1GMX
 
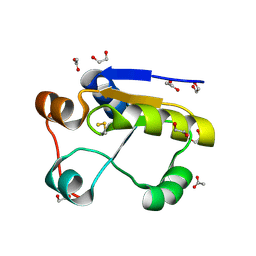 | | Escherichia coli GlpE sulfurtransferase | | Descriptor: | 1,2-ETHANEDIOL, ACETATE ION, THIOSULFATE SULFURTRANSFERASE GLPE | | Authors: | Spallarossa, A, Donahue, J.T, Larson, T.J, Bolognesi, M, Bordo, D. | | Deposit date: | 2001-09-25 | | Release date: | 2001-11-28 | | Last modified: | 2019-07-24 | | Method: | X-RAY DIFFRACTION (1.1 Å) | | Cite: | Escherichia Coli Glpe is a Prototype Sulfurtransferase for the Single-Domain Rhodanese Homology Superfamily
Structure, 9, 2001
|
|
1HZM
 
 | |
2HHG
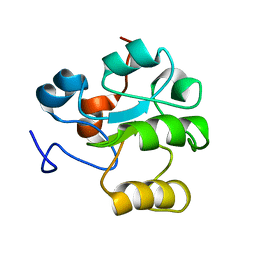
 | | Structure of Protein of Unknown Function RPA3614, Possible Tyrosine Phosphatase, from Rhodopseudomonas palustris CGA009 | | Descriptor: | Hypothetical protein RPA3614, PHOSPHATE ION, SODIUM ION | | Authors: | Binkowski, T.A, Evdokimova, E, Savchenko, A, Edwards, A, Joachimiak, A, Midwest Center for Structural Genomics (MCSG) | | Deposit date: | 2006-06-28 | | Release date: | 2006-07-25 | | Last modified: | 2011-07-13 | | Method: | X-RAY DIFFRACTION (1.2 Å) | | Cite: | Hypothetical protein RPA3614 from Rhodopseudomonas palustris CGA009
To be published
|
|
1YML
 
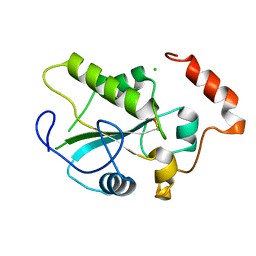 | | Crystal Structure of the CDC25B phosphatase catalytic domain with the active site cysteine in the sulfenic form | | Descriptor: | CHLORIDE ION, M-phase inducer phosphatase 2 | | Authors: | Buhrman, G.K, Parker, B, Sohn, J, Rudolph, J, Mattos, C. | | Deposit date: | 2005-01-21 | | Release date: | 2005-04-12 | | Last modified: | 2023-11-15 | | Method: | X-RAY DIFFRACTION (1.7 Å) | | Cite: | Structural Mechanism of Oxidative Regulation of the Phosphatase Cdc25B via an Intramolecular Disulfide Bond
Biochemistry, 44, 2005
|
|
1YS0
 
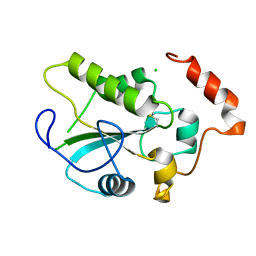 | | Crystal Structure of the CDC25B phosphatase catalytic domain with the active site cysteine in the disulfide form | | Descriptor: | CHLORIDE ION, M-phase inducer phosphatase 2 | | Authors: | Buhrman, G.K, Parker, B, Sohn, J, Rudolph, J, Mattos, C. | | Deposit date: | 2005-02-05 | | Release date: | 2005-04-12 | | Last modified: | 2023-08-23 | | Method: | X-RAY DIFFRACTION (2 Å) | | Cite: | Structural Mechanism of Oxidative Regulation of the Phosphatase Cdc25B via an Intramolecular Disulfide Bond
Biochemistry, 44, 2005
|
|
1YMK
 
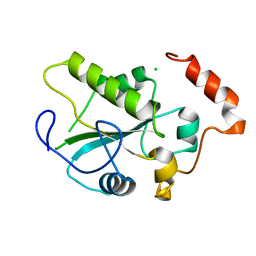 | | Crystal Structure of the CDC25B phosphatase catalytic domain in the apo form | | Descriptor: | CHLORIDE ION, M-phase inducer phosphatase 2 | | Authors: | Buhrman, G.K, Parker, B, Sohn, J, Rudolph, J, Mattos, C. | | Deposit date: | 2005-01-21 | | Release date: | 2005-04-12 | | Last modified: | 2023-08-23 | | Method: | X-RAY DIFFRACTION (1.7 Å) | | Cite: | Structural Mechanism of Oxidative Regulation of the Phosphatase Cdc25B via an Intramolecular Disulfide Bond
Biochemistry, 44, 2005
|
|
1QXN
 
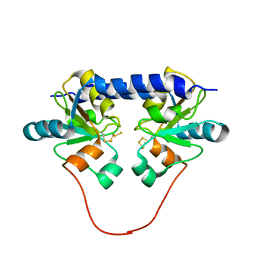 | | Solution Structure of the 30 kDa Polysulfide-sulfur Transferase Homodimer from Wolinella Succinogenes | | Descriptor: | PENTASULFIDE-SULFUR, sulfide dehydrogenase | | Authors: | Lin, Y.J, Dancea, F, Loehr, F, Klimmek, O, Pfeiffer-Marek, S, Nilges, M, Wienk, H, Kroeger, A, Rueterjans, H. | | Deposit date: | 2003-09-08 | | Release date: | 2004-02-24 | | Last modified: | 2022-03-02 | | Method: | SOLUTION NMR | | Cite: | Solution Structure of the 30 kDa Polysulfide-Sulfur Transferase Homodimer from Wolinella succinogenes
Biochemistry, 43, 2004
|
|
1QB0
 
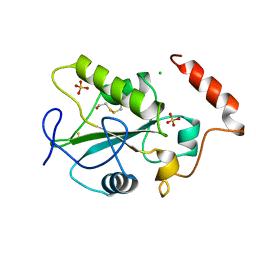 | | HUMAN CDC25B CATALYTIC DOMAIN | | Descriptor: | BETA-MERCAPTOETHANOL, CHLORIDE ION, PROTEIN (M-PHASE INDUCER PHOSPHATASE 2 (CDC25B)), ... | | Authors: | Watenpaugh, K.D, Reynolds, R.A. | | Deposit date: | 1999-04-29 | | Release date: | 2000-04-29 | | Last modified: | 2018-02-28 | | Method: | X-RAY DIFFRACTION (1.91 Å) | | Cite: | Crystal structure of the catalytic subunit of Cdc25B required for G2/M phase transition of the cell cycle.
J.Mol.Biol., 293, 1999
|
|
1YMD
 
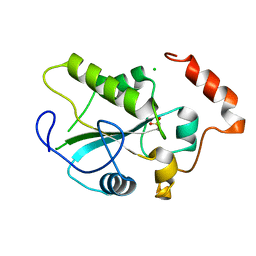 | | Crystal Structure of the CDC25B phosphatase catalytic domain with the active site cysteine in the sulfonic form | | Descriptor: | CHLORIDE ION, M-phase inducer phosphatase 2 | | Authors: | Buhrman, G.K, Parker, B, Sohn, J, Rudolph, J, Mattos, C. | | Deposit date: | 2005-01-20 | | Release date: | 2005-04-12 | | Last modified: | 2023-11-15 | | Method: | X-RAY DIFFRACTION (1.7 Å) | | Cite: | Structural Mechanism of Oxidative Regulation of the Phosphatase Cdc25B via an Intramolecular Disulfide Bond
Biochemistry, 44, 2005
|
|
1YT8
 
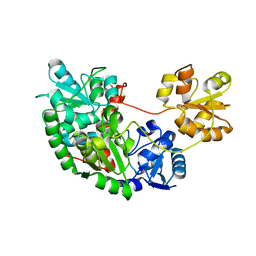 | |
2A2K
 
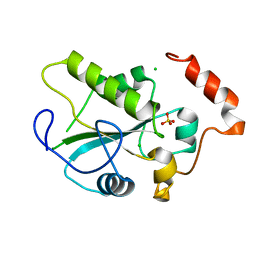 | | Crystal Structure of an active site mutant, C473S, of Cdc25B Phosphatase Catalytic Domain | | Descriptor: | CHLORIDE ION, M-phase inducer phosphatase 2, SULFATE ION | | Authors: | Sohn, J, Parks, J, Buhrman, G, Brown, P, Kristjansdottir, K, Safi, A, Yang, W, Edelsbrunner, H, Rudolph, J. | | Deposit date: | 2005-06-22 | | Release date: | 2006-01-03 | | Last modified: | 2023-08-23 | | Method: | X-RAY DIFFRACTION (1.52 Å) | | Cite: | Experimental Validation of the Docking Orientation of Cdc25 with Its Cdk2-CycA Protein Substrate.
Biochemistry, 44, 2005
|
|
1RHD
 
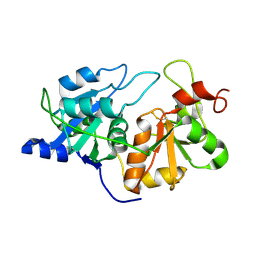 | |
1RHS
 
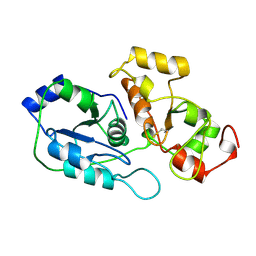 | | SULFUR-SUBSTITUTED RHODANESE | | Descriptor: | SULFUR-SUBSTITUTED RHODANESE | | Authors: | Zanotti, G, Gliubich, F, Colapietro, M, Barba, L. | | Deposit date: | 1997-07-16 | | Release date: | 1998-01-21 | | Last modified: | 2024-06-05 | | Method: | X-RAY DIFFRACTION (1.36 Å) | | Cite: | Structure of sulfur-substituted rhodanese at 1.36 A resolution.
Acta Crystallogr.,Sect.D, 54, 1998
|
|
1ORB
 
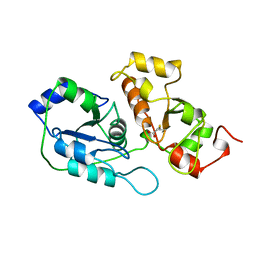 | | ACTIVE SITE STRUCTURAL FEATURES FOR CHEMICALLY MODIFIED FORMS OF RHODANESE | | Descriptor: | ACETATE ION, CARBOXYMETHYLATED RHODANESE | | Authors: | Gliubich, F, Gazerro, M, Zanotti, G, Delbono, S, Berni, R. | | Deposit date: | 1995-07-24 | | Release date: | 1995-10-15 | | Last modified: | 2024-06-05 | | Method: | X-RAY DIFFRACTION (2 Å) | | Cite: | Active site structural features for chemically modified forms of rhodanese.
J.Biol.Chem., 271, 1996
|
|
1T3K
 
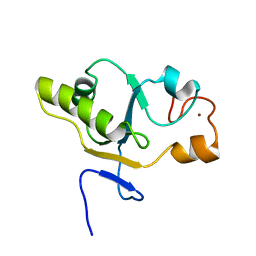 | | NMR structure of a CDC25-like dual-specificity tyrosine phosphatase of Arabidopsis thaliana | | Descriptor: | Dual-specificity tyrosine phosphatase, ZINC ION | | Authors: | Landrieu, I, da Costa, M, De Veylder, L, Dewitte, F, Vandepoele, K, Hassan, S, Wieruszeski, J.M, Faure, J.D, Inze, D, Lippens, G. | | Deposit date: | 2004-04-27 | | Release date: | 2004-09-07 | | Last modified: | 2024-05-22 | | Method: | SOLUTION NMR | | Cite: | A small CDC25 dual-specificity tyrosine-phosphatase isoform in Arabidopsis thaliana.
Proc.Natl.Acad.Sci.Usa, 101, 2004
|
|
3P3A
 | |
3OP3
 
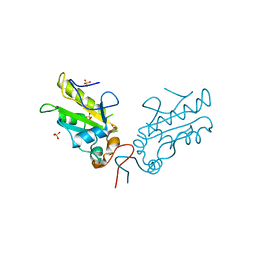 | | Crystal Structure of Cell Division Cycle 25C Protein Isoform A from Homo sapiens | | Descriptor: | M-phase inducer phosphatase 3, SULFATE ION | | Authors: | Kim, Y, Weger, A, Hatzos, C, Savitsky, P, Johansson, C, Ball, L, Barr, A, Vollmar, M, Muniz, J, Weigelt, J, Arrowsmith, C.H, Edwards, A, Bountra, C, Gileadi, O, von Delft, F, Knapp, S, Joachimiak, A, Structural Genomics Consortium (SGC) | | Deposit date: | 2010-08-31 | | Release date: | 2010-09-29 | | Last modified: | 2023-09-06 | | Method: | X-RAY DIFFRACTION (2.63 Å) | | Cite: | Crystal Structure of Cell Division Cycle 25C Protein Isoform A from Homo sapiens
TO BE PUBLISHED
|
|
2MOI
 
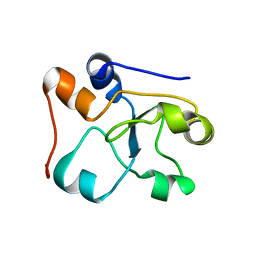 | | 3D NMR structure of the cytoplasmic rhodanese domain of the inner membrane protein YgaP from Escherichia coli | | Descriptor: | Inner membrane protein YgaP | | Authors: | Eichmann, C, Tzitzilonis, C, Bordignon, E, Maslennikov, I, Choe, S, Riek, R. | | Deposit date: | 2014-04-26 | | Release date: | 2014-06-25 | | Last modified: | 2024-05-01 | | Method: | SOLUTION NMR | | Cite: | Solution NMR Structure and Functional Analysis of the Integral Membrane Protein YgaP from Escherichia coli.
J.Biol.Chem., 289, 2014
|
|
2MOL
 
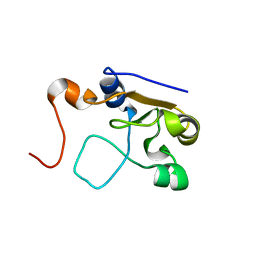 | | 3D NMR structure of the cytoplasmic rhodanese domain of the full-length inner membrane protein YgaP from Escherichia coli | | Descriptor: | Inner membrane protein YgaP | | Authors: | Eichmann, C, Tzitzilonis, C, Bordignon, E, Maslennikov, I, Choe, S, Riek, R. | | Deposit date: | 2014-04-27 | | Release date: | 2014-06-25 | | Last modified: | 2024-05-15 | | Method: | SOLUTION NMR | | Cite: | Solution NMR Structure and Functional Analysis of the Integral Membrane Protein YgaP from Escherichia coli.
J.Biol.Chem., 289, 2014
|
|
2JTR
 
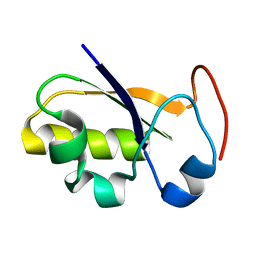 | | rhodanese persulfide from E. coli | | Descriptor: | Phage shock protein E | | Authors: | Jin, C, Li, H. | | Deposit date: | 2007-08-06 | | Release date: | 2008-06-17 | | Last modified: | 2024-05-29 | | Method: | SOLUTION NMR | | Cite: | Solution structures and backbone dynamics of Escherichia coli rhodanese PspE in its sulfur-free and persulfide-intermediate forms: implications for the catalytic mechanism of rhodanese.
Biochemistry, 47, 2008
|
|
3O3W
 
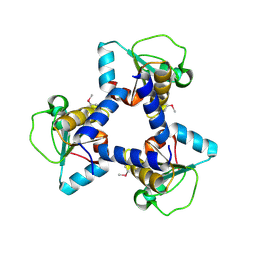 | | Crystal Structure of BH2092 protein (residues 14-131) from Bacillus halodurans, Northeast Structural Genomics Consortium Target BhR228A | | Descriptor: | BH2092 protein | | Authors: | Forouhar, F, Neely, H, Seetharaman, J, Sahdev, S, Xiao, R, Ciccosanti, C, Lee, D, Everett, J.K, Nair, R, Acton, T.B, Rost, B, Montelione, G.T, Tong, L, Hunt, J.F, Northeast Structural Genomics Consortium (NESG) | | Deposit date: | 2010-07-26 | | Release date: | 2010-09-01 | | Last modified: | 2019-07-17 | | Method: | X-RAY DIFFRACTION (2.91 Å) | | Cite: | Crystal Structure of BH2092 protein (residues 14-131) from Bacillus halodurans, Northeast Structural Genomics Consortium Target BhR228A
To be Published
|
|
2JTQ
 
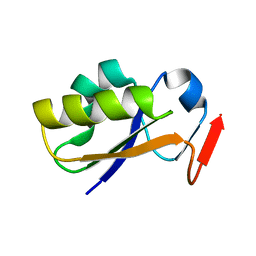 | | Rhodanese from E.coli | | Descriptor: | Phage shock protein E | | Authors: | Jin, C, Li, H. | | Deposit date: | 2007-08-06 | | Release date: | 2008-06-17 | | Last modified: | 2024-05-29 | | Method: | SOLUTION NMR | | Cite: | Solution structures and backbone dynamics of Escherichia coli rhodanese PspE in its sulfur-free and persulfide-intermediate forms: implications for the catalytic mechanism of rhodanese.
Biochemistry, 47, 2008
|
|
3OLH
 
 | | Human 3-mercaptopyruvate sulfurtransferase | | Descriptor: | 3-mercaptopyruvate sulfurtransferase, SODIUM ION, SULFATE ION | | Authors: | Karlberg, T, Collins, R, Arrowsmith, C.H, Berglund, H, Bountra, C, Edwards, A.M, Flodin, S, Flores, A, Graslund, S, Hammarstrom, M, Johansson, I, Kotenyova, T, Kouznetsova, E, Moche, M, Nordlund, P, Nyman, T, Persson, C, Schutz, P, Sehic, A, Siponen, M.I, Thorsell, A.G, Tresaugues, L, Van Den Berg, S, Wahlberg, E, Weigelt, J, Welin, M, Schuler, H, Structural Genomics Consortium (SGC) | | Deposit date: | 2010-08-26 | | Release date: | 2010-09-29 | | Last modified: | 2023-09-06 | | Method: | X-RAY DIFFRACTION (2.5 Å) | | Cite: | Human 3-Mercaptopyruvate Sulfurtransferase
To be Published
|
|
2JTS
 
 | | rhodanese with anions from E. coli | | Descriptor: | Phage shock protein E | | Authors: | Jin, C, Li, H. | | Deposit date: | 2007-08-06 | | Release date: | 2008-06-17 | | Last modified: | 2024-05-29 | | Method: | SOLUTION NMR | | Cite: | Solution structures and backbone dynamics of Escherichia coli rhodanese PspE in its sulfur-free and persulfide-intermediate forms: implications for the catalytic mechanism of rhodanese.
Biochemistry, 47, 2008
|
|
2K0Z
 
 | | Solution NMR structure of protein hp1203 from Helicobacter pylori 26695. Northeast Structural Genomics Consortium (NESG) target PT1/Ontario Center for Structural Proteomics target hp1203 | | Descriptor: | Uncharacterized protein hp1203 | | Authors: | Wu, B, Yee, A, Lemak, A, Cort, J, Semest, A, Kenney, M.A, Arrowsmith, C.H, Northeast Structural Genomics Consortium (NESG) | | Deposit date: | 2008-02-18 | | Release date: | 2008-03-04 | | Last modified: | 2024-05-01 | | Method: | SOLUTION NMR | | Cite: | Solution NMR structure of protein hp1203 from Helicobacter pylori 26695. Northeast Structural Genomics Consortium (NESG) target PT1/Ontario Center for Structural Proteomics target hp1203.
To be Published
|
|
