1MWQ
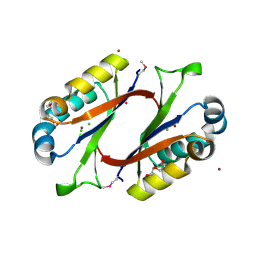 
 | | Structure of HI0828, a Hypothetical Protein from Haemophilus influenzae with a Putative Active-Site Phosphohistidine | | Descriptor: | CACODYLATE ION, CHLORIDE ION, DI(HYDROXYETHYL)ETHER, ... | | Authors: | Willis, M.A, Krajewski, W, Chalamasetty, V.R, Reddy, P, Howard, A, Herzberg, O, Structure 2 Function Project (S2F) | | Deposit date: | 2002-09-30 | | Release date: | 2003-11-25 | | Last modified: | 2019-07-24 | | Method: | X-RAY DIFFRACTION (0.99 Å) | | Cite: | Structure of YciI from Haemophilus influenzae (HI0828) reveals a ferredoxin-like alpha/beta-fold with a histidine/aspartate centered catalytic site
Proteins, 59, 2005
|
|
1SFD
 
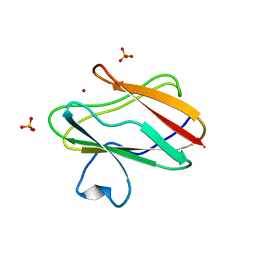 | | oxidized form of amicyanin mutant P94F | | Descriptor: | Amicyanin, COPPER (II) ION, SULFATE ION | | Authors: | Carrell, C.J, Sun, D, Jiang, S, Davidson, V.L, Mathews, F.S. | | Deposit date: | 2004-02-19 | | Release date: | 2004-07-27 | | Last modified: | 2023-08-23 | | Method: | X-RAY DIFFRACTION (0.99 Å) | | Cite: | Structural Studies of Two Mutants of Amicyanin from Paracoccus denitrificans That Stabilize the Reduced State of the Copper.
Biochemistry, 43, 2004
|
|
7P1U
 
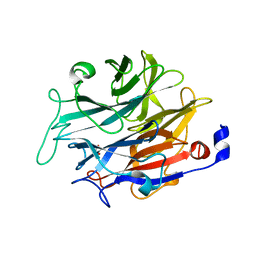 | |
6G1I
 
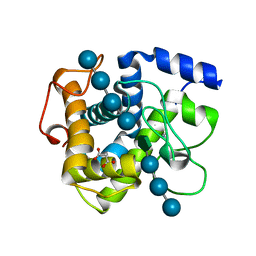 | | GH124 cellulase from Ruminiclostridium thermocellum in complex with Mn and fructosylated cellopentaose | | Descriptor: | Glycosyl Hydrolase, MALONIC ACID, MANGANESE (II) ION, ... | | Authors: | Urresti, S, Davies, G.J, Walton, P.H. | | Deposit date: | 2018-03-21 | | Release date: | 2018-08-15 | | Last modified: | 2024-01-17 | | Method: | X-RAY DIFFRACTION (0.99 Å) | | Cite: | Structural studies of the unusual metal-ion site of the GH124 endoglucanase from Ruminiclostridium thermocellum.
Acta Crystallogr F Struct Biol Commun, 74, 2018
|
|
5O45
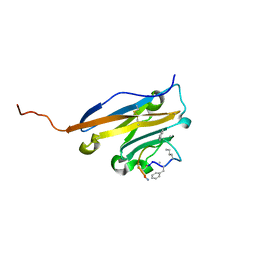 
 | | Structure of human PD-L1 in complex with inhibitor | | Descriptor: | PHE-MEA-9KK-SAR-ASP-VAL-MEA-TYR-SAR-TRP-TYR-LEU-CCS-GLY-NH2, Programmed cell death 1 ligand 1 | | Authors: | Magiera, K, Grudnik, P, Dubin, G, Holak, T.A. | | Deposit date: | 2017-05-26 | | Release date: | 2017-09-20 | | Last modified: | 2024-01-17 | | Method: | X-RAY DIFFRACTION (0.99 Å) | | Cite: | Bioactive Macrocyclic Inhibitors of the PD-1/PD-L1 Immune Checkpoint.
Angew. Chem. Int. Ed. Engl., 56, 2017
|
|
5II8
 
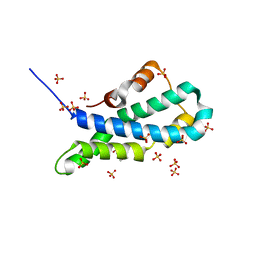 | | Orthorhombic crystal structure of red abalone lysin at 0.99 A resolution | | Descriptor: | 2-(N-MORPHOLINO)-ETHANESULFONIC ACID, Egg-lysin, SULFATE ION | | Authors: | Sadat Al-Hosseini, H, Raj, I, Nishimura, K, De Sanctis, D, Jovine, L. | | Deposit date: | 2016-03-01 | | Release date: | 2017-06-14 | | Last modified: | 2024-01-10 | | Method: | X-RAY DIFFRACTION (0.99 Å) | | Cite: | Structural Basis of Egg Coat-Sperm Recognition at Fertilization.
Cell, 169, 2017
|
|
6YZK
 
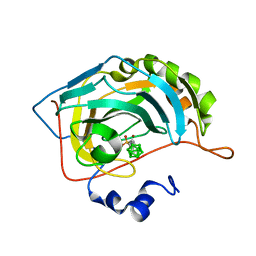 | | Carborane closo-pentyl-sulfonamide in complex with CA IX mimic | | Descriptor: | Carbonic anhydrase 2, Carborane closo-pentyl-sulfonamide, ZINC ION | | Authors: | Kugler, M, Brynda, J, Pospisilova, K, Rezacova, P. | | Deposit date: | 2020-05-07 | | Release date: | 2020-10-28 | | Last modified: | 2024-01-24 | | Method: | X-RAY DIFFRACTION (0.99 Å) | | Cite: | The structural basis for the selectivity of sulfonamido dicarbaboranes toward cancer-associated carbonic anhydrase IX.
J Enzyme Inhib Med Chem, 35, 2020
|
|
7TB7
 
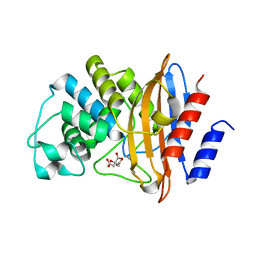 | | Crystal structure of D179N KPC-2 beta-lactamase | | Descriptor: | CITRIC ACID, Carbapenem-hydrolyzing beta-lactamase KPC | | Authors: | van den Akker, F, Alsenani, T. | | Deposit date: | 2021-12-21 | | Release date: | 2022-03-02 | | Last modified: | 2023-10-18 | | Method: | X-RAY DIFFRACTION (0.99 Å) | | Cite: | Structural Characterization of the D179N and D179Y Variants of KPC-2 beta-Lactamase: Omega-Loop Destabilization as a Mechanism of Resistance to Ceftazidime-Avibactam.
Antimicrob.Agents Chemother., 66, 2022
|
|
3DW3
 
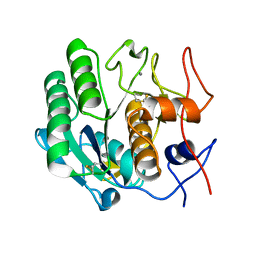 | |
6JJQ
 
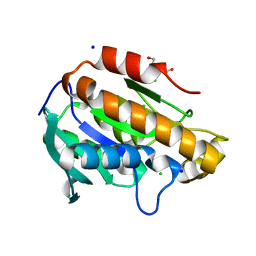 | | Crystal structure of peptidyl-tRNA hydrolase from Acinetobacter baumannii at 0.99 A resolution. | | Descriptor: | CHLORIDE ION, DI(HYDROXYETHYL)ETHER, Peptidyl-tRNA hydrolase, ... | | Authors: | Viswanathan, V, Bairagya, H.R, Sharma, P, Sharma, S, Singh, T.P. | | Deposit date: | 2019-02-26 | | Release date: | 2019-03-06 | | Last modified: | 2023-11-22 | | Method: | X-RAY DIFFRACTION (0.99 Å) | | Cite: | Crystal structure of peptidyl-tRNA hydrolase from Acinetobacter baumannii at 0.99 A resolution.
To Be Published
|
|
7LOG
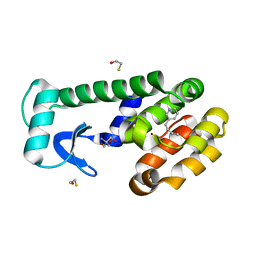 
 | | T4 lysozyme mutant L99A in complex with 3-butylpyridine | | Descriptor: | 2-AMINO-2-HYDROXYMETHYL-PROPANE-1,3-DIOL, 3-butylpyridine, BETA-MERCAPTOETHANOL, ... | | Authors: | Kamenik, A.S, Singh, I, Lak, P, Balius, T.E, Liedl, K.R, Shoichet, B.K. | | Deposit date: | 2021-02-10 | | Release date: | 2021-08-25 | | Last modified: | 2023-10-18 | | Method: | X-RAY DIFFRACTION (0.99 Å) | | Cite: | Energy penalties enhance flexible receptor docking in a model cavity.
Proc.Natl.Acad.Sci.USA, 118, 2021
|
|
8GIV
 
 | |
7G0Y
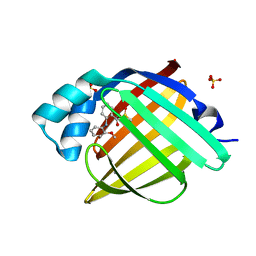 
 | | Crystal Structure of human FABP4 in complex with 2-(2-methoxyanilino)benzoic acid | | Descriptor: | 2-(2-methoxyanilino)benzoic acid, DIMETHYL SULFOXIDE, FORMIC ACID, ... | | Authors: | Ehler, A, Benz, J, Obst, U, Gurien, H, Rudolph, M.G. | | Deposit date: | 2023-04-27 | | Release date: | 2023-06-14 | | Last modified: | 2024-04-03 | | Method: | X-RAY DIFFRACTION (0.99 Å) | | Cite: | Crystal Structure of a human FABP4 complex
To be published
|
|
7FZ1
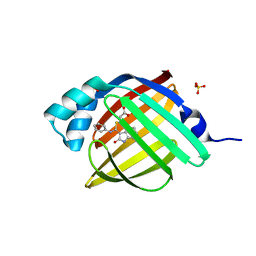 
 | | Crystal Structure of human FABP4 in complex with 3-[(E)-anilino-(2-oxo-1H-indol-3-ylidene)methyl]sulfanylpropanoic acid | | Descriptor: | 3-{[(E)-anilino(2-oxo-1,2-dihydro-3H-indol-3-ylidene)methyl]sulfanyl}propanoic acid, FORMIC ACID, Fatty acid-binding protein, ... | | Authors: | Ehler, A, Benz, J, Obst, U, Mischke, S, Rudolph, M.G. | | Deposit date: | 2023-04-27 | | Release date: | 2023-06-14 | | Last modified: | 2024-04-03 | | Method: | X-RAY DIFFRACTION (0.99 Å) | | Cite: | Crystal Structure of a human FABP4 complex
To be published
|
|
7G08
 
 | | Crystal Structure of human FABP4 in complex with 2-(3-chloro-2-methylanilino)benzoic acid | | Descriptor: | 2-[(3-chloro-2-methylphenyl)amino]benzoic acid, Fatty acid-binding protein, adipocyte, ... | | Authors: | Ehler, A, Benz, J, Obst, U, Boehringer, M, Rudolph, M.G. | | Deposit date: | 2023-04-27 | | Release date: | 2023-06-14 | | Last modified: | 2024-04-03 | | Method: | X-RAY DIFFRACTION (0.99 Å) | | Cite: | Crystal Structure of a human FABP4 complex
To be published
|
|
7FX1
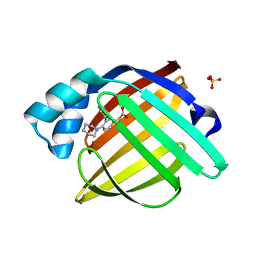 
 | | Crystal Structure of human FABP4 in complex with 2-(3-phenyl-4-piperidin-1-ylphenyl)acetic acid | | Descriptor: | Fatty acid-binding protein, adipocyte, SULFATE ION, ... | | Authors: | Ehler, A, Benz, J, Obst, U, Pfister, R, Rudolph, M.G. | | Deposit date: | 2023-04-27 | | Release date: | 2023-06-14 | | Last modified: | 2024-04-03 | | Method: | X-RAY DIFFRACTION (0.99 Å) | | Cite: | Crystal Structure of a human FABP4 complex
To be published
|
|
7FXR
 
 | | Crystal Structure of human FABP4 in complex with 2-[rac-(9R,10S)-10-benzyl-3,3-dimethyl-1,5-dioxaspiro[5.5]undecan-9-yl]acetic acid | | Descriptor: | FORMIC ACID, Fatty acid-binding protein, adipocyte, ... | | Authors: | Ehler, A, Benz, J, Obst, U, Chen, J, Rudolph, M.G. | | Deposit date: | 2023-04-27 | | Release date: | 2023-06-14 | | Last modified: | 2024-04-03 | | Method: | X-RAY DIFFRACTION (0.99 Å) | | Cite: | Crystal Structure of a human FABP4 complex
To be published
|
|
1P1X
 
 | |
7FZO
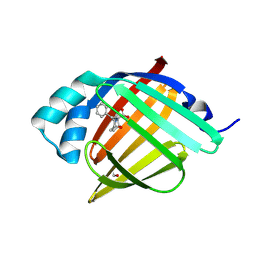 
 | | Crystal Structure of human FABP4 in complex with 4-oxo-3-(2-phenylethyl)-10-oxa-3-azatricyclo[5.2.1.01,5]dec-8-ene-6-carboxylic acid | | Descriptor: | (3aR,6S,7R,7aS)-1-oxo-2-(2-phenylethyl)-1,2,3,6,7,7a-hexahydro-3a,6-epoxyisoindole-7-carboxylic acid, FORMIC ACID, Fatty acid-binding protein, ... | | Authors: | Ehler, A, Benz, J, Obst, U, Brunner, M, Rudolph, M.G. | | Deposit date: | 2023-04-27 | | Release date: | 2023-06-14 | | Last modified: | 2024-04-03 | | Method: | X-RAY DIFFRACTION (0.99 Å) | | Cite: | Crystal Structure of a human FABP4 complex
To be published
|
|
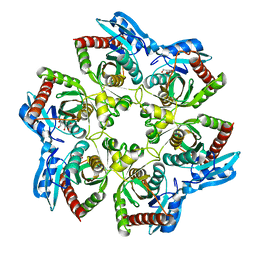
4RJ2
 | |
3DWE
 
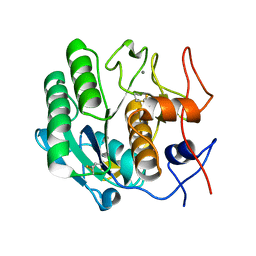 | |
5RCB
 
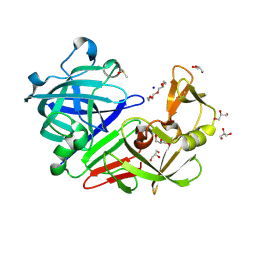 | | PanDDA analysis group deposition -- Endothiapepsin changed state model for fragment F2X-Entry Library G09a | | Descriptor: | (2R)-2-(acetylamino)-4-phenylbutanoic acid, ACETATE ION, DIMETHYL SULFOXIDE, ... | | Authors: | Weiss, M.S, Wollenhaupt, J, Metz, A, Barthel, T, Lima, G.M.A, Heine, A, Mueller, U, Klebe, G. | | Deposit date: | 2020-03-24 | | Release date: | 2020-06-03 | | Last modified: | 2020-06-17 | | Method: | X-RAY DIFFRACTION (0.99 Å) | | Cite: | F2X-Universal and F2X-Entry: Structurally Diverse Compound Libraries for Crystallographic Fragment Screening.
Structure, 28, 2020
|
|
5JEV
 
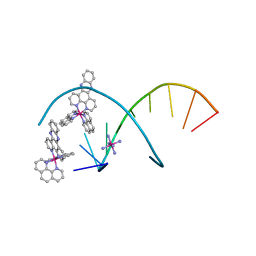 | | del-[Ru(phen)2(dppz]2+ bound to d(TCGGCGCCGA) with Cobalt hexammine | | Descriptor: | COBALT HEXAMMINE(III), DNA (5'-D(*TP*CP*GP*GP*CP*GP*CP*CP*GP*A)-3'), Delta-Ru(phen)2(dppz) complex | | Authors: | Hall, J.P, Cardin, C.J. | | Deposit date: | 2016-04-19 | | Release date: | 2016-09-14 | | Last modified: | 2024-05-08 | | Method: | X-RAY DIFFRACTION (0.99 Å) | | Cite: | Delta chirality ruthenium 'light-switch' complexes can bind in the minor groove of DNA with five different binding modes.
Nucleic Acids Res., 44, 2016
|
|
6RK0
 
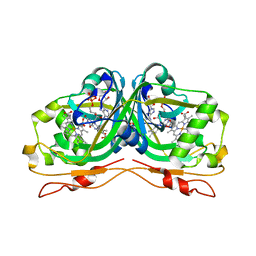 | | Structure of the Flavocytochrome Anf3 from Azotobacter vinelandii | | Descriptor: | FLAVIN-ADENINE DINUCLEOTIDE, PROTOPORPHYRIN IX CONTAINING FE, SULFATE ION, ... | | Authors: | Murray, J.W, Varghese, F, Kabasakal, B. | | Deposit date: | 2019-04-29 | | Release date: | 2019-05-08 | | Last modified: | 2024-01-24 | | Method: | X-RAY DIFFRACTION (0.99 Å) | | Cite: | A low-potential terminal oxidase associated with the iron-only nitrogenase from the nitrogen-fixing bacteriumAzotobacter vinelandii.
J.Biol.Chem., 294, 2019
|
|
6TD0
 
 | | Crystal structure of vaborbactam bound to KPC-2 | | Descriptor: | Carbapenem-hydrolyzing beta-lactamase KPC, GLYCEROL, SULFATE ION, ... | | Authors: | Tooke, C.L, Hinchliffe, P, Spencer, J. | | Deposit date: | 2019-11-07 | | Release date: | 2020-01-22 | | Last modified: | 2024-01-24 | | Method: | X-RAY DIFFRACTION (0.99 Å) | | Cite: | Cyclic boronates as versatile scaffolds for KPC-2 beta-lactamase inhibition.
Rsc Med Chem, 11, 2020
|
|
