5DRE
 
 | |
5HHJ
 | |
4PVF
 
 | | Crystal structure of Homo sapiens holo serine hydroxymethyltransferase 2 (mitochondrial) (SHMT2), isoform 3, transcript variant 5, 483 aa, at 2.6 ang. resolution | | 分子名称: | DI(HYDROXYETHYL)ETHER, GLYCEROL, Serine hydroxymethyltransferase, ... | | 著者 | Giardina, G, Brunotti, P, Fiascarelli, A, Contestabile, R, Cutruzzola, F. | | 登録日 | 2014-03-17 | | 公開日 | 2015-01-28 | | 最終更新日 | 2023-12-06 | | 実験手法 | X-RAY DIFFRACTION (2.6 Å) | | 主引用文献 | How pyridoxal 5'-phosphate differentially regulates human cytosolic and mitochondrial serine hydroxymethyltransferase oligomeric state.
Febs J., 282, 2015
|
|
4U0I
 
 | | Crystal structure of KIT in complex with ponatinib | | 分子名称: | 3-(imidazo[1,2-b]pyridazin-3-ylethynyl)-4-methyl-N-{4-[(4-methylpiperazin-1-yl)methyl]-3-(trifluoromethyl)phenyl}benzam ide, Mast/stem cell growth factor receptor Kit,Mast/stem cell growth factor receptor Kit, PHOSPHATE ION | | 著者 | Zhou, T, Zhu, X, Dalgarno, D.C. | | 登録日 | 2014-07-11 | | 公開日 | 2014-10-08 | | 最終更新日 | 2023-12-27 | | 実験手法 | X-RAY DIFFRACTION (2 Å) | | 主引用文献 | Ponatinib Inhibits Polyclonal Drug-Resistant KIT Oncoproteins and Shows Therapeutic Potential in Heavily Pretreated Gastrointestinal Stromal Tumor (GIST) Patients.
Clin.Cancer Res., 20, 2014
|
|
3TOP
 
 | | Crystral Structure of the C-terminal Subunit of Human Maltase-Glucoamylase in Complex with Acarbose | | 分子名称: | 4,6-dideoxy-4-{[(1S,4R,5S,6S)-4,5,6-trihydroxy-3-(hydroxymethyl)cyclohex-2-en-1-yl]amino}-alpha-D-glucopyranose-(1-4)-alpha-D-glucopyranose-(1-4)-alpha-D-glucopyranose, Maltase-glucoamylase, intestinal | | 著者 | Shen, Y, Qin, X.H, Ren, L.M. | | 登録日 | 2011-09-06 | | 公開日 | 2011-11-23 | | 最終更新日 | 2023-11-01 | | 実験手法 | X-RAY DIFFRACTION (2.881 Å) | | 主引用文献 | Structural insight into substrate specificity of human intestinal maltase-glucoamylase
Protein Cell, 2, 2011
|
|
3TON
 
 | |
3V0D
 
 | | Crystal structure of Ciona intestinalis voltage sensor-containing phosphatase (Ci-VSP), residues 241-576(C363S) | | 分子名称: | PHOSPHATE ION, Voltage-sensor containing phosphatase | | 著者 | Liu, L, Kohout, S.C, Xu, Q, Muller, S, Kimberlin, C, Isacoff, E.Y, Minor, D.L. | | 登録日 | 2011-12-07 | | 公開日 | 2012-05-09 | | 最終更新日 | 2024-04-03 | | 実験手法 | X-RAY DIFFRACTION (1.1 Å) | | 主引用文献 | A glutamate switch controls voltage-sensitive phosphatase function.
Nat.Struct.Mol.Biol., 19, 2012
|
|
3V0J
 
 | | Crystal structure of Ciona intestinalis voltage sensor-containing phosphatase (Ci-VSP), residues 241-576(C363S), Deletion of 401-405 | | 分子名称: | PHOSPHATE ION, Voltage-sensor containing phosphatase | | 著者 | Liu, L, Kohout, S.C, Xu, Q, Muller, S, Kimberlin, C, Isacoff, E.Y, Minor, D.L. | | 登録日 | 2011-12-08 | | 公開日 | 2012-05-09 | | 最終更新日 | 2023-09-13 | | 実験手法 | X-RAY DIFFRACTION (1.72 Å) | | 主引用文献 | A glutamate switch controls voltage-sensitive phosphatase function.
Nat.Struct.Mol.Biol., 19, 2012
|
|
3V0I
 
 | | Crystal structure of Ciona intestinalis voltage sensor-containing phosphatase (Ci-VSP), residues 256-576, E411F | | 分子名称: | SULFATE ION, Voltage-sensor containing phosphatase | | 著者 | Liu, L, Kohout, S.C, Xu, Q, Muller, S, Kimberlin, C, Isacoff, E.Y, Minor, D.L. | | 登録日 | 2011-12-08 | | 公開日 | 2012-05-09 | | 最終更新日 | 2023-09-13 | | 実験手法 | X-RAY DIFFRACTION (1.95 Å) | | 主引用文献 | A glutamate switch controls voltage-sensitive phosphatase function.
Nat.Struct.Mol.Biol., 19, 2012
|
|
3V0E
 
 | | Crystal structure of Ciona intestinalis voltage sensor-containing phosphatase (Ci-VSP), residues 256-576(C363S) | | 分子名称: | CHLORIDE ION, GLYCEROL, SULFATE ION, ... | | 著者 | Liu, L, Kohout, S.C, Xu, Q, Muller, S, Kimberlin, C, Isacoff, E.Y, Minor, D.L. | | 登録日 | 2011-12-07 | | 公開日 | 2012-05-09 | | 最終更新日 | 2023-09-13 | | 実験手法 | X-RAY DIFFRACTION (1.65 Å) | | 主引用文献 | A glutamate switch controls voltage-sensitive phosphatase function.
Nat.Struct.Mol.Biol., 19, 2012
|
|
3V0H
 
 | | Crystal structure of Ciona intestinalis voltage sensor-containing phosphatase (Ci-VSP), residues 241-576(C363S), complexed with D-MYO-inositol-1,4,5-triphosphate | | 分子名称: | D-MYO-INOSITOL-1,4,5-TRIPHOSPHATE, Voltage-sensor containing phosphatase | | 著者 | Liu, L, Kohout, S.C, Xu, Q, Muller, S, Kimberlin, C, Isacoff, E.Y, Minor, D.L. | | 登録日 | 2011-12-08 | | 公開日 | 2012-05-09 | | 最終更新日 | 2023-09-13 | | 実験手法 | X-RAY DIFFRACTION (1.85 Å) | | 主引用文献 | A glutamate switch controls voltage-sensitive phosphatase function.
Nat.Struct.Mol.Biol., 19, 2012
|
|
3V0F
 
 | | Crystal structure of Ciona intestinalis voltage sensor-containing phosphatase (Ci-VSP), residues 241-576(C363S), form II | | 分子名称: | PHOSPHATE ION, Voltage-sensor containing phosphatase | | 著者 | Liu, L, Kohout, S.C, Xu, Q, Muller, S, Kimberlin, C, Isacoff, E.Y, Minor, D.L. | | 登録日 | 2011-12-07 | | 公開日 | 2012-05-09 | | 最終更新日 | 2023-09-13 | | 実験手法 | X-RAY DIFFRACTION (1.3 Å) | | 主引用文献 | A glutamate switch controls voltage-sensitive phosphatase function.
Nat.Struct.Mol.Biol., 19, 2012
|
|
3V0G
 
 | | Crystal structure of Ciona intestinalis voltage sensor-containing phosphatase (Ci-VSP), residues 241-576(C363S), form III | | 分子名称: | PHOSPHATE ION, Voltage-sensor containing phosphatase | | 著者 | Liu, L, Kohout, S.C, Xu, Q, Muller, S, Kimberlin, C, Isacoff, E.Y, Minor, D.L. | | 登録日 | 2011-12-08 | | 公開日 | 2012-05-09 | | 最終更新日 | 2023-09-13 | | 実験手法 | X-RAY DIFFRACTION (1.6 Å) | | 主引用文献 | A glutamate switch controls voltage-sensitive phosphatase function.
Nat.Struct.Mol.Biol., 19, 2012
|
|
4ADW
 
 | | CRYSTAL STRUCTURE OF LEISHMANIA INFANTUM TRYPANOTHIONE REDUCTASE IN COMPLEX WITH NADPH AND TRYPANOTHIONE | | 分子名称: | BIS(GAMMA-GLUTAMYL-CYSTEINYL-GLYCINYL)SPERMIDINE, FLAVIN-ADENINE DINUCLEOTIDE, NADPH DIHYDRO-NICOTINAMIDE-ADENINE-DINUCLEOTIDE PHOSPHATE, ... | | 著者 | Baiocco, P, Ilari, A, Colotti, G, Malatesta, F, Fiorillo, A. | | 登録日 | 2012-01-04 | | 公開日 | 2013-01-16 | | 最終更新日 | 2023-12-20 | | 実験手法 | X-RAY DIFFRACTION (3.61 Å) | | 主引用文献 | Inhibition of Leishmania Infantum Trypanothione Reductase by Azole-Based Compounds: A Comparative Analysis with its Physiological Substrate by X-Ray Crystallography.
Chemmedchem, 8, 2013
|
|
2BIF
 
 | | 6-PHOSPHOFRUCTO-2-KINASE/FRUCTOSE-2,6-BISPHOSPHATASE H256A MUTANT WITH F6P IN PHOSPHATASE ACTIVE SITE | | 分子名称: | 6-O-phosphono-beta-D-fructofuranose, MAGNESIUM ION, PHOSPHATE ION, ... | | 著者 | Yuen, M.H, Hasemann, C.A. | | 登録日 | 1998-10-26 | | 公開日 | 1999-02-16 | | 最終更新日 | 2023-08-23 | | 実験手法 | X-RAY DIFFRACTION (2.4 Å) | | 主引用文献 | Crystal structure of the H256A mutant of rat testis fructose-6-phosphate,2-kinase/fructose-2,6-bisphosphatase. Fructose 6-phosphate in the active site leads to mechanisms for both mutant and wild type bisphosphatase activities.
J.Biol.Chem., 274, 1999
|
|
4BGL
 
 | | Superoxide reductase (Neelaredoxin) from Archaeoglobus fulgidus | | 分子名称: | FE (III) ION, GLYCEROL, SUPEROXIDE REDUCTASE | | 著者 | Bandeiras, T.M, Rodrigues, J.V, Sousa, C.M, Barradas, A.R, Pinho, F.G, Pinto, A.F, Teixeira, M, Matias, P.M, Romao, C.V. | | 登録日 | 2013-03-27 | | 公開日 | 2014-04-09 | | 最終更新日 | 2024-05-08 | | 実験手法 | X-RAY DIFFRACTION (1.9 Å) | | 主引用文献 | Superoxide Reductase from Giardia Intestinalis: Structural Characterization of the First Sor from a Eukaryotic Organism Shows an Iron Centre that is Highly Sensitive to Photoreduction
Acta Crystallogr.,Sect.D, 71, 2015
|
|
3UZZ
 
 | | Crystal structure of 5beta-reductase (AKR1D1) E120H mutant in complex with NADP+ and delta4-androstenedione | | 分子名称: | 3-oxo-5-beta-steroid 4-dehydrogenase, 4-ANDROSTENE-3-17-DIONE, CHLORIDE ION, ... | | 著者 | Chen, M, Christianson, D.W, Penning, T.M. | | 登録日 | 2011-12-07 | | 公開日 | 2012-03-21 | | 最終更新日 | 2023-09-13 | | 実験手法 | X-RAY DIFFRACTION (1.82 Å) | | 主引用文献 | Conversion of Human Steroid 5beta-Reductase (AKR1D1) into 3β-Hydroxysteroid Dehydrogenase by Single Point Mutation E120H: EXAMPLE OF PERFECT ENZYME ENGINEERING.
J.Biol.Chem., 287, 2012
|
|
5GLU
 
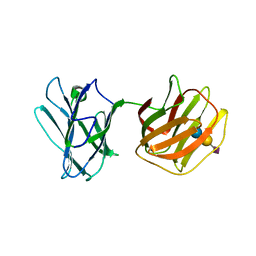 | | Tl-gal with SiaLac | | 分子名称: | GALECTIN, N-acetyl-alpha-neuraminic acid-(2-3)-beta-D-galactopyranose-(1-4)-alpha-D-glucopyranose | | 著者 | Jang, S.B, Hwang, E.Y. | | 登録日 | 2016-07-12 | | 公開日 | 2016-11-09 | | 最終更新日 | 2023-11-08 | | 実験手法 | X-RAY DIFFRACTION (2.1 Å) | | 主引用文献 | Structural Basis for Carbohydrate Recognition and Anti-inflammatory Modulation by Gastrointestinal Nematode Parasite Toxascaris leonina Galectin
J. Biol. Chem., 291, 2016
|
|
5GLZ
 
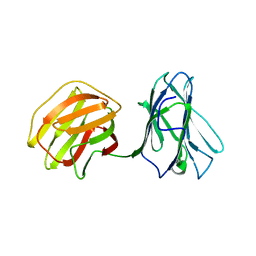 | | Tl-gal with Glucose | | 分子名称: | beta-D-glucopyranose, galectin | | 著者 | Jang, S.B, Hwang, E.Y. | | 登録日 | 2016-07-12 | | 公開日 | 2016-11-09 | | 最終更新日 | 2023-11-08 | | 実験手法 | X-RAY DIFFRACTION (2 Å) | | 主引用文献 | Structural Basis for Carbohydrate Recognition and Anti-inflammatory Modulation by Gastrointestinal Nematode Parasite Toxascaris leonina Galectin
J. Biol. Chem., 291, 2016
|
|
5GLT
 
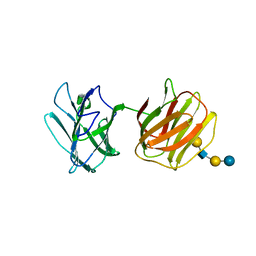 | | Tl-gal with LNT | | 分子名称: | Galectin, beta-D-galactopyranose-(1-3)-2-acetamido-2-deoxy-beta-D-glucopyranose-(1-3)-beta-D-galactopyranose-(1-4)-beta-D-glucopyranose | | 著者 | Jang, S.B, Hwang, E.Y. | | 登録日 | 2016-07-12 | | 公開日 | 2016-11-09 | | 最終更新日 | 2023-11-08 | | 実験手法 | X-RAY DIFFRACTION (2 Å) | | 主引用文献 | Structural Basis for Carbohydrate Recognition and Anti-inflammatory Modulation by Gastrointestinal Nematode Parasite Toxascaris leonina Galectin
J. Biol. Chem., 291, 2016
|
|
5GM0
 
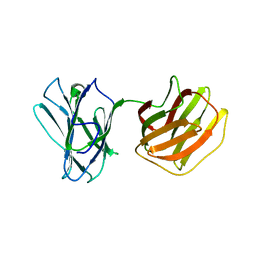 | | Tl-gal with lactose | | 分子名称: | beta-D-galactopyranose-(1-4)-beta-D-glucopyranose, galectin | | 著者 | Hwang, E.Y. | | 登録日 | 2016-07-12 | | 公開日 | 2016-11-09 | | 最終更新日 | 2023-11-08 | | 実験手法 | X-RAY DIFFRACTION (1.7 Å) | | 主引用文献 | Structural Basis for Carbohydrate Recognition and Anti-inflammatory Modulation by Gastrointestinal Nematode Parasite Toxascaris leonina Galectin
J. Biol. Chem., 291, 2016
|
|
5GLV
 
 | | Tl-gal | | 分子名称: | galectin | | 著者 | Jang, S.B, Hwang, E.Y. | | 登録日 | 2016-07-12 | | 公開日 | 2016-11-09 | | 最終更新日 | 2024-03-20 | | 実験手法 | X-RAY DIFFRACTION (1.8 Å) | | 主引用文献 | Structural Basis for Carbohydrate Recognition and Anti-inflammatory Modulation by Gastrointestinal Nematode Parasite Toxascaris leonina Galectin
J. Biol. Chem., 291, 2016
|
|
5GT1
 
 | | crystal structure of cbpa from L. salivarius REN | | 分子名称: | 1,2-ETHANEDIOL, ACETATE ION, Choline binding protein A, ... | | 著者 | Jiang, L, Ren, F. | | 登録日 | 2016-08-18 | | 公開日 | 2017-07-19 | | 最終更新日 | 2023-11-08 | | 実験手法 | X-RAY DIFFRACTION (1.85 Å) | | 主引用文献 | The Adhesion of Lactobacillus salivarius REN to a Human Intestinal Epithelial Cell Line Requires S-layer Proteins
Sci Rep, 7, 2017
|
|
5GLW
 
 | | Tl-gal with LacNAc | | 分子名称: | beta-D-galactopyranose-(1-4)-2-acetamido-2-deoxy-beta-D-glucopyranose, galectin | | 著者 | Jang, S.B, Hwang, E.Y. | | 登録日 | 2016-07-12 | | 公開日 | 2016-11-09 | | 最終更新日 | 2023-11-08 | | 実験手法 | X-RAY DIFFRACTION (2 Å) | | 主引用文献 | Structural Basis for Carbohydrate Recognition and Anti-inflammatory Modulation by Gastrointestinal Nematode Parasite Toxascaris leonina Galectin
J. Biol. Chem., 291, 2016
|
|
4MT5
 | |
