6UUR
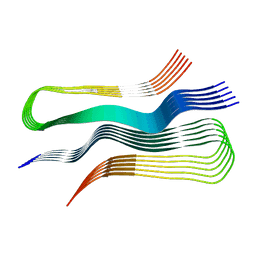 
 | | Human prion protein fibril, M129 variant | | 分子名称: | Major prion protein | | 著者 | Glynn, C, Sawaya, M.R, Ge, P, Zhou, Z.H, Rodriguez, J.A. | | 登録日 | 2019-10-31 | | 公開日 | 2020-04-15 | | 最終更新日 | 2024-03-06 | | 実験手法 | ELECTRON MICROSCOPY (3.5 Å) | | 主引用文献 | Cryo-EM structure of a human prion fibril with a hydrophobic, protease-resistant core.
Nat.Struct.Mol.Biol., 27, 2020
|
|
3TF6
 
 | |
3EHF
 
 | |
6PF0
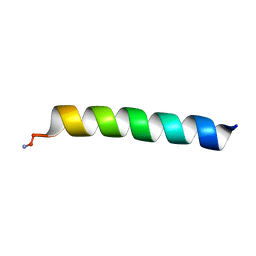 
 | |
6PEZ
 
 | |
4R6R
 
 | | Jacalin-carbohydrate interactions. Distortion of the ligand as a determinant of affinity. | | 分子名称: | 1,2-ETHANEDIOL, 4-nitrophenyl beta-D-galactopyranoside, Agglutinin alpha chain, ... | | 著者 | Abhinav, K.V, Sharma, K, Swaminathan, C.P, Surolia, A, Vijayan, M. | | 登録日 | 2014-08-26 | | 公開日 | 2015-02-18 | | 最終更新日 | 2023-11-08 | | 実験手法 | X-RAY DIFFRACTION (1.38 Å) | | 主引用文献 | Jacalin-carbohydrate interactions: distortion of the ligand molecule as a determinant of affinity.
Acta Crystallogr.,Sect.D, 71, 2015
|
|
6VJN
 
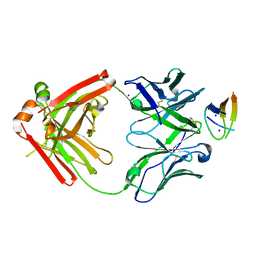 | | Structure of NHP D11A.B5Fab in complex with 16055 V2b peptide | | 分子名称: | D11A.B5 Fab Heavy chain, D11A.B5 Fab Light chain, SODIUM ION, ... | | 著者 | Singh, S, Liban, T.J, Pancera, M. | | 登録日 | 2020-01-16 | | 公開日 | 2021-01-20 | | 最終更新日 | 2023-10-11 | | 実験手法 | X-RAY DIFFRACTION (2 Å) | | 主引用文献 | Structurally related but genetically unrelated antibody lineages converge on an immunodominant HIV-1 Env neutralizing determinant following trimer immunization.
Plos Pathog., 17, 2021
|
|
4R6Q
 
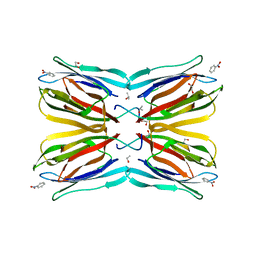 | | Jacalin-carbohydrate interactions. Distortion of the ligand as a determinant of affinity. | | 分子名称: | 1,2-ETHANEDIOL, Agglutinin alpha chain, Agglutinin beta-3 chain, ... | | 著者 | Abhinav, K.V, Sharma, K, Swaminathan, C.P, Surolia, A, Vijayan, M. | | 登録日 | 2014-08-26 | | 公開日 | 2015-02-18 | | 最終更新日 | 2023-11-08 | | 実験手法 | X-RAY DIFFRACTION (1.6 Å) | | 主引用文献 | Jacalin-carbohydrate interactions: distortion of the ligand molecule as a determinant of affinity.
Acta Crystallogr.,Sect.D, 71, 2015
|
|
6T7J
 
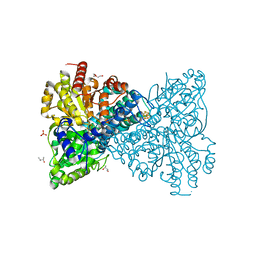 | | As-isolated Ni-free crystal structure of carbon monoxide dehydrogenase from Thermococcus sp. AM4 produced without CooC maturase | | 分子名称: | (R,R)-2,3-BUTANEDIOL, Broken Fe4S4 cluster, CITRIC ACID, ... | | 著者 | Dobbek, H, Jeoung, J.-H. | | 登録日 | 2019-10-22 | | 公開日 | 2020-11-18 | | 最終更新日 | 2024-01-24 | | 実験手法 | X-RAY DIFFRACTION (2.43 Å) | | 主引用文献 | The two CO-dehydrogenases of Thermococcus sp. AM4.
Biochim Biophys Acta Bioenerg, 1861, 2020
|
|
6WAS
 
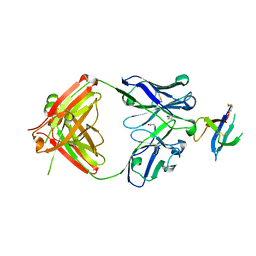 | | Structure of D19.PA8 Fab in complex with 1FD6 16055 V1V2 scaffold | | 分子名称: | 1FD6 16055 V1V2 scaffold, 2-acetamido-2-deoxy-beta-D-glucopyranose, GN1_PA8 Fab Heavy chain, ... | | 著者 | Singh, S, Liban, T.J, Pancera, M. | | 登録日 | 2020-03-26 | | 公開日 | 2021-03-31 | | 最終更新日 | 2023-10-18 | | 実験手法 | X-RAY DIFFRACTION (1.904 Å) | | 主引用文献 | Structurally related but genetically unrelated antibody lineages converge on an immunodominant HIV-1 Env neutralizing determinant following trimer immunization.
Plos Pathog., 17, 2021
|
|
3FOC
 
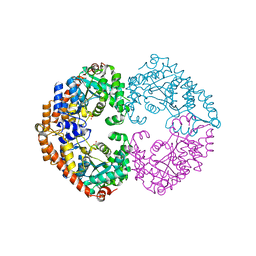 | |
4R6O
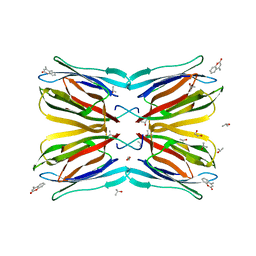 
 | | Jacalin-carbohydrate interactions. Distortion of the ligand as a determinant of affinity. | | 分子名称: | 1,2-ETHANEDIOL, 4-METHYL-2H-CHROMEN-2-ONE, Agglutinin alpha chain, ... | | 著者 | Abhinav, K.V, Sharma, K, Swaminathan, C.P, Surolia, A, Vijayan, M. | | 登録日 | 2014-08-26 | | 公開日 | 2015-02-18 | | 最終更新日 | 2023-11-08 | | 実験手法 | X-RAY DIFFRACTION (1.6 Å) | | 主引用文献 | Jacalin-carbohydrate interactions: distortion of the ligand molecule as a determinant of affinity.
Acta Crystallogr.,Sect.D, 71, 2015
|
|
4R6P
 
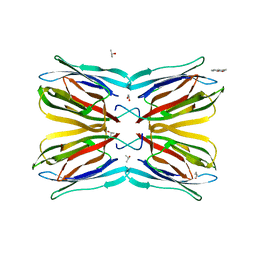 | | Jacalin-carbohydrate interactions. Distortion of the ligand as a determinant of affinity. | | 分子名称: | 1,2-ETHANEDIOL, 4-METHYL-2H-CHROMEN-2-ONE, Agglutinin alpha chain, ... | | 著者 | Abhinav, K.V, Sharma, K, Swaminathan, C.P, Surolia, A, Vijayan, M. | | 登録日 | 2014-08-26 | | 公開日 | 2015-02-18 | | 最終更新日 | 2023-11-08 | | 実験手法 | X-RAY DIFFRACTION (1.7 Å) | | 主引用文献 | Jacalin-carbohydrate interactions: distortion of the ligand molecule as a determinant of affinity.
Acta Crystallogr.,Sect.D, 71, 2015
|
|
4R6N
 
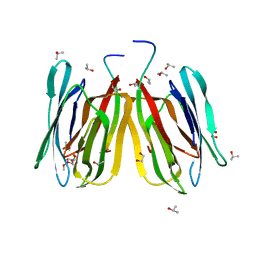 | | Jacalin-carbohydrate interactions. Distortion of the ligand as a determinant of affinity | | 分子名称: | 1,2-ETHANEDIOL, Agglutinin alpha chain, Agglutinin beta-3 chain, ... | | 著者 | Abhinav, K.V, Sharma, K, Swaminathan, C.P, Surolia, A, Vijayan, M. | | 登録日 | 2014-08-26 | | 公開日 | 2015-02-18 | | 最終更新日 | 2023-11-08 | | 実験手法 | X-RAY DIFFRACTION (1.67 Å) | | 主引用文献 | Jacalin-carbohydrate interactions: distortion of the ligand molecule as a determinant of affinity.
Acta Crystallogr.,Sect.D, 71, 2015
|
|
6U7U
 
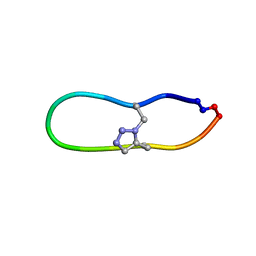 | | NMR solution structure of triazole bridged matriptase inhibitor | | 分子名称: | 1-methyl-1H-1,2,3-triazole, GLY-ARG-ALA-THR-LYS-SER-ILE-PRO-PRO-ARG-ALA-PHE-PRO-ASP | | 著者 | White, A.M, Harvey, P.J, Durek, T, Craik, D.J. | | 登録日 | 2019-09-03 | | 公開日 | 2020-04-22 | | 最終更新日 | 2020-07-15 | | 実験手法 | SOLUTION NMR | | 主引用文献 | Application and Structural Analysis of Triazole-Bridged Disulfide Mimetics in Cyclic Peptides.
Angew.Chem.Int.Ed.Engl., 59, 2020
|
|
6U7S
 
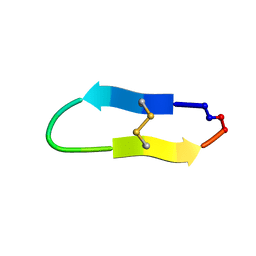 | |
6GIK
 | |
6WIQ
 
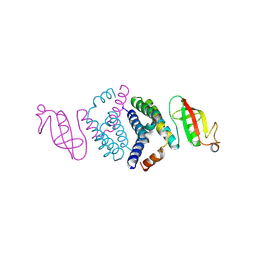 | | Crystal structure of the co-factor complex of NSP7 and the C-terminal domain of NSP8 from SARS CoV-2 | | 分子名称: | Non-structural protein 7, Non-structural protein 8 | | 著者 | Wilamowski, M, Kim, Y, Jedrzejczak, R, Maltseva, N, Endres, M, Godzik, A, Michalska, K, Joachimiak, A, Center for Structural Genomics of Infectious Diseases (CSGID) | | 登録日 | 2020-04-10 | | 公開日 | 2020-04-22 | | 最終更新日 | 2023-10-18 | | 実験手法 | X-RAY DIFFRACTION (2.85 Å) | | 主引用文献 | Transient and stabilized complexes of Nsp7, Nsp8, and Nsp12 in SARS-CoV-2 replication.
Biophys.J., 120, 2021
|
|
7MC3
 
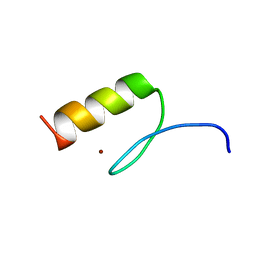 | | Solution structure of Miz-1 zinc finger 12 | | 分子名称: | Isoform 2 of Zinc finger and BTB domain-containing protein 17, ZINC ION | | 著者 | Boisvert, O, Lavigne, P. | | 登録日 | 2021-04-01 | | 公開日 | 2021-04-21 | | 最終更新日 | 2024-05-15 | | 実験手法 | SOLUTION NMR | | 主引用文献 | Zinc Fingers 10 and 11 of Miz-1 undergo conformational exchange to achieve specific DNA binding.
Structure, 30, 2022
|
|
6T3I
 
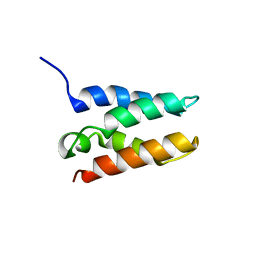 | | Solution structure of the HRP2 IBD | | 分子名称: | Hepatoma-derived growth factor-related protein 2 | | 著者 | Veverka, V. | | 登録日 | 2019-10-11 | | 公開日 | 2020-11-18 | | 最終更新日 | 2024-06-19 | | 実験手法 | SOLUTION NMR | | 主引用文献 | Unlike Its Paralog LEDGF/p75, HRP-2 Is Dispensable for MLL-R Leukemogenesis but Important for Leukemic Cell Survival.
Cells, 10, 2021
|
|
6U7Q
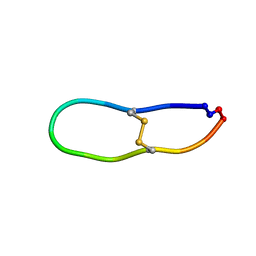 
 | |
6U24
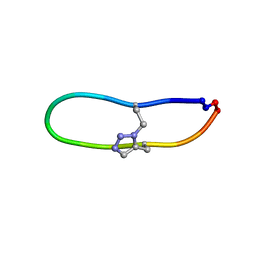 
 | | NMR solution structure of triazole bridged SFTI-1 | | 分子名称: | 1-methyl-1H-1,2,3-triazole, GLY-ARG-ALA-THR-LYS-SER-ILE-PRO-PRO-ILE-ALA-PHE-PRO-ASP | | 著者 | White, A.M, Harvey, P.J, Durek, T, Craik, D.J. | | 登録日 | 2019-08-19 | | 公開日 | 2020-07-01 | | 最終更新日 | 2020-07-15 | | 実験手法 | SOLUTION NMR | | 主引用文献 | Application and Structural Analysis of Triazole-Bridged Disulfide Mimetics in Cyclic Peptides.
Angew.Chem.Int.Ed.Engl., 59, 2020
|
|
7MC1
 
 | | Solution structure of Miz-1 Zinc finger 10 | | 分子名称: | Isoform 2 of Zinc finger and BTB domain-containing protein 17, ZINC ION | | 著者 | Boisvert, O, Lavigne, P. | | 登録日 | 2021-04-01 | | 公開日 | 2021-04-28 | | 最終更新日 | 2024-05-15 | | 実験手法 | SOLUTION NMR | | 主引用文献 | Zinc Fingers 10 and 11 of Miz-1 undergo conformational exchange to achieve specific DNA binding.
Structure, 30, 2022
|
|
7MC2
 
 | |
6U7W
 
 | | NMR solution structure of a triazole bridged KLK7 inhibitor | | 分子名称: | 1-methyl-1H-1,2,3-triazole, GLY-LYS-ALA-LEU-PHE-SER-ASN-PRO-PRO-ILE-ALA-PHE-PRO-ASN | | 著者 | White, A.M, Harvey, P.J, Durek, T, Craik, D.J. | | 登録日 | 2019-09-03 | | 公開日 | 2020-04-22 | | 最終更新日 | 2020-07-15 | | 実験手法 | SOLUTION NMR | | 主引用文献 | Application and Structural Analysis of Triazole-Bridged Disulfide Mimetics in Cyclic Peptides.
Angew.Chem.Int.Ed.Engl., 59, 2020
|
|
