2RLG
 
 | |
2RLH
 | |
1GA3
 
 | |
5ZB6
 | |
6ALK
 | | NMR solution structure of the major beech pollen allergen Fag s 1 | | Descriptor: | Fag s 1 pollen allergen | | Authors: | Moraes, A.H, Asam, A, Almeida, F.C.L, Wallner, M, Ferreira, F, Valente, A.P. | | Deposit date: | 2017-08-08 | | Release date: | 2018-08-08 | | Last modified: | 2024-05-15 | | Method: | SOLUTION NMR | | Cite: | Structural basis for cross-reactivity and conformation fluctuation of the major beech pollen allergen Fag s 1.
Sci Rep, 8, 2018
|
|
1QSK
 
 | | NMR-DERIVED SOLUTION STRUCTURE OF A FIVE-ADENINE BULGE LOOP WITHIN A DNA DUPLEX | | Descriptor: | 5'-D(CP*GP*TP*AP*GP*CP*CP*GP*AP*TP*GP*CP)-3', 5'-D(GP*CP*AP*TP*CP*GP*AP*AP*AP*AP*AP*GP*CP*TP*AP*CP*GP)-3' | | Authors: | Dornberger, U, Hillisch, A, Gollmick, F, Fritzsche, H, Diekmann, S. | | Deposit date: | 1999-06-22 | | Release date: | 1999-11-05 | | Last modified: | 2024-05-22 | | Method: | SOLUTION NMR | | Cite: | Solution structure of a five-adenine bulge loop within a DNA duplex.
Biochemistry, 38, 1999
|
|
2G35
 
 | | NMR structure of talin-PTB in complex with PIPKI | | Descriptor: | Talin-1, peptide | | Authors: | Kong, X, Wang, X, Misra, S, Qin, J. | | Deposit date: | 2006-02-17 | | Release date: | 2006-05-02 | | Last modified: | 2022-03-09 | | Method: | SOLUTION NMR | | Cite: | Structural Basis for the Phosphorylation-regulated Focal Adhesion Targeting of Type Igamma Phosphatidylinositol Phosphate Kinase (PIPKIgamma) by Talin.
J.Mol.Biol., 359, 2006
|
|
1QM9
 
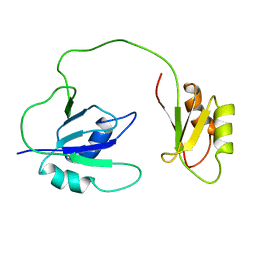 | | NMR, REPRESENTATIVE STRUCTURE | | Descriptor: | POLYPYRIMIDINE TRACT-BINDING PROTEIN | | Authors: | Conte, M.R, Grune, T, Curry, S, Matthews, S. | | Deposit date: | 1999-09-22 | | Release date: | 2000-07-03 | | Last modified: | 2024-05-15 | | Method: | SOLUTION NMR | | Cite: | Structure of Tandem RNA Recognition Motifs from Polypyrimidine Tract Binding Protein Reveals Novel Features of the Rrm Fold
Embo J., 19, 2000
|
|
2RLJ
 
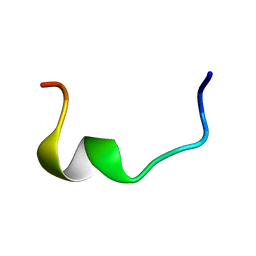 | | NMR Structure of Ebola fusion peptide in SDS micelles at pH 7 | | Descriptor: | Envelope glycoprotein | | Authors: | Freitas, M.S, Gaspar, L.P, Lorenzoni, M, Almeida, F.C, Tinoco, L.W, Almeida, M.S, Maia, L.F, Degreve, L, Valente, A.P, Silva, J.L. | | Deposit date: | 2007-07-05 | | Release date: | 2007-08-07 | | Last modified: | 2024-05-29 | | Method: | SOLUTION NMR | | Cite: | Structure of the Ebola fusion peptide in a membrane-mimetic environment and the interaction with lipid rafts.
J.Biol.Chem., 282, 2007
|
|
7NRN
 | | NMR structure of GIPC1-GH2 domain | | Descriptor: | PDZ domain-containing protein GIPC1 | | Authors: | Barthe, P, Roumestand, C. | | Deposit date: | 2021-03-04 | | Release date: | 2021-04-14 | | Last modified: | 2024-06-19 | | Method: | SOLUTION NMR | | Cite: | Pressure and Chemical Unfolding of an alpha-Helical Bundle Protein: The GH2 Domain of the Protein Adaptor GIPC1.
Int J Mol Sci, 22, 2021
|
|
1N5G
 
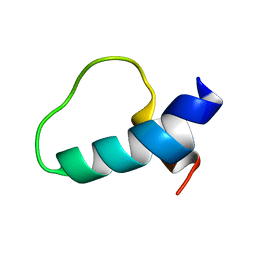 | |
1OVX
 
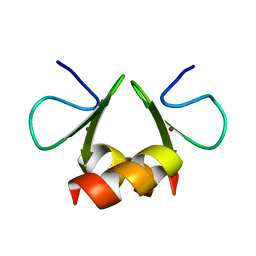 | | NMR structure of the E. coli ClpX chaperone zinc binding domain dimer | | Descriptor: | ATP-dependent Clp protease ATP-binding subunit clpX, ZINC ION | | Authors: | Donaldson, L.W, Kwan, J, Wojtyra, U, Houry, W.A. | | Deposit date: | 2003-03-27 | | Release date: | 2003-12-30 | | Last modified: | 2024-05-01 | | Method: | SOLUTION NMR | | Cite: | Solution structure of the dimeric zinc binding domain of the chaperone ClpX.
J.Biol.Chem., 278, 2003
|
|
1D1D
 
 | |
1P9F
 
 | |
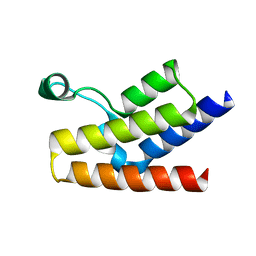
1F68
 | |
1E5U
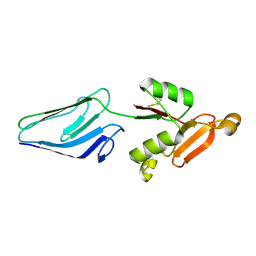 
 | | NMR Representative Structure of Intimin-190 (Int190) from Enteropathogenic E. coli | | Descriptor: | INTIMIN | | Authors: | Prasannan, S, Matthews, S.J, Batchelor, M, Daniell, S, Reece, S, Frankel, G, Dougan, G, Connerton, I, Bloomberg, G. | | Deposit date: | 2000-08-02 | | Release date: | 2000-08-16 | | Last modified: | 2011-07-13 | | Method: | SOLUTION NMR | | Cite: | Structural Basis for Recognition of the Translocated Intimin Receptor (Tir) by Intimin from Enteropathogenic E. Coli
Embo J., 19, 2000
|
|
1DQB
 
 | |
7D0X
 
 | |
1GO0
 
 | |
1QWB
 
 | |
1PV3
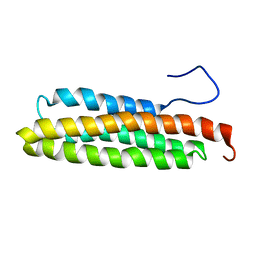 
 | | NMR Solution Structure of the Avian FAT-domain of Focal Adhesion Kinase | | Descriptor: | Focal adhesion kinase 1 | | Authors: | Prutzman, K.C, Gao, G, King, M.L, Iyer, V.V, Mueller, G.A, Schaller, M.D, Campbell, S.L. | | Deposit date: | 2003-06-26 | | Release date: | 2004-05-25 | | Last modified: | 2024-05-01 | | Method: | SOLUTION NMR | | Cite: | The Focal Adhesion Targeting Domain of Focal Adhesion Kinase Contains a Hinge Region that Modulates Tyrosine 926 Phosphorylation.
STRUCTURE, 12, 2004
|
|
2RLQ
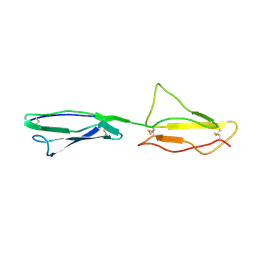 
 | | NMR structure of CCP modules 2-3 of complement factor H | | Descriptor: | Complement factor H | | Authors: | Hocking, H.G, Herbert, A.P, Pangburn, M.K, Kavanagh, D, Barlow, P.N, Uhrin, D. | | Deposit date: | 2007-07-29 | | Release date: | 2008-02-19 | | Last modified: | 2022-03-16 | | Method: | SOLUTION NMR | | Cite: | Structure of the N-terminal region of complement factor H and conformational implications of disease-linked sequence variations.
J.Biol.Chem., 283, 2008
|
|
2RQK
 
 | | NMR Solution Structure of Mesoderm Development (MESD) - closed conformation | | Descriptor: | Mesoderm development candidate 2 | | Authors: | Koehler, C, Lighthouse, J.K, Werther, T, Andersen, O.M, Diehl, A, Schmieder, P, Holdener, B.C, Oschkinat, H. | | Deposit date: | 2009-08-06 | | Release date: | 2009-08-25 | | Last modified: | 2024-05-01 | | Method: | SOLUTION NMR | | Cite: | The Structure of MESD45-184 Brings Light into the Mechanism of LDLR Family Folding
Structure, 19, 2011
|
|
2RLP
 
 | | NMR structure of CCP modules 1-2 of complement factor H | | Descriptor: | Complement factor H | | Authors: | Hocking, H.G, Herbert, A.P, Pangburn, M.K, Kavanagh, D, Barlow, P.N, Uhrin, D. | | Deposit date: | 2007-07-28 | | Release date: | 2008-02-19 | | Last modified: | 2022-03-16 | | Method: | SOLUTION NMR | | Cite: | Structure of the N-terminal region of complement factor H and conformational implications of disease-linked sequence variations.
J.Biol.Chem., 283, 2008
|
|
1F5X
 
 | | NMR STRUCTURE OF THE Y174 AUTOINHIBITED DBL HOMOLOGY DOMAIN | | Descriptor: | RHO-GEF VAV | | Authors: | Aghazadeh, B, Rosen, M.K, Lowry, W.E, Huang, X.Y. | | Deposit date: | 2000-06-18 | | Release date: | 2000-09-15 | | Last modified: | 2024-05-22 | | Method: | SOLUTION NMR | | Cite: | Structural basis for relief of autoinhibition of the Dbl homology domain of proto-oncogene Vav by tyrosine phosphorylation.
Cell(Cambridge,Mass.), 102, 2000
|
|
