1IQT
 
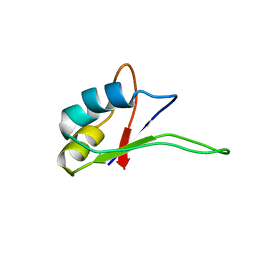 | | Solution structure of the C-terminal RNA-binding domain of heterogeneous nuclear ribonucleoprotein D0 (AUF1) | | 分子名称: | heterogeneous nuclear ribonucleoprotein D0 | | 著者 | Katahira, M, Miyanoiri, Y, Enokizono, Y, Matsuda, G, Nagata, T, Ishikawa, F, Uesugi, S. | | 登録日 | 2001-08-01 | | 公開日 | 2002-08-07 | | 最終更新日 | 2023-12-27 | | 実験手法 | SOLUTION NMR | | 主引用文献 | Structure of the C-terminal RNA-binding domain of hnRNP D0 (AUF1), its interactions with RNA and DNA, and change in backbone dynamics upon complex formation with DNA.
J.Mol.Biol., 311, 2001
|
|
1B7F
 
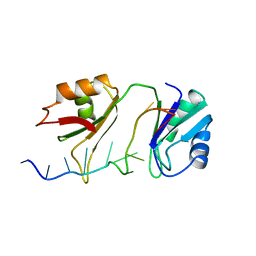 | | SXL-LETHAL PROTEIN/RNA COMPLEX | | 分子名称: | PROTEIN (SXL-LETHAL PROTEIN), RNA (5'-R(P*GP*UP*UP*GP*UP*UP*UP*UP*UP*UP*UP*U)-3') | | 著者 | Handa, N, Nureki, O, Kurimoto, K, Kim, I, Sakamoto, H, Shimura, Y, Muto, Y, Yokoyama, S, RIKEN Structural Genomics/Proteomics Initiative (RSGI) | | 登録日 | 1999-01-23 | | 公開日 | 1999-05-03 | | 最終更新日 | 2023-12-27 | | 実験手法 | X-RAY DIFFRACTION (2.6 Å) | | 主引用文献 | Structural basis for recognition of the tra mRNA precursor by the Sex-lethal protein.
Nature, 398, 1999
|
|
1DZ5
 
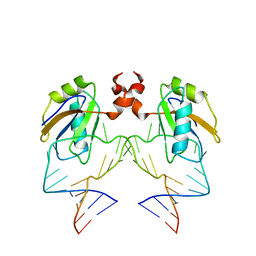 | | The NMR structure of the 38KDa U1A protein-PIE RNA complex reveals the basis of cooperativity in regulation of polyadenylation by human U1A protein | | 分子名称: | PIE, RNA (5'-R(*GP*AP*GP*AP*CP*AP*UP*UP*GP*CP*AP*CP*CP* CP*GP*GP*AP*GP*UP*CP*UP*C)-3'), U1 SMALL NUCLEAR RIBONUCLEOPROTEIN A | | 著者 | Varani, L, Gunderson, S.I, Mattaj, I.W, Kay, L.E, Neuhaus, D, Varani, G. | | 登録日 | 2000-02-16 | | 公開日 | 2000-03-29 | | 最終更新日 | 2024-05-15 | | 実験手法 | SOLUTION NMR | | 主引用文献 | The NMR Structure of the 38kDa U1A Protein-Pie RNA Complex Reveals the Basis of Cooperativity in Regulation of Polyadenylation by Human U1A Protein
Nat.Struct.Biol., 7, 2000
|
|
1M5K
 
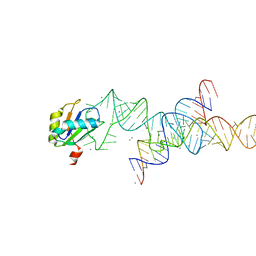 | |
1M5V
 
 | | Transition State Stabilization by a Catalytic RNA | | 分子名称: | (4S)-2-METHYL-2,4-PENTANEDIOL, CALCIUM ION, RNA HAIRPIN RIBOZYME, ... | | 著者 | Rupert, P.B, Massey, A.P, Sigurdsson, S.T, Ferre-D'Amare, A.R. | | 登録日 | 2002-07-09 | | 公開日 | 2002-10-12 | | 最終更新日 | 2024-02-14 | | 実験手法 | X-RAY DIFFRACTION (2.4 Å) | | 主引用文献 | Transition state stabilization by a catalytic RNA
Science, 298, 2002
|
|
1DRZ
 
 | | U1A SPLICEOSOMAL PROTEIN/HEPATITIS DELTA VIRUS GENOMIC RIBOZYME COMPLEX | | 分子名称: | MAGNESIUM ION, PROTEIN (U1 SMALL RIBONUCLEOPROTEIN A), RNA (HEPATITIS DELTA VIRUS GENOMIC RIBOZYME), ... | | 著者 | Ferre-D'Amare, A.R, Zhou, K, Doudna, J.A. | | 登録日 | 1998-09-01 | | 公開日 | 1999-02-16 | | 最終更新日 | 2021-11-03 | | 実験手法 | X-RAY DIFFRACTION (2.3 Å) | | 主引用文献 | Crystal structure of a hepatitis delta virus ribozyme.
Nature, 395, 1998
|
|
1M5O
 
 | | Transition State Stabilization by a Catalytic RNA | | 分子名称: | CALCIUM ION, RNA SUBSTRATE, RNA HAIRPIN RIBOZYME, ... | | 著者 | Rupert, P.B, Massey, A.P, Sigurdsson, S.T, Ferre-D'Amare, A.R. | | 登録日 | 2002-07-09 | | 公開日 | 2002-10-12 | | 最終更新日 | 2024-02-14 | | 実験手法 | X-RAY DIFFRACTION (2.2 Å) | | 主引用文献 | Transition state stabilization by a catalytic RNA
Science, 298, 2002
|
|
1M5P
 
 | | Transition State Stabilization by a Catalytic RNA | | 分子名称: | CALCIUM ION, RNA HAIRPIN RIBOZYME, RNA INHIBITOR SUBSTRATE, ... | | 著者 | Rupert, P.B, Massey, A, Sigurdsson, S.T, Ferre-D'Amare, A.R. | | 登録日 | 2002-07-09 | | 公開日 | 2002-10-12 | | 最終更新日 | 2024-02-14 | | 実験手法 | X-RAY DIFFRACTION (2.6 Å) | | 主引用文献 | Transition state stabilization by a catalytic RNA
Science, 298, 2002
|
|
3IRW
 
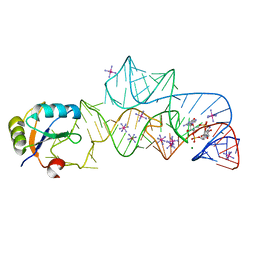 | | Structure of a c-di-GMP riboswitch from V. cholerae | | 分子名称: | 9,9'-[(2R,3R,3aS,5S,7aR,9R,10R,10aS,12S,14aR)-3,5,10,12-tetrahydroxy-5,12-dioxidooctahydro-2H,7H-difuro[3,2-d:3',2'-j][1,3,7,9,2,8]tetraoxadiphosphacyclododecine-2,9-diyl]bis(2-amino-1,9-dihydro-6H-purin-6-one), IRIDIUM HEXAMMINE ION, MAGNESIUM ION, ... | | 著者 | Smith, K.D. | | 登録日 | 2009-08-24 | | 公開日 | 2009-11-10 | | 最終更新日 | 2024-02-21 | | 実験手法 | X-RAY DIFFRACTION (2.7 Å) | | 主引用文献 | Structural basis of ligand binding by a c-di-GMP riboswitch.
Nat.Struct.Mol.Biol., 16, 2009
|
|
5IQQ
 
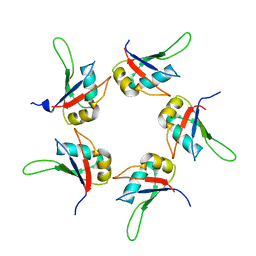 | |
5ITH
 
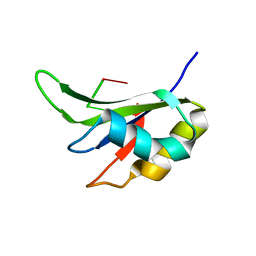 | |
3IIN
 
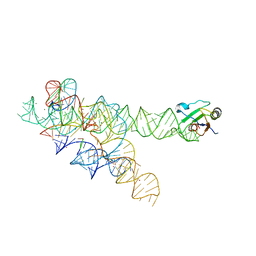 | | Plasticity of the kink turn structural motif | | 分子名称: | DNA/RNA (5'-R(*AP*AP*GP*CP*CP*AP*CP*AP*CP*AP*GP*AP*CP*C)-D(P*AP*GP*A)-R(P*CP*GP*GP*CP*C)-3'), DNA/RNA (5'-R(*CP*A)-D(P*T)-3'), Group I intron, ... | | 著者 | Lipchock, S.V, Strobel, S.A, Antonioli, A.H, Cochrane, J.C. | | 登録日 | 2009-08-02 | | 公開日 | 2010-03-09 | | 最終更新日 | 2023-09-06 | | 実験手法 | X-RAY DIFFRACTION (4.18 Å) | | 主引用文献 | Plasticity of the RNA kink turn structural motif.
Rna, 16, 2010
|
|
3K0J
 
 | | Crystal structure of the E. coli ThiM riboswitch in complex with thiamine pyrophosphate and the U1A crystallization module | | 分子名称: | MAGNESIUM ION, RNA (87-MER), THIAMINE DIPHOSPHATE, ... | | 著者 | Kulshina, N, Edwards, T.E, Ferre-D'Amare, A.R. | | 登録日 | 2009-09-24 | | 公開日 | 2009-12-22 | | 最終更新日 | 2024-02-21 | | 実験手法 | X-RAY DIFFRACTION (3.1 Å) | | 主引用文献 | Thermodynamic analysis of ligand binding and ligand binding-induced tertiary structure formation by the thiamine pyrophosphate riboswitch.
Rna, 16, 2010
|
|
5TBX
 
 | | hnRNP A18 RNA Recognition Motif | | 分子名称: | ACETATE ION, Cold-inducible RNA-binding protein, NICKEL (II) ION | | 著者 | Coburn, K.M, Melville, Z, Aligholizadeh, E, Roth, B.M, Varney, K.M, Weber, D.J. | | 登録日 | 2016-09-13 | | 公開日 | 2017-04-12 | | 最終更新日 | 2024-04-03 | | 実験手法 | X-RAY DIFFRACTION (1.767 Å) | | 主引用文献 | Crystal structure of the human heterogeneous ribonucleoprotein A18 RNA-recognition motif.
Acta Crystallogr F Struct Biol Commun, 73, 2017
|
|
5TKZ
 
 | |
5UZG
 
 | |
3L3C
 
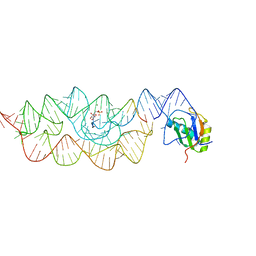 | | Crystal structure of the Bacillus anthracis glmS ribozyme bound to Glc6P | | 分子名称: | 6-O-phosphono-alpha-D-glucopyranose, GLMS RIBOZYME, MAGNESIUM ION, ... | | 著者 | Strobel, S.A, Cochrane, J.C, Lipchock, S.V, Smith, K.D. | | 登録日 | 2009-12-16 | | 公開日 | 2009-12-29 | | 最終更新日 | 2024-02-21 | | 実験手法 | X-RAY DIFFRACTION (2.85 Å) | | 主引用文献 | Structural and chemical basis for glucosamine 6-phosphate binding and activation of the glmS ribozyme
Biochemistry, 48, 2009
|
|
3IWN
 
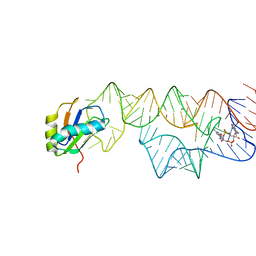 | | Co-crystal structure of a bacterial c-di-GMP riboswitch | | 分子名称: | 9,9'-[(2R,3R,3aS,5S,7aR,9R,10R,10aS,12S,14aR)-3,5,10,12-tetrahydroxy-5,12-dioxidooctahydro-2H,7H-difuro[3,2-d:3',2'-j][1,3,7,9,2,8]tetraoxadiphosphacyclododecine-2,9-diyl]bis(2-amino-1,9-dihydro-6H-purin-6-one), C-di-GMP riboswitch, U1 small nuclear ribonucleoprotein A | | 著者 | Kulshina, N, Baird, N.J, Ferre-D'Amare, A.R. | | 登録日 | 2009-09-02 | | 公開日 | 2009-11-10 | | 最終更新日 | 2024-02-21 | | 実験手法 | X-RAY DIFFRACTION (3.2 Å) | | 主引用文献 | Recognition of the bacterial second messenger cyclic diguanylate by its cognate riboswitch.
Nat.Struct.Mol.Biol., 16, 2009
|
|
5KW1
 
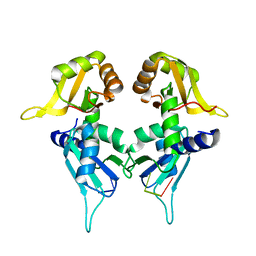 | | Crystal Structure of the Two Tandem RRM Domains of PUF60 Bound to a Modified AdML Pre-mRNA 3' Splice Site Analogue | | 分子名称: | CHLORIDE ION, DNA/RNA (30-MER), Poly(U)-binding-splicing factor PUF60 | | 著者 | Crichlow, G.V, Hsiao, H.-H, Albright, R, Lolis, E.J, Braddock, D.T. | | 登録日 | 2016-07-15 | | 公開日 | 2017-08-23 | | 最終更新日 | 2023-10-04 | | 実験手法 | X-RAY DIFFRACTION (2.1 Å) | | 主引用文献 | Unraveling the mechanism of recognition of the 3' splice site of the adenovirus major late promoter intron by the alternative splicing factor PUF60.
Plos One, 15, 2020
|
|
5KVY
 
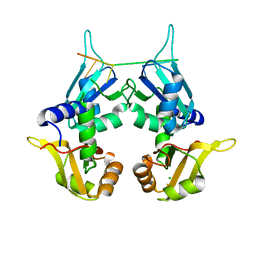 | | CRYSTAL STRUCTURE OF THE TWO TANDEM RRM DOMAINS OF PUF60 BOUND TO A PORTION OF AN ADML PRE-MRNA 3' SPLICE SITE ANALOG | | 分子名称: | CHLORIDE ION, DNA (30-MER), Poly(U)-binding-splicing factor PUF60 | | 著者 | Hsiao, H.-H, Crichlow, G.V, Albright, R.A, Murphy, J.W, Lolis, E.J, Braddock, D.T. | | 登録日 | 2016-07-15 | | 公開日 | 2017-08-23 | | 最終更新日 | 2023-10-04 | | 実験手法 | X-RAY DIFFRACTION (1.95 Å) | | 主引用文献 | Unraveling the mechanism of recognition of the 3' splice site of the adenovirus major late promoter intron by the alternative splicing factor PUF60.
Plos One, 15, 2020
|
|
5UZM
 
 | |
5SZW
 
 | | NMR solution structure of the RRM1 domain of the post-transcriptional regulator HuR | | 分子名称: | ELAV-like protein 1 | | 著者 | Lixa, C, Mujo, A, Jendiroba, K.A, Almeida, F.C.L, Lima, L.M.T.R, Pinheiro, A.S. | | 登録日 | 2016-08-15 | | 公開日 | 2017-09-06 | | 最終更新日 | 2024-05-01 | | 実験手法 | SOLUTION NMR | | 主引用文献 | Oligomeric transition and dynamics of RNA binding by the HuR RRM1 domain in solution.
J. Biomol. NMR, 72, 2018
|
|
5KWQ
 
 | | Two Tandem RRM Domains of FBP-Interacting Repressor (FIR), also Known as PUF60 | | 分子名称: | Poly(U)-binding-splicing factor PUF60 | | 著者 | Crichlow, G.V, Yang, Y, Zhou, H, Lolis, E.J, Braddock, D.T. | | 登録日 | 2016-07-18 | | 公開日 | 2017-08-23 | | 最終更新日 | 2023-10-04 | | 実験手法 | X-RAY DIFFRACTION (2.8 Å) | | 主引用文献 | Unraveling the mechanism of recognition of the 3' splice site of the adenovirus major late promoter intron by the alternative splicing factor PUF60.
Plos One, 15, 2020
|
|
3LPY
 
 | | Crystal structure of the RRM domain of CyP33 | | 分子名称: | 4-(2-HYDROXYETHYL)-1-PIPERAZINE ETHANESULFONIC ACID, Peptidyl-prolyl cis-trans isomerase E, SULFATE ION | | 著者 | Wang, Z, Patel, D.J. | | 登録日 | 2010-02-07 | | 公開日 | 2010-07-07 | | 最終更新日 | 2024-02-21 | | 実験手法 | X-RAY DIFFRACTION (2 Å) | | 主引用文献 | Pro isomerization in MLL1 PHD3-bromo cassette connects H3K4me readout to CyP33 and HDAC-mediated repression.
Cell(Cambridge,Mass.), 141, 2010
|
|
5T9P
 
 | |
