyde-3-phosphate dehydrogenase from Plasmodium falciparum at 2.25 A resolution reveals intriguing extra electron density in the active site
Proteins, 62, 2006
|
|
2B4U
 
 | | Structure of the C252S mutant of Selenomonas ruminantium PTP-like phytase | | Descriptor: | CHLORIDE ION, MALONATE ION, SULFATE ION, ... | | Authors: | Gruninger, R.J, Selinger, L.B, Mosimann, S.C. | | Deposit date: | 2005-09-26 | | Release date: | 2006-11-07 | | Last modified: | 2021-10-20 | | Method: | X-RAY DIFFRACTION (2 Å) | | Cite: | Kinetic and structural analysis of a bacterial protein tyrosine phosphatase-like myo-inositol polyphosphatase.
Protein Sci., 16, 2007
|
|
2B4V
 
 | | Structural Basis for UTP Specificity of RNA Editing TUTases From Trypanosoma Brucei | | Descriptor: | POTASSIUM ION, RNA editing complex protein MP57 | | Authors: | Deng, J, Ernst, N.L, Turley, S, Stuart, K.D, Hol, W.G. | | Deposit date: | 2005-09-26 | | Release date: | 2005-11-22 | | Last modified: | 2021-10-20 | | Method: | X-RAY DIFFRACTION (1.8 Å) | | Cite: | Structural basis for UTP specificity of RNA editing TUTases from Trypanosoma brucei.
Embo J., 24, 2005
|
|
2B4W
 
 | |
2B4X
 
 | |
2B4Y
 
 | | Crystal Structure of Human Sirtuin homolog 5 | | Descriptor: | 4-(2-HYDROXYETHYL)-1-PIPERAZINE ETHANESULFONIC ACID, ADENOSINE-5-DIPHOSPHORIBOSE, NAD-dependent deacetylase sirtuin-5, ... | | Authors: | Min, J.R, Antoshenko, T, Dong, A, Schuetz, A, Loppnau, P, Weigelt, J, Sundstrom, M, Arrowsmith, C.H, Edwards, A.M, Bochkarev, A, Plotnikov, A.N, Structural Genomics Consortium (SGC) | | Deposit date: | 2005-09-27 | | Release date: | 2006-02-28 | | Last modified: | 2023-11-15 | | Method: | X-RAY DIFFRACTION (1.9 Å) | | Cite: | Crystal Structure of Human Sirtuin homolog 5 in complex with NAD
To be Published
|
|
2B4Z
 
 | | Crystal structure of cytochrome C from bovine heart at 1.5 A resolution. | | Descriptor: | Cytochrome c, PROTOPORPHYRIN IX CONTAINING FE | | Authors: | Mirkin, N, Jakoncic, J, Stojanoff, V, Moreno, A. | | Deposit date: | 2005-09-27 | | Release date: | 2005-10-11 | | Last modified: | 2023-08-23 | | Method: | X-RAY DIFFRACTION (1.5 Å) | | Cite: | High resolution X-ray crystallographic structure of bovine heart cytochrome c and its application to the design of an electron transfer biosensor.
Proteins, 70, 2008
|
|
2B50
 
 | | Human Nuclear Receptor-Ligand Complex 2 | | Descriptor: | CALCIUM ION, Peroxisome proliferator activated receptor delta, VACCENIC ACID, ... | | Authors: | Fyffe, S.A, Alphey, M.S, Buetow, L, Smith, T.K, Ferguson, M.A.J, Sorensen, M.D, Bjorkling, F, Hunter, W.N. | | Deposit date: | 2005-09-27 | | Release date: | 2006-02-14 | | Last modified: | 2024-03-13 | | Method: | X-RAY DIFFRACTION (2 Å) | | Cite: | Recombinant Human PPAR-beta/delta Ligand-binding Domain is Locked in an Activated Conformation by Endogenous Fatty Acids
J.Mol.Biol., 356, 2006
|
|
2B51
 
 | | Structural Basis for UTP Specificity of RNA Editing TUTases from Trypanosoma Brucei | | Descriptor: | MANGANESE (II) ION, RNA editing complex protein MP57, URIDINE 5'-TRIPHOSPHATE | | Authors: | Deng, J, Ernst, N.L, Turley, S, Stuart, K.D, Hol, W.G. | | Deposit date: | 2005-09-27 | | Release date: | 2005-11-22 | | Last modified: | 2024-02-14 | | Method: | X-RAY DIFFRACTION (2.05 Å) | | Cite: | Structural basis for UTP specificity of RNA editing TUTases from Trypanosoma brucei.
Embo J., 24, 2005
|
|
2B52
 
 | | Human cyclin dependent kinase 2 (CDK2) complexed with DPH-042562 | | Descriptor: | 1-(3-(2,4-DIMETHYLTHIAZOL-5-YL)-4-OXO-2,4-DIHYDROINDENO[1,2-C]PYRAZOL-5-YL)-3-(4-METHYLPIPERAZIN-1-YL)UREA, Cell division protein kinase 2 | | Authors: | Muckelbauer, J. | | Deposit date: | 2005-09-27 | | Release date: | 2005-10-11 | | Last modified: | 2024-02-14 | | Method: | X-RAY DIFFRACTION (1.88 Å) | | Cite: | Synthesis and Evaluation of Indenopyrazoles as Cyclin-Dependent Kinase Inhibitors. Part 4: Heterocycles at C3
Bioorg.Med.Chem.Lett., 14, 2004
|
|
2B53
 
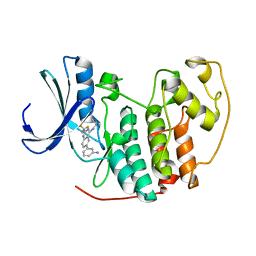 | |
2B54
 
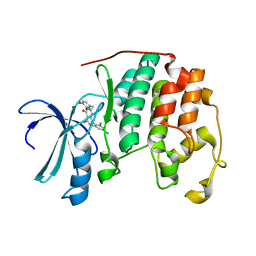 | | Human cyclin dependent kinase 2 (CKD2)complexed with DIN-232305 | | Descriptor: | 6-(3,4-DIHYDROXYBENZYL)-3-ETHYL-1-(2,4,6-TRICHLOROPHENYL)-1H-PYRAZOLO[3,4-D]PYRIMIDIN-4(5H)-ONE, Cell division protein kinase 2 | | Authors: | Chang, C.-C. | | Deposit date: | 2005-09-27 | | Release date: | 2005-10-11 | | Last modified: | 2024-04-03 | | Method: | X-RAY DIFFRACTION (1.85 Å) | | Cite: | Synthesis and biological evaluation of 1-aryl-4,5-dihydro-1h-pyraxolo[3,4-d]pyrimidin-4-one inhibitors of cyclin dependent kinases
J.Med.Chem., 47, 2004
|
|
2B55
 
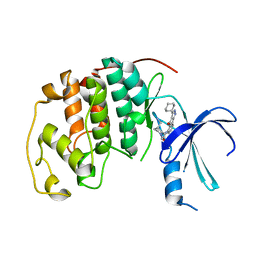 | | Human cyclin dependent kinase 2 (cdk2) complexed with indenopyraxole DIN-101312 | | Descriptor: | 2-(4-(AMINOMETHYL)PIPERIDIN-1-YL)-N-(3_CYCLOHEXYL-4-OXO-2,4-DIHYDROINDENO[1,2-C]PYRAZOL-5-YL)ACETAMIDE, Cell division protein kinase 2 | | Authors: | Muckelbauer, J. | | Deposit date: | 2005-09-27 | | Release date: | 2005-10-11 | | Last modified: | 2024-02-14 | | Method: | X-RAY DIFFRACTION (1.85 Å) | | Cite: | Synthesis and Evaluation of Indenopyrazoles as Cyclin-Dependent Kinase Inhibitors. 3. Structure Activity Relationships at C3
J.Med.Chem., 45, 2002
|
|
2B56
 
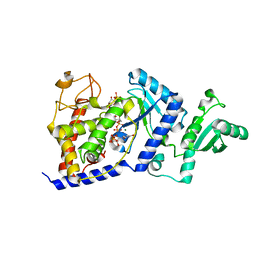 | | Structural Basis for UTP Specificity of RNA Editing TUTases From Trypanosoma Brucei | | Descriptor: | MAGNESIUM ION, RNA editing complex protein MP57, URIDINE 5'-TRIPHOSPHATE, ... | | Authors: | Deng, J, Ernst, N.L, Turley, S, Stuart, K.D, Hol, W.G. | | Deposit date: | 2005-09-27 | | Release date: | 2005-11-22 | | Last modified: | 2024-02-14 | | Method: | X-RAY DIFFRACTION (1.97 Å) | | Cite: | Structural basis for UTP specificity of RNA editing TUTases from Trypanosoma brucei.
Embo J., 24, 2005
|
|
2B57
 
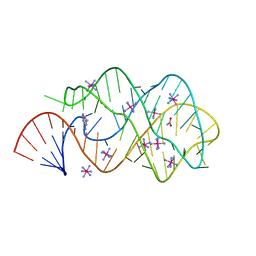 | | Guanine Riboswitch C74U mutant bound to 2,6-diaminopurine | | Descriptor: | 65-MER, 9H-PURINE-2,6-DIAMINE, ACETATE ION, ... | | Authors: | Gilbert, S.D, Stoddard, C.D, Wise, S.J, Batey, R.T. | | Deposit date: | 2005-09-27 | | Release date: | 2006-05-23 | | Last modified: | 2024-02-14 | | Method: | X-RAY DIFFRACTION (2.15 Å) | | Cite: | Thermodynamic and kinetic characterization of ligand binding to the purine riboswitch aptamer domain.
J.Mol.Biol., 359, 2006
|
|
2B58
 
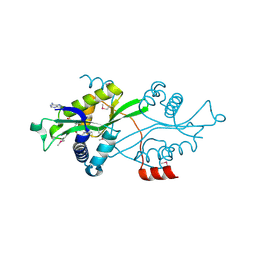 | | SSAT with coa_sp, spermine disordered, K26R mutant | | Descriptor: | COENZYME A, Diamine acetyltransferase 1 | | Authors: | Bewley, M.C, Graziano, V, Jiang, J.S, Matz, E, Studier, F.W, Pegg, A.P, Coleman, C.S, Flanagan, J.M, Burley, S.K, New York SGX Research Center for Structural Genomics (NYSGXRC) | | Deposit date: | 2005-09-27 | | Release date: | 2006-01-24 | | Last modified: | 2021-02-03 | | Method: | X-RAY DIFFRACTION (1.95 Å) | | Cite: | Structures of wild-type and mutant human spermidine/spermine N1-acetyltransferase, a potential therapeutic drug target
Proc.Natl.Acad.Sci.Usa, 103, 2006
|
|
2B59
 
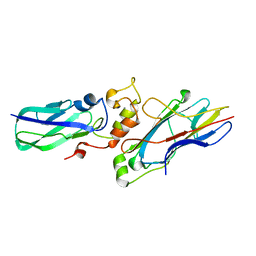 | |
2B5A
 
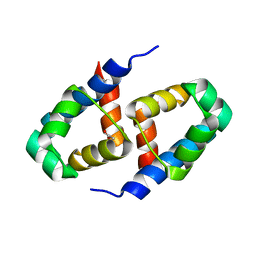 | | C.BclI, Control Element of the BclI Restriction-Modification System | | Descriptor: | ACETIC ACID, C.BclI | | Authors: | Sawaya, M.R, Zhu, Z, Mersha, F, Chan, S.H, Dabur, R, Xu, S.Y, Balendiran, G.K. | | Deposit date: | 2005-09-28 | | Release date: | 2006-01-03 | | Last modified: | 2024-02-14 | | Method: | X-RAY DIFFRACTION (1.543 Å) | | Cite: | Crystal Structure of the Restriction-Modification System Control Element C.BclI and Mapping of Its Binding Site.
Structure, 13, 2005
|
|
