1S7A
 
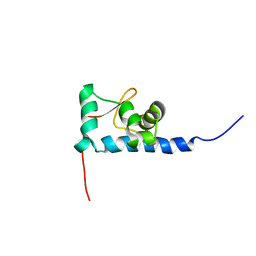 | | NMR structure of the La motif of human La protein | | Descriptor: | Lupus La protein | | Authors: | Alfano, C, Sanfelice, D, Babon, J, Kelly, G, Jacks, A, Curry, S, Conte, M.R. | | Deposit date: | 2004-01-29 | | Release date: | 2004-04-06 | | Last modified: | 2024-05-22 | | Method: | SOLUTION NMR | | Cite: | Structural analysis of cooperative RNA binding by the La motif and central RRM domain of human La protein.
Nat.Struct.Mol.Biol., 11, 2004
|
|
7OB2
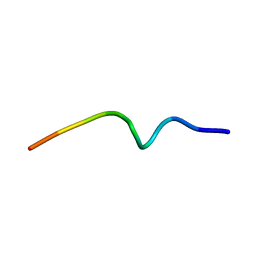 
 | | NMR structure of the antimicrobial RiLK1 peptide in SDS micelles | | Descriptor: | RiLK1 | | Authors: | Falcigno, L, D'Auria, G, Palmieri, G, Gogliettino, M, Agrillo, B. | | Deposit date: | 2021-04-20 | | Release date: | 2021-11-17 | | Last modified: | 2024-06-19 | | Method: | SOLUTION NMR | | Cite: | Key Physicochemical Determinants in the Antimicrobial Peptide RiLK1 Promote Amphipathic Structures.
Int J Mol Sci, 22, 2021
|
|
7L8V
 
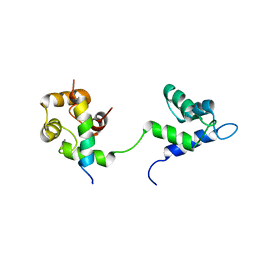 | |
1LUI
 
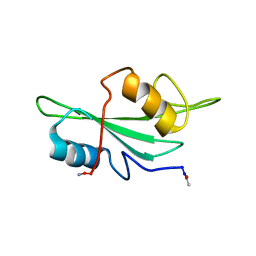 | | NMR Structures of Itk SH2 domain, Pro287cis isoform, ensemble of 20 low energy structures | | Descriptor: | Tyrosine-protein kinase ITK/TSK | | Authors: | Mallis, R.J, Brazin, K.N, Fulton, B.F, Andreotti, A.M. | | Deposit date: | 2002-05-22 | | Release date: | 2002-11-27 | | Last modified: | 2022-02-23 | | Method: | SOLUTION NMR | | Cite: | Structural characterization of a proline-driven conformational switch
within the Itk SH2 domain
Nat.Struct.Biol., 9, 2002
|
|
1LUK
 
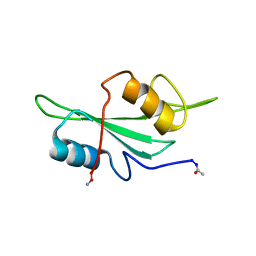 | | NMR Structure of the Itk SH2 domain, Pro287cis, Energy minimized average structure | | Descriptor: | Tyrosine-protein kinase ITK/TSK | | Authors: | Mallis, R.J, Brazin, K.N, Fulton, B.F, Andreotti, A.H. | | Deposit date: | 2002-05-22 | | Release date: | 2002-11-27 | | Last modified: | 2022-02-23 | | Method: | SOLUTION NMR | | Cite: | Structural characterization of a proline-driven conformational switch
within the Itk SH2 domain
Nat.Struct.Biol., 9, 2002
|
|
1LUN
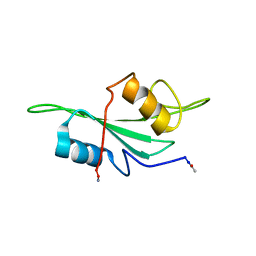 
 | | NMR Structure of the Itk SH2 domain, Pro287trans, energy minimized average structure | | Descriptor: | Tyrosine-protein kinase ITK/TSK | | Authors: | Mallis, R.J, Brazin, K.N, Fulton, D.B, Andreotti, A.H. | | Deposit date: | 2002-05-22 | | Release date: | 2002-11-27 | | Last modified: | 2022-02-23 | | Method: | SOLUTION NMR | | Cite: | Structural characterization of a proline-driven conformational switch
within the Itk SH2 domain
Nat.Struct.Biol., 9, 2002
|
|
1LUM
 
 | | NMR Structure of the Itk SH2 domain, Pro287trans, 20 low energy structures | | Descriptor: | Tyrosine-protein kinase ITK/TSK | | Authors: | Mallis, R.J, Brazin, K.N, Fulton, D.B, Andreotti, A.H. | | Deposit date: | 2002-05-22 | | Release date: | 2002-11-27 | | Last modified: | 2022-02-23 | | Method: | SOLUTION NMR | | Cite: | Structural characterization of a proline-driven conformational switch
within the Itk SH2 domain
Nat.Struct.Biol., 9, 2002
|
|
1EKH
 
 | | NMR STRUCTURE OF D(TTGGCCAA)2 BOUND TO CHROMOMYCIN-A3 AND COBALT | | Descriptor: | (1S)-5-deoxy-1-O-methyl-1-C-[(2R,3S)-3,5,7,10-tetrahydroxy-6-methyl-4-oxo-1,2,3,4-tetrahydroanthracen-2-yl]-D-xylulose, 2,6-dideoxy-4-O-methyl-alpha-D-galactopyranose-(1-3)-4-O-acetyl-2,6-dideoxy-beta-D-galactopyranose, 3-C-methyl-4-O-acetyl-alpha-L-Olivopyranose-(1-3)-beta-D-Olivopyranose-(1-3)-beta-D-Olivopyranose, ... | | Authors: | Gochin, M. | | Deposit date: | 2000-03-08 | | Release date: | 2000-03-20 | | Last modified: | 2024-05-01 | | Method: | SOLUTION NMR | | Cite: | A high-resolution structure of a DNA-chromomycin-Co(II) complex determined from pseudocontact shifts in nuclear magnetic resonance.
Structure Fold.Des., 8, 2000
|
|
1DG4
 
 | | NMR STRUCTURE OF THE SUBSTRATE BINDING DOMAIN OF DNAK IN THE APO FORM | | Descriptor: | DNAK | | Authors: | Pellecchia, M, Montgomery, D.L, Stevens, S.Y, Van der Kooi, C.W, Feng, H, Gierasch, L.M, Zuiderweg, E.R.P. | | Deposit date: | 1999-11-23 | | Release date: | 1999-12-08 | | Last modified: | 2024-05-22 | | Method: | SOLUTION NMR | | Cite: | Structural insights into substrate binding by the molecular chaperone DnaK.
Nat.Struct.Biol., 7, 2000
|
|
1HCS
 
 | |
1HCT
 
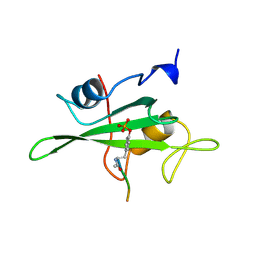 | |
1JAU
 | |
7ALU
 
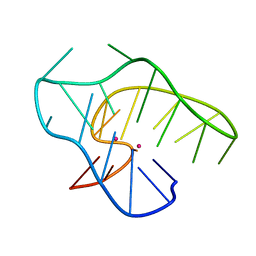 | |
7AY8
 
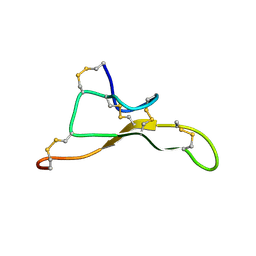 | | NMR solution structure of Tbo-IT2 | | Descriptor: | Tbo-IT2 | | Authors: | Mineev, K.S, Kornilov, F.D, Lushpa, V.A, Korolkova, Y.A, Maleeva, E.E, Andreev, Y.A, Kozlov, S.A, Arseniev, A.S. | | Deposit date: | 2020-11-11 | | Release date: | 2021-03-03 | | Last modified: | 2023-06-14 | | Method: | SOLUTION NMR | | Cite: | New Insectotoxin from Tibellus Oblongus Spider Venom Presents Novel Adaptation of ICK Fold.
Toxins, 13, 2021
|
|
1J5L
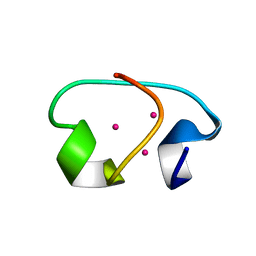 
 | | NMR STRUCTURE OF THE ISOLATED BETA_C DOMAIN OF LOBSTER METALLOTHIONEIN-1 | | Descriptor: | CADMIUM ION, METALLOTHIONEIN-1 | | Authors: | Munoz, A, Forsterling, F.H, Shaw III, C.F, Petering, D.H. | | Deposit date: | 2002-05-16 | | Release date: | 2002-05-22 | | Last modified: | 2024-05-08 | | Method: | SOLUTION NMR | | Cite: | Structure of the (113)Cd(3)beta domains from Homarus americanus metallothionein-1: hydrogen bonding and solvent accessibility of sulfur atoms
J.Biol.Inorg.Chem., 7, 2002
|
|
1N1U
 
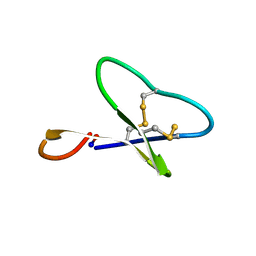 | | NMR structure of [Ala1,15]kalata B1 | | Descriptor: | kalata B1 | | Authors: | Daly, N.L, Clark, R.J, Craik, D.J. | | Deposit date: | 2002-10-20 | | Release date: | 2003-02-25 | | Last modified: | 2022-02-23 | | Method: | SOLUTION NMR | | Cite: | Disulfide Folding Pathways of Cystine Knot Proteins. TYING THE KNOT WITHIN THE CIRCULAR BACKBONE OF THE CYCLOTIDES
J.Biol.Chem., 278, 2003
|
|
6O3Q
 
 | |
6OQ2
 
 | |
1OLD
 
 | |
6AST
 
 | | NMR and Restrained Molecular Dynamics Determination of the Structure of an Aza-Benzimidazole Derivative Complex with the DNA Minor Groove of an -AAGATA Sequence | | Descriptor: | DNA (5'-D(*CP*CP*AP*AP*GP*AP*TP*AP*G)-3'), DNA (5'-D(*CP*TP*AP*TP*CP*TP*TP*GP*G)-3'), amino(4-{[(2-{4-[amino(iminio)methyl]phenyl}-3H-imidazo[4,5-b]pyridin-5-yl)oxy]methyl}phenyl)methaniminium | | Authors: | Harika, N.K, Germann, M.W, Wilson, W.D. | | Deposit date: | 2017-08-25 | | Release date: | 2018-01-31 | | Last modified: | 2024-05-01 | | Method: | SOLUTION NMR | | Cite: | First Structure of a Designed Minor Groove Binding Heterocyclic Cation that Specifically Recognizes Mixed DNA Base Pair Sequences.
Chemistry, 23, 2017
|
|
1Q02
 
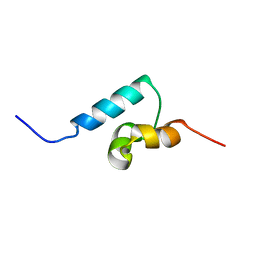 | | NMR structure of the UBA domain of p62 (SQSTM1) | | Descriptor: | sequestosome 1 | | Authors: | Ciani, B, Layfield, R, Cavey, J.R, Sheppard, P.W, Searle, M.S. | | Deposit date: | 2003-07-15 | | Release date: | 2003-09-30 | | Last modified: | 2024-05-22 | | Method: | SOLUTION NMR | | Cite: | Structure of the Ubiquitin-associated Domain of p62 (SQSTM1) and Implications for Mutations That Cause Paget's
Disease of Bone
J.Biol.Chem., 278, 2003
|
|
1NBL
 
 | |
6ASF
 
 | |
6O2L
 
 | |
1QC8
 
 | | NMR STRUCTURE OF TAU EXON 10 SPLICING REGULATORY ELEMENT RNA | | Descriptor: | TAU EXON 10 SPLICING REGULATORY ELEMENT RNA | | Authors: | Varani, L, Spillantini, M.G, Klug, A, Goedert, M, Varani, G. | | Deposit date: | 1999-05-18 | | Release date: | 1999-08-31 | | Last modified: | 2022-03-02 | | Method: | SOLUTION NMR | | Cite: | Structure of tau exon 10 splicing regulatory element RNA and destabilization by mutations of frontotemporal dementia and parkinsonism linked to chromosome 17.
Proc.Natl.Acad.Sci.USA, 96, 1999
|
|
