2G71
 
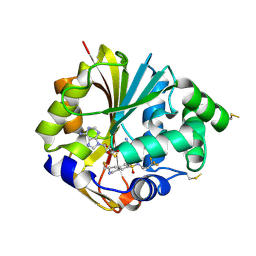 | | Structure of hPNMT with inhibitor 3-fluoromethyl-7-trifluoropropyl-THIQ and AdoHcy | | Descriptor: | (3R)-3-(FLUOROMETHYL)-N-(3,3,3-TRIFLUOROPROPYL)-1,2,3,4-TETRAHYDROISOQUINOLINE-7-SULFONAMIDE, GLYCEROL, Phenylethanolamine N-methyltransferase, ... | | Authors: | Tyndall, J.D.A, Gee, C.L, Martin, J.L. | | Deposit date: | 2006-02-27 | | Release date: | 2007-02-13 | | Last modified: | 2023-10-25 | | Method: | X-RAY DIFFRACTION (2.2 Å) | | Cite: | Enzyme Adaptation to Inhibitor Binding: A Cryptic Binding Site in Phenylethanolamine N-Methyltransferase
J.Med.Chem., 50, 2007
|
|
2G72
 
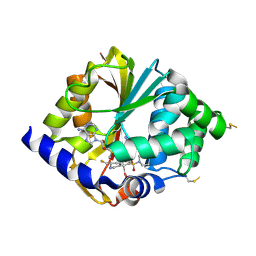 | | Structure of hPNMT with inhibitor 3-fluoromethyl-7-thiomorpholinosulfonamide-THIQ and AdoMet | | Descriptor: | (3R)-3-(FLUOROMETHYL)-7-(THIOMORPHOLIN-4-YLSULFONYL)-1,2,3,4-TETRAHYDROISOQUINOLINE, Phenylethanolamine N-methyltransferase, S-ADENOSYLMETHIONINE | | Authors: | Tyndall, J.D.A, Gee, C.L, Martin, J.L. | | Deposit date: | 2006-02-27 | | Release date: | 2007-02-13 | | Last modified: | 2023-10-25 | | Method: | X-RAY DIFFRACTION (2 Å) | | Cite: | Enzyme Adaptation to Inhibitor Binding: A Cryptic Binding Site in Phenylethanolamine N-Methyltransferase
J.Med.Chem., 50, 2007
|
|
2G73
 
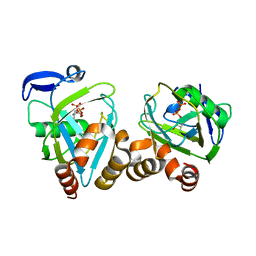 | | Y104F mutant type 1 IPP isomerase complex with EIPP | | Descriptor: | 4-HYDROXY-3-METHYL BUTYL DIPHOSPHATE, Isopentenyl-diphosphate delta-isomerase, MAGNESIUM ION, ... | | Authors: | de Ruyck, J, Wouters, J. | | Deposit date: | 2006-02-27 | | Release date: | 2006-04-18 | | Last modified: | 2023-10-25 | | Method: | X-RAY DIFFRACTION (1.97 Å) | | Cite: | Structural role for Tyr-104 in Escherichia coli isopentenyl-diphosphate isomerase: site-directed mutagenesis, enzymology, and protein crystallography.
J.Biol.Chem., 281, 2006
|
|
2G74
 
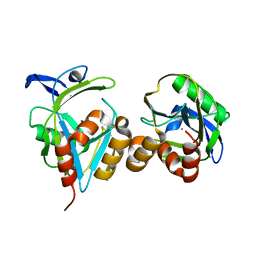 | |
2G75
 
 | | Crystal Structure of anti-SARS m396 Antibody | | Descriptor: | IGG Heavy Chain, IGG Light Chain | | Authors: | Prabakaran, P, Gan, J.H, Feng, Y, Zhu, Z.Y, Xiao, X.D, Ji, X, Dimitrov, D.S. | | Deposit date: | 2006-02-27 | | Release date: | 2006-04-04 | | Last modified: | 2023-08-30 | | Method: | X-RAY DIFFRACTION (2.28 Å) | | Cite: | Structure of severe acute respiratory syndrome coronavirus receptor-binding domain complexed with neutralizing antibody.
J.Biol.Chem., 281, 2006
|
|
2G76
 
 | | Crystal structure of human 3-phosphoglycerate dehydrogenase | | Descriptor: | D-3-phosphoglycerate dehydrogenase, D-MALATE, NICOTINAMIDE-ADENINE-DINUCLEOTIDE | | Authors: | Turnbull, A.P, Salah, E, Savitsky, P, Gileadi, O, von Delft, F, Edwards, A, Arrowsmith, C, Weigelt, J, Sundstrom, M, Oppermann, U, Structural Genomics Consortium (SGC) | | Deposit date: | 2006-02-27 | | Release date: | 2006-03-21 | | Last modified: | 2023-08-30 | | Method: | X-RAY DIFFRACTION (1.7 Å) | | Cite: | Crystal structure of human 3-phosphoglycerate dehydrogenase
To be Published
|
|
2G77
 
 | | Crystal Structure of Gyp1 TBC domain in complex with Rab33 GTPase bound to GDP and AlF3 | | Descriptor: | ALUMINUM FLUORIDE, GTPase-activating protein GYP1, GUANOSINE-5'-DIPHOSPHATE, ... | | Authors: | Pan, X, Eathiraj, S, Munson, M, Lambright, D.G. | | Deposit date: | 2006-02-27 | | Release date: | 2006-07-25 | | Last modified: | 2021-10-20 | | Method: | X-RAY DIFFRACTION (2.26 Å) | | Cite: | TBC-domain GAPs for Rab GTPases accelerate GTP hydrolysis by a dual-finger mechanism.
Nature, 442, 2006
|
|
2G78
 | |
2G79
 | |
2G7B
 | |
2G7C
 
 | | Clostridium difficile Toxin A Fragment Bound to aGal(1,3)bGal(1,4)bGlcNAc | | Descriptor: | GLYCEROL, Toxin A, alpha-D-galactopyranose-(1-3)-beta-D-galactopyranose-(1-4)-2-acetamido-2-deoxy-beta-D-glucopyranose | | Authors: | Greco, A, Ho, J.G.S, Lin, S.J, Palcic, M.M, Rupnik, M, Ng, K.K.S. | | Deposit date: | 2006-02-28 | | Release date: | 2006-04-18 | | Last modified: | 2023-09-20 | | Method: | X-RAY DIFFRACTION (2 Å) | | Cite: | Carbohydrate recognition by Clostridium difficile toxin A.
Nat.Struct.Mol.Biol., 13, 2006
|
|
2G7E
 
 | | The 1.6 A crystal structure of Vibrio cholerae extracellular endonuclease I | | Descriptor: | CHLORIDE ION, Endonuclease I | | Authors: | Altermark, B, Smalaas, A.O, Willassen, N.P, Helland, R. | | Deposit date: | 2006-02-28 | | Release date: | 2006-10-31 | | Last modified: | 2023-08-30 | | Method: | X-RAY DIFFRACTION (1.6 Å) | | Cite: | The structure of Vibrio cholerae extracellular endonuclease I reveals the presence of a buried chloride ion.
Acta Crystallogr.,Sect.D, 62, 2006
|
|
2G7F
 
 | | The 1.95 A crystal structure of Vibrio cholerae extracellular endonuclease I | | Descriptor: | CHLORIDE ION, Endonuclease I, MAGNESIUM ION | | Authors: | Altermark, B, Smalaas, A.O, Willassen, N.P, Helland, R. | | Deposit date: | 2006-02-28 | | Release date: | 2006-10-31 | | Last modified: | 2023-08-30 | | Method: | X-RAY DIFFRACTION (1.95 Å) | | Cite: | The structure of Vibrio cholerae extracellular endonuclease I reveals the presence of a buried chloride ion.
Acta Crystallogr.,Sect.D, 62, 2006
|
|
2G7G
 
 | | The Crystal Structure of the Putative Transcriptional Regulator Rha04620 from Rhodococcus sp. RHA1 | | Descriptor: | ACETIC ACID, Rha04620, Putative Transcriptional Regulator | | Authors: | Kim, Y, Joachimiak, A, Evdokimova, E, Kagan, O, Savchenko, A, Edwards, A.M, Midwest Center for Structural Genomics (MCSG) | | Deposit date: | 2006-02-28 | | Release date: | 2006-03-28 | | Last modified: | 2011-07-13 | | Method: | X-RAY DIFFRACTION (2.01 Å) | | Cite: | The Crystal Structure of the Putative Transcriptional Regulator Rha04620 from Rhodococcus sp. RHA1
To be Published
|
|
2G7H
 
 | |
2G7I
 
 | |
2G7J
 
 | | Solution NMR structure of the putative cytoplasmic protein ygaC from Salmonella typhimurium. Northeast Structural Genomics target StR72. | | Descriptor: | putative cytoplasmic protein | | Authors: | Aramini, J.M, Swapna, G.V.T, Ramelot, T.A, Ho, C.K, Cunningham, K, Ma, L.-C, Shetty, K, Xiao, R, Acton, T.B, Kennedy, M.A, Montelione, G.T, Northeast Structural Genomics Consortium (NESG) | | Deposit date: | 2006-02-28 | | Release date: | 2006-05-02 | | Last modified: | 2024-05-01 | | Method: | SOLUTION NMR | | Cite: | Solution NMR structure of the putative cytoplasmic protein ygaC from Salmonella typhimurium. Northeast Structural Genomics target StR72.
To be Published
|
|
2G7K
 
 | | Structure of the Light Chain of Botulinum Neurotoxin, Serotype A Bound to small Molecule Inhibitors | | Descriptor: | Botulinum neurotoxin type A | | Authors: | Fu, Z, Baldwin, M.R, Boldt, G.E, Crawford, A, Janda, K.D, Barbieri, J.T, Kim, J.-J.P. | | Deposit date: | 2006-02-28 | | Release date: | 2006-08-15 | | Last modified: | 2023-08-30 | | Method: | X-RAY DIFFRACTION (2.8 Å) | | Cite: | Light chain of botulinum neurotoxin serotype A: structural resolution of a catalytic intermediate.
Biochemistry, 45, 2006
|
|
2G7L
 
 | | Crystal structure of putative transcription regulator SCO7704 from Streptomyces coelicor | | Descriptor: | TetR-family transcriptional regulator | | Authors: | Ezersky, A, Lunin, V.V, Skarina, T, Wierzbicka, M, Joachimiak, A, Edwards, A.M, Savchenko, A, Midwest Center for Structural Genomics (MCSG) | | Deposit date: | 2006-02-28 | | Release date: | 2006-03-14 | | Last modified: | 2017-10-18 | | Method: | X-RAY DIFFRACTION (2.1 Å) | | Cite: | Crystal structure of putative transcription regulator SCO7704 from Streptomyces coelicor
To be Published
|
|
2G7M
 | | Crystal structure of B. fragilis N-succinylornithine transcarbamylase P90E mutant complexed with carbamoyl phosphate and N-acetylnorvaline | | Descriptor: | N-ACETYL-L-NORVALINE, PHOSPHORIC ACID MONO(FORMAMIDE)ESTER, SULFATE ION, ... | | Authors: | Shi, D, Yu, X, Roth, L, Morizono, H, Allewell, N.M, Tuchman, M. | | Deposit date: | 2006-02-28 | | Release date: | 2007-03-06 | | Last modified: | 2023-08-30 | | Method: | X-RAY DIFFRACTION (2.9 Å) | | Cite: | A single mutation in the active site swaps the substrate specificity of N-acetyl-L-ornithine transcarbamylase and N-succinyl-L-ornithine transcarbamylase.
Protein Sci., 16, 2007
|
|
2G7N
 
 | | Structure of the Light Chain of Botulinum Neurotoxin Serotype A Bound to small Molecule Inhibitors | | Descriptor: | Botulinum neurotoxin type A, SILVER ION, ZINC ION | | Authors: | Fu, Z, Baldwin, M.R, Boldt, G.E, Janda, K.D, Barbieri, J.T, Kim, J.-J.P. | | Deposit date: | 2006-02-28 | | Release date: | 2007-03-13 | | Last modified: | 2023-08-30 | | Method: | X-RAY DIFFRACTION (1.9 Å) | | Cite: | Structure of the Light Chain of Botulinum Neurotoxin Serotype A bound to Small Molecule Inhibitors
To be Published
|
|
2G7O
 
 | | Protonation-mediated structural flexibility in the F conjugation regulatory protein, TraM | | Descriptor: | Protein traM | | Authors: | Lu, J, Edwards, R.A, Wong, J.J, Manchak, J, Scott, P.G, Frost, L.S, Glover, J.N. | | Deposit date: | 2006-02-28 | | Release date: | 2006-06-13 | | Last modified: | 2024-02-14 | | Method: | X-RAY DIFFRACTION (1.4 Å) | | Cite: | Protonation-mediated structural flexibility in the F conjugation regulatory protein, TraM.
Embo J., 25, 2006
|
|
2G7P
 
 | | Structure of the Light Chain of Botulinum Neurotoxin Serotype A Bound to Small Molecule Inhibitors | | Descriptor: | Botulinum neurotoxin type A, ZINC ION | | Authors: | Fu, Z, Baldwin, M.R, Boldt, G.E, Janda, K.D, Barbieri, J.T, Kim, J.-J.P. | | Deposit date: | 2006-02-28 | | Release date: | 2006-08-15 | | Last modified: | 2023-08-30 | | Method: | X-RAY DIFFRACTION (2.3 Å) | | Cite: | Light chain of botulinum neurotoxin serotype A: structural resolution of a catalytic intermediate.
Biochemistry, 45, 2006
|
|
2G7Q
 
 | | Structure of the Light Chain of Botulinum Neurotoxin Serotype A Bound to Small Molecule Inhibitors | | Descriptor: | Botulinum neurotoxin type A, N-HYDROXY-L-ARGININAMIDE, ZINC ION | | Authors: | Fu, Z, Baldwin, M.R, Boldt, G.E, Janda, K.D, Barbieri, J.T, Kim, J.-J.P. | | Deposit date: | 2006-02-28 | | Release date: | 2006-08-15 | | Last modified: | 2023-08-30 | | Method: | X-RAY DIFFRACTION (2.41 Å) | | Cite: | Light chain of botulinum neurotoxin serotype A: structural resolution of a catalytic intermediate.
Biochemistry, 45, 2006
|
|
2G7R
 
 | | X-ray structure of the death domain of the human mucosa associated lymphoid tissue lymphoma translocation protein 1 | | Descriptor: | Mucosa-associated lymphoid tissue lymphoma translocation protein 1 | | Authors: | Walker, J.R, Wybenga-Groot, L, Newman, E.M, Finerty Jr, P.J, Butler-Cole, C, Weigelt, J, Sundstrom, M, Arrowsmith, C, Edwards, A, Bochkarev, A, Dhe-Paganon, S, Structural Genomics Consortium (SGC) | | Deposit date: | 2006-02-28 | | Release date: | 2006-04-04 | | Last modified: | 2011-07-13 | | Method: | X-RAY DIFFRACTION (2.7 Å) | | Cite: | X-ray structure of the death domain of the human mucosa associated lymphoid tissue lymphoma translocation protein 1
To be Published
|
|
