4E19
 
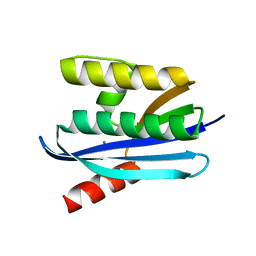 | |
7XX5
 
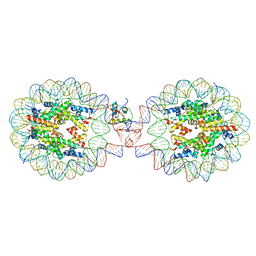 | | Crystal Structure of Nucleosome-H1.3 Linker Histone Assembly (sticky-169a DNA fragment) | | Descriptor: | CALCIUM ION, DNA (169-MER), Histone H1.3, ... | | Authors: | Adhireksan, Z, Qiuye, B, Lee, P.L, Sharma, D, Padavattan, S, Davey, C.A. | | Deposit date: | 2022-05-28 | | Release date: | 2023-05-31 | | Last modified: | 2023-11-29 | | Method: | X-RAY DIFFRACTION (3.19 Å) | | Cite: | Crystal Structure of Nucleosome-H1.0 Linker Histone Assembly (sticky-169a DNA fragment)
To Be Published
|
|
6HKT
 
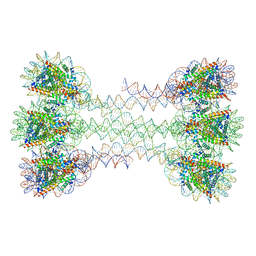 | | Structure of an H1-bound 6-nucleosome array | | Descriptor: | DNA (1122-MER), Histone H2A type 1-B/E, Histone H2B type 1-J, ... | | Authors: | Garcia-Saez, I, Dimitrov, S, Petosa, C. | | Deposit date: | 2018-09-08 | | Release date: | 2018-10-31 | | Last modified: | 2024-01-17 | | Method: | X-RAY DIFFRACTION (9.7 Å) | | Cite: | Structure of an H1-Bound 6-Nucleosome Array Reveals an Untwisted Two-Start Chromatin Fiber Conformation.
Mol. Cell, 72, 2018
|
|
1GHC
 
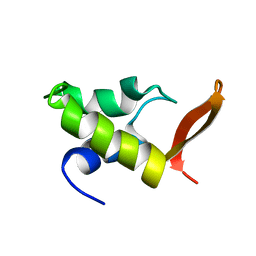 | | HOMO-AND HETERONUCLEAR TWO-DIMENSIONAL NMR STUDIES OF THE GLOBULAR DOMAIN OF HISTONE H1: FULL ASSIGNMENT, TERTIARY STRUCTURE, AND COMPARISON WITH THE GLOBULAR DOMAIN OF HISTONE H5 | | Descriptor: | GH1 | | Authors: | Cerf, C, Lippens, G, Ramakrishnan, V, Muyldermans, S, Segers, A, Wyns, L, Wodak, S.J, Hallenga, K. | | Deposit date: | 1994-05-16 | | Release date: | 1994-08-31 | | Last modified: | 2024-05-22 | | Method: | SOLUTION NMR | | Cite: | Homo- and heteronuclear two-dimensional NMR studies of the globular domain of histone H1: full assignment, tertiary structure, and comparison with the globular domain of histone H5.
Biochemistry, 33, 1994
|
|
6Q6V
 
 | |
6Q47
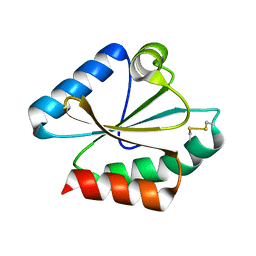 
 | |
6Q6T
 
 | |
6Q46
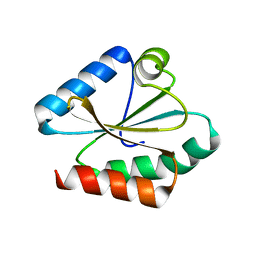 
 | |
6Q6U
 
 | |
4J4Z
 
 | | Crystal structure of the improved variant of the evolved serine hydrolase, OSH55.4_H1.2, bond with sulfate ion in the active site, Northeast Structural Genomics Consortium (NESG) Target OR301 | | Descriptor: | 4-(2-HYDROXYETHYL)-1-PIPERAZINE ETHANESULFONIC ACID, DI(HYDROXYETHYL)ETHER, Designed serine hydrolase variant OSH55.4_H1.2, ... | | Authors: | Kuzin, A.P, Lew, S, Rajagopalan, S, Maglaqui, M, Xiao, R, Lee, D, Everett, J.K, Acton, T.B, Baker, D, Montelione, G.T, Tong, L, Hunt, J.F, Northeast Structural Genomics Consortium (NESG) | | Deposit date: | 2013-02-07 | | Release date: | 2013-03-06 | | Last modified: | 2024-10-09 | | Method: | X-RAY DIFFRACTION (1.26 Å) | | Cite: | Crystal structure of the improved variant of the evolved serine hydrolase, OSH55.4_H1.2, bond with sulfate ion in the active site, Northeast Structural Genomics Consortium (NESG) Target OR301
To be Published
|
|
1TI8
 
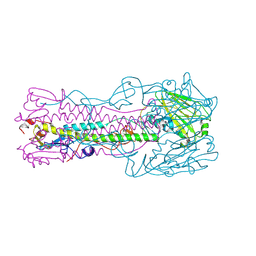 | | H7 Haemagglutinin | | Descriptor: | 2-acetamido-2-deoxy-alpha-D-glucopyranose, 2-acetamido-2-deoxy-beta-D-glucopyranose, alpha-D-mannopyranose, ... | | Authors: | Russell, R.J, Gamblin, S.J, Haire, L.F, Stevens, D.J, Xaio, B, Ha, Y, Skehel, J.J. | | Deposit date: | 2004-06-02 | | Release date: | 2005-06-21 | | Last modified: | 2020-07-29 | | Method: | X-RAY DIFFRACTION (3 Å) | | Cite: | H1 and H7 influenza haemagglutinin structures extend a structural classification of haemagglutinin subtypes.
Virology, 325, 2004
|
|
1YQA
 
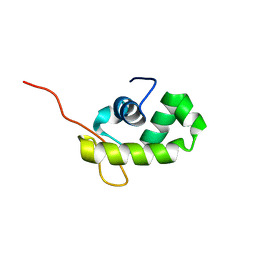 | |
8GZ4
 
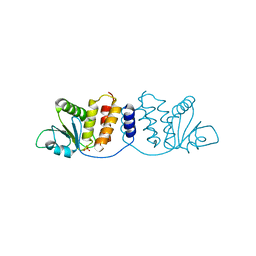 | | Crystal structure of MPXV phosphatase | | Descriptor: | Dual specificity protein phosphatase H1, PHOSPHATE ION | | Authors: | Yang, H.T, Wang, W, Huang, H.J, Ji, X.Y. | | Deposit date: | 2022-09-25 | | Release date: | 2023-05-17 | | Last modified: | 2023-12-06 | | Method: | X-RAY DIFFRACTION (1.802 Å) | | Cite: | Crystal structure of monkeypox H1 phosphatase, an antiviral drug target.
Protein Cell, 14, 2023
|
|
6N89
 
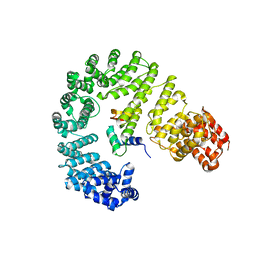 | |
6N88
 
 | |
8D21
 
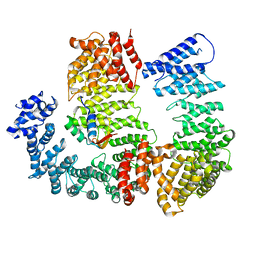
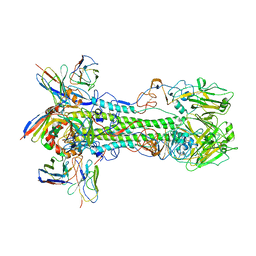 | | Cryo-EM structure of the VRC321 clinical trial, vaccine-elicited, human antibody 1B06 in complex with a stabilized NC99 HA trimer | | Descriptor: | 1B06 Heavy Chain, 1B06 Light Chain, 2-acetamido-2-deoxy-beta-D-glucopyranose, ... | | Authors: | Gorman, J, Kwong, P.D. | | Deposit date: | 2022-05-27 | | Release date: | 2023-04-12 | | Last modified: | 2023-05-03 | | Method: | ELECTRON MICROSCOPY (3.96 Å) | | Cite: | An influenza H1 hemagglutinin stem-only immunogen elicits a broadly cross-reactive B cell response in humans.
Sci Transl Med, 15, 2023
|
|
6H8P
 
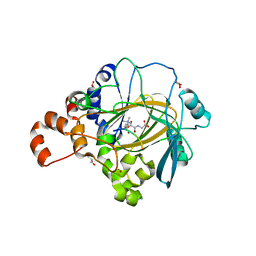 | | JMJD2A/ KDM4A COMPLEXED WITH NI(II), NOG AND Histone H1.4(18-32)K26me3 peptide (15-mer) | | Descriptor: | CHLORIDE ION, GLYCEROL, Histone H1.4, ... | | Authors: | Chowdhury, R, Walport, L.J, Schofield, C.J. | | Deposit date: | 2018-08-03 | | Release date: | 2018-08-15 | | Last modified: | 2024-01-17 | | Method: | X-RAY DIFFRACTION (1.983 Å) | | Cite: | Mechanistic and structural studies of KDM-catalysed demethylation of histone 1 isotype 4 at lysine 26.
FEBS Lett., 592, 2018
|
|
2LB4
 
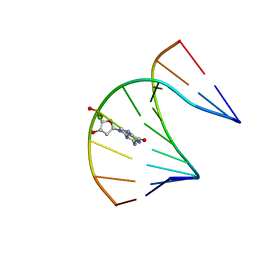 | | DNA / RNA Hybrid containing a central stereo specific Sp borano phosphate linkage | | Descriptor: | DNA_(5'-D(*AP*TP*GP*GP*TP*BGR*CP*TP*C)-3')_, RNA_(5'-R(*GP*AP*GP*CP*AP*CP*CP*AP*U)-3')_ | | Authors: | Johnson, C.N, Spring, A.M, Shaw, B.R, Germann, M.W. | | Deposit date: | 2011-03-22 | | Release date: | 2011-06-29 | | Last modified: | 2024-05-01 | | Method: | SOLUTION NMR | | Cite: | Structural Basis of the RNase H1 Activity on Stereo Regular Borano Phosphonate DNA/RNA Hybrids.
Biochemistry, 50, 2011
|
|
2LAR
 
 | | DNA / RNA Hybrid containing a central stereo specific Rp borano phosphate linkage | | Descriptor: | DNA_(5'-D(*AP*TP*GP*GP*TP*BGR*CP*TP*C)-3')_, RNA_(5'-R(*GP*AP*GP*CP*AP*CP*CP*AP*U)-3')_ | | Authors: | Johnson, C.N, Spring, A.M, Shaw, B.R, Germann, M.W. | | Deposit date: | 2011-03-18 | | Release date: | 2011-10-12 | | Last modified: | 2024-05-01 | | Method: | SOLUTION NMR | | Cite: | Structural basis of the RNase H1 activity on stereo regular borano phosphonate DNA/RNA hybrids.
Biochemistry, 50, 2011
|
|
3L7U
 
 | | Crystal structure of human NM23-H1 | | Descriptor: | Nucleoside diphosphate kinase A, PHOSPHATE ION | | Authors: | Han, B.G, Min, K, Lee, B.I, Lee, S. | | Deposit date: | 2009-12-29 | | Release date: | 2010-11-10 | | Last modified: | 2023-11-01 | | Method: | X-RAY DIFFRACTION (2.1 Å) | | Cite: | Refined Structure of Human NM23-H1 from a Hexagonal Crystal
BULL.KOREAN CHEM.SOC., 31, 2010
|
|
7K5X
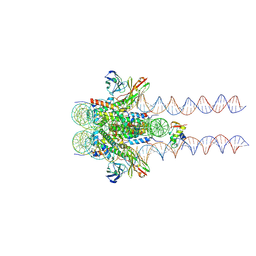 
 | |
7K5Y
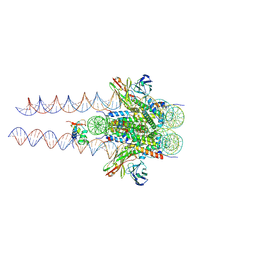 
 | |
7K60
 
 | |
3H08
 
 | | Crystal structure of the Ribonuclease H1 from Chlorobium tepidum | | Descriptor: | MAGNESIUM ION, Rnh (Ribonuclease H) | | Authors: | Ratcliff, K, Corn, J, Marqusee, S. | | Deposit date: | 2009-04-08 | | Release date: | 2009-05-12 | | Last modified: | 2024-02-21 | | Method: | X-RAY DIFFRACTION (1.6 Å) | | Cite: | Structure, stability, and folding of ribonuclease H1 from the moderately thermophilic chlorobium tepidum: comparison with thermophilic and mesophilic homologues.
Biochemistry, 48, 2009
|
|
6QZL
 
 | | Structure of the H1 domain of human KCTD12 | | Descriptor: | BTB/POZ domain-containing protein KCTD12 | | Authors: | Pinkas, D.M, Bufton, J.C, Fox, A.E, Newman, J.A, Kupinska, K, Burgess-Brown, N.A, von Delft, F, Arrowsmith, C.H, Edwards, A.M, Bountra, C, Bullock, A.N, Structural Genomics Consortium (SGC) | | Deposit date: | 2019-03-11 | | Release date: | 2019-03-20 | | Last modified: | 2024-01-24 | | Method: | X-RAY DIFFRACTION (1.98 Å) | | Cite: | Structure of the H1 domain of human KCTD12
To be published
|
|
