8UCR
 
 | |
8UCQ
 
 | |
8UCI
 
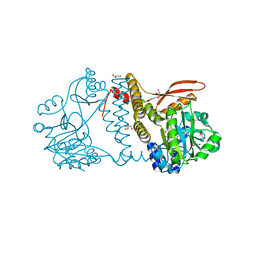 | | Thermophilic RNA Ligase from Palaeococcus pacificus K238G + AMP | | Descriptor: | ADENOSINE MONOPHOSPHATE, ATP dependent DNA ligase, GLYCEROL, ... | | Authors: | Rousseau, M.D, Hicks, J.L, Oulavallickal, T, Williamson, A, Arcus, V.L, Patrick, M.W. | | Deposit date: | 2023-09-26 | | Release date: | 2024-02-21 | | Last modified: | 2024-05-08 | | Method: | X-RAY DIFFRACTION (2.14 Å) | | Cite: | Characterisation and engineering of a thermophilic RNA ligase from Palaeococcus pacificus.
Nucleic Acids Res., 52, 2024
|
|
8UCH
 
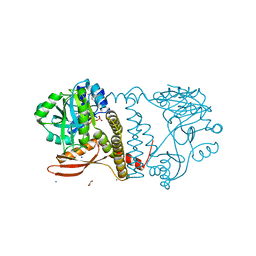 | | Thermophilic RNA Ligase from Palaeococcus pacificus K92A + ATP | | Descriptor: | ADENOSINE-5'-TRIPHOSPHATE, ATP dependent DNA ligase, GLYCEROL, ... | | Authors: | Rousseau, M.D, Hicks, J.L, Oulavallickal, T, Williamson, A, Arcus, V.L, Patrick, M.W. | | Deposit date: | 2023-09-26 | | Release date: | 2024-02-21 | | Last modified: | 2024-05-08 | | Method: | X-RAY DIFFRACTION (2.14 Å) | | Cite: | Characterisation and engineering of a thermophilic RNA ligase from Palaeococcus pacificus.
Nucleic Acids Res., 52, 2024
|
|
8UCG
 
 | | Thermophilic RNA Ligase from Palaeococcus pacificus K92A | | Descriptor: | ATP dependent DNA ligase, GLYCEROL, MAGNESIUM ION, ... | | Authors: | Rousseau, M.D, Hicks, J.L, Oulavallickal, T, Williamson, A, Arcus, V.L, Patrick, M.W. | | Deposit date: | 2023-09-26 | | Release date: | 2024-02-21 | | Last modified: | 2024-05-08 | | Method: | X-RAY DIFFRACTION (1.86 Å) | | Cite: | Characterisation and engineering of a thermophilic RNA ligase from Palaeococcus pacificus.
Nucleic Acids Res., 52, 2024
|
|
8UCF
 
 | | Thermophilic RNA Ligase from Palaeococcus pacificus K238G | | Descriptor: | ATP dependent DNA ligase, GLYCEROL, MAGNESIUM ION | | Authors: | Rousseau, M.D, Hicks, J.L, Oulavallickal, T, Williamson, A, Arcus, V.L, Patrick, M.W. | | Deposit date: | 2023-09-26 | | Release date: | 2024-02-21 | | Last modified: | 2024-05-08 | | Method: | X-RAY DIFFRACTION (1.9 Å) | | Cite: | Characterisation and engineering of a thermophilic RNA ligase from Palaeococcus pacificus.
Nucleic Acids Res., 52, 2024
|
|
8UCE
 
 | | Thermophilic RNA Ligase from Palaeococcus pacificus + AMP | | Descriptor: | ADENOSINE MONOPHOSPHATE, ATP dependent DNA ligase, GLYCEROL, ... | | Authors: | Rousseau, M.D, Hicks, J.L, Oulavallickal, T, Williamson, A, Arcus, V.L, Patrick, M.W. | | Deposit date: | 2023-09-26 | | Release date: | 2024-02-21 | | Last modified: | 2024-05-08 | | Method: | X-RAY DIFFRACTION (2.12 Å) | | Cite: | Characterisation and engineering of a thermophilic RNA ligase from Palaeococcus pacificus.
Nucleic Acids Res., 52, 2024
|
|
8UCD
 
 | | Cryo-EM structure of human STEAP1 in complex with AMG 509 Fab | | Descriptor: | 1-palmitoyl-2-oleoyl-sn-glycero-3-phosphocholine, AMG 509 anti-STEAP1 Fab, heavy chain, ... | | Authors: | Li, F, Bailis, J.M, Zhang, H. | | Deposit date: | 2023-09-26 | | Release date: | 2023-11-22 | | Last modified: | 2024-01-24 | | Method: | ELECTRON MICROSCOPY (3 Å) | | Cite: | AMG 509 (Xaluritamig), an Anti-STEAP1 XmAb 2+1 T-cell Redirecting Immune Therapy with Avidity-Dependent Activity against Prostate Cancer.
Cancer Discov, 14, 2024
|
|
8UCC
 
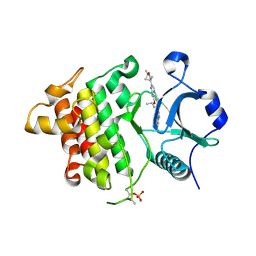 | | IRAK4 in complex with compound 20 | | Descriptor: | (4S)-2-[(1R,4s)-1-methyl-2-oxabicyclo[2.1.1]hexan-4-yl]-N-[(8S)-6-methylpyrazolo[1,5-a]pyrimidin-3-yl]-7-[(propan-2-yl)oxy]imidazo[1,2-a]pyrimidine-6-carboxamide, Interleukin-1 receptor-associated kinase 4 | | Authors: | Metrick, C.M, Chodaparambil, J.V. | | Deposit date: | 2023-09-26 | | Release date: | 2024-06-26 | | Method: | X-RAY DIFFRACTION (1.8 Å) | | Cite: | The Discovery of 7-Isopropoxy-2-(1-methyl-2-oxabicyclo[2.1.1]hexan-4-yl)- N -(6-methylpyrazolo[1,5- a ]pyrimidin-3-yl)imidazo[1,2- a ]pyrimidine-6-carboxamide (BIO-7488), a Potent, Selective, and CNS-Penetrant IRAK4 Inhibitor for the Treatment of Ischemic Stroke.
J.Med.Chem., 67, 2024
|
|
8UCB
 
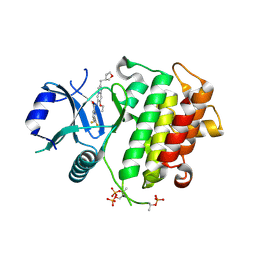 | | IRAK4 in complex with compound 8 | | Descriptor: | 6-(difluoromethyl)-N-[(4R)-7-ethoxy-2-{[(3R)-oxolan-3-yl]methyl}imidazo[1,2-a]pyridin-6-yl]pyridine-2-carboxamide, Interleukin-1 receptor-associated kinase 4 | | Authors: | Metrick, C.M, Chodaparambil, J.V. | | Deposit date: | 2023-09-26 | | Release date: | 2024-06-26 | | Method: | X-RAY DIFFRACTION (1.85 Å) | | Cite: | The Discovery of 7-Isopropoxy-2-(1-methyl-2-oxabicyclo[2.1.1]hexan-4-yl)- N -(6-methylpyrazolo[1,5- a ]pyrimidin-3-yl)imidazo[1,2- a ]pyrimidine-6-carboxamide (BIO-7488), a Potent, Selective, and CNS-Penetrant IRAK4 Inhibitor for the Treatment of Ischemic Stroke.
J.Med.Chem., 67, 2024
|
|
8UC9
 
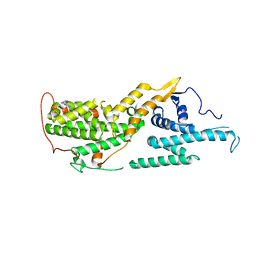 | |
8UC6
 
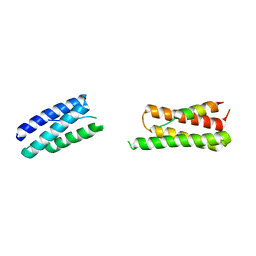 | | Calpain-7:IST1 Complex | | Descriptor: | Calpain-7, IST1 homolog | | Authors: | Paine, E, Whitby, F.G, Hill, C.P, Sunquist, W.I. | | Deposit date: | 2023-09-25 | | Release date: | 2023-10-25 | | Last modified: | 2024-05-15 | | Method: | X-RAY DIFFRACTION (2.701 Å) | | Cite: | The Calpain-7 protease functions together with the ESCRT-III protein IST1 within the midbody to regulate the timing and completion of abscission.
Elife, 12, 2023
|
|
8UC3
 
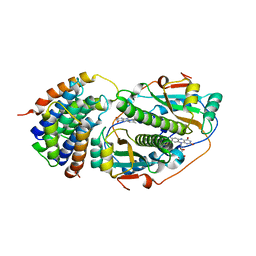 | |
8UC2
 
 | | Ethylene forming enzyme (EFE) R171A variant in complex with nickel and Benzoic acid | | Descriptor: | 1,2-ETHANEDIOL, 2-oxoglutarate-dependent ethylene/succinate-forming enzyme, BENZOIC ACID, ... | | Authors: | Chatterjee, S, Rankin, J.A, Hu, J, Hausinger, R. | | Deposit date: | 2023-09-25 | | Release date: | 2023-10-04 | | Last modified: | 2024-05-01 | | Method: | X-RAY DIFFRACTION (1.6 Å) | | Cite: | Structural, Spectroscopic, and Computational Insights from Canavanine-Bound and Two Catalytically Compromised Variants of the Ethylene-Forming Enzyme.
Biochemistry, 63, 2024
|
|
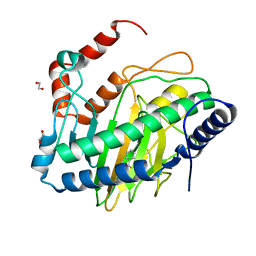
8UC1
 
 | |
8UC0
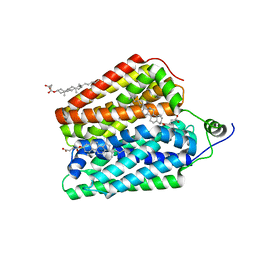 
 | | Endogenous ligand bound FLVCR1 | | Descriptor: | CHOLESTEROL HEMISUCCINATE, Heme transporter FLVCR1 | | Authors: | Hite, R.K, Son, Y. | | Deposit date: | 2023-09-25 | | Release date: | 2024-03-27 | | Last modified: | 2024-05-29 | | Method: | ELECTRON MICROSCOPY (2.42 Å) | | Cite: | Structural basis of lipid head group entry to the Kennedy pathway by FLVCR1.
Nature, 629, 2024
|
|
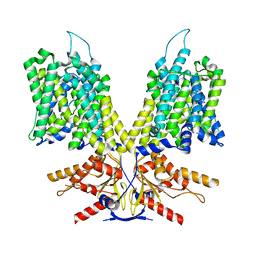
8UBZ
 
 | | Choline-bound FLVCR1 | | Descriptor: | CHOLESTEROL HEMISUCCINATE, CHOLINE ION, Heme transporter FLVCR1 | | Authors: | Hite, R.K, Son, Y. | | Deposit date: | 2023-09-25 | | Release date: | 2024-03-27 | | Last modified: | 2024-05-29 | | Method: | ELECTRON MICROSCOPY (3.02 Å) | | Cite: | Structural basis of lipid head group entry to the Kennedy pathway by FLVCR1.
Nature, 629, 2024
|
|
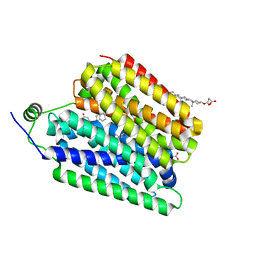
8UBY
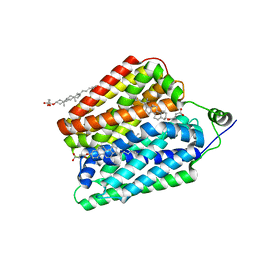 
 | | Choline-bound FLVCR1 | | Descriptor: | CHOLESTEROL HEMISUCCINATE, CHOLINE ION, Heme transporter FLVCR1 | | Authors: | Hite, R.K, Son, Y. | | Deposit date: | 2023-09-25 | | Release date: | 2024-03-27 | | Last modified: | 2024-05-29 | | Method: | ELECTRON MICROSCOPY (2.67 Å) | | Cite: | Structural basis of lipid head group entry to the Kennedy pathway by FLVCR1.
Nature, 629, 2024
|
|
8UBX
 
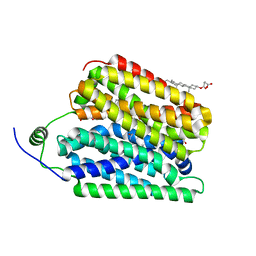 | | Ethanolamine-bound FLVCR1 | | Descriptor: | CHOLESTEROL HEMISUCCINATE, ETHANOLAMINE, Heme transporter FLVCR1 | | Authors: | Hite, R.K, Son, Y. | | Deposit date: | 2023-09-25 | | Release date: | 2024-03-27 | | Last modified: | 2024-05-29 | | Method: | ELECTRON MICROSCOPY (2.5 Å) | | Cite: | Structural basis of lipid head group entry to the Kennedy pathway by FLVCR1.
Nature, 629, 2024
|
|
8UBW
 
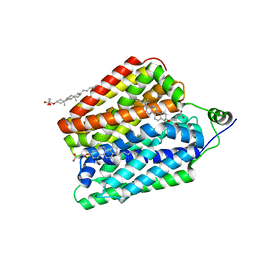 | | Choline-bound FLVCR1 | | Descriptor: | CHOLESTEROL HEMISUCCINATE, CHOLINE ION, Heme transporter FLVCR1 | | Authors: | Hite, R.K, Son, Y. | | Deposit date: | 2023-09-25 | | Release date: | 2024-03-27 | | Last modified: | 2024-05-29 | | Method: | ELECTRON MICROSCOPY (2.59 Å) | | Cite: | Structural basis of lipid head group entry to the Kennedy pathway by FLVCR1.
Nature, 629, 2024
|
|
8UBR
 
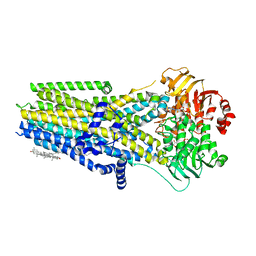 | | Complex of the phosphorylated human cystic fibrosis transmembrane conductance regulator (CFTR) with CFTRinh-172 and ATP/Mg | | Descriptor: | 1-palmitoyl-2-oleoyl-sn-glycero-3-phosphocholine, 4-[(Z)-{(3M)-4-oxo-2-sulfanylidene-3-[3-(trifluoromethyl)phenyl]-1,3-thiazolidin-5-ylidene}methyl]benzoic acid, ADENOSINE-5'-TRIPHOSPHATE, ... | | Authors: | Young, P.G, Fiedorczuk, K, Chen, J. | | Deposit date: | 2023-09-24 | | Release date: | 2024-02-21 | | Last modified: | 2024-03-20 | | Method: | ELECTRON MICROSCOPY (2.7 Å) | | Cite: | Structural basis for CFTR inhibition by CFTR inh -172.
Proc.Natl.Acad.Sci.USA, 121, 2024
|
|
8UBH
 
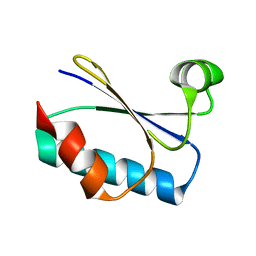 | |
8UBG
 
 | | DpHF19 filament | | Descriptor: | DpHF19,Green fluorescent protein (Fragment) | | Authors: | Lynch, E.M, Shen, H, Kollman, J.M, Baker, D. | | Deposit date: | 2023-09-22 | | Release date: | 2024-04-10 | | Method: | ELECTRON MICROSCOPY (3.5 Å) | | Cite: | De novo design of pH-responsive self-assembling helical protein filaments.
Nat Nanotechnol, 2024
|
|
8UB6
 
 | |
8UB3
 
 | | DpHF7 filament | | Descriptor: | DpHF7 filament | | Authors: | Lynch, E.M, Farrell, D, Shen, H, Kollman, J.M, DiMaio, F, Baker, D. | | Deposit date: | 2023-09-22 | | Release date: | 2024-04-10 | | Method: | ELECTRON MICROSCOPY (3.3 Å) | | Cite: | De novo design of pH-responsive self-assembling helical protein filaments.
Nat Nanotechnol, 2024
|
|
