1SLJ
 
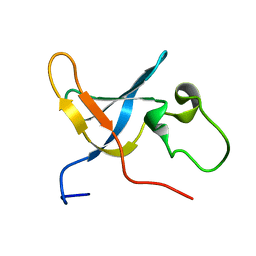 | | Solution structure of the S1 domain of RNase E from E. coli | | Descriptor: | Ribonuclease E | | Authors: | Schubert, M, Edge, R.E, Lario, P, Cook, M.A, Strynadka, N.C.J, Mackie, G.A, McIntosh, L.P. | | Deposit date: | 2004-03-05 | | Release date: | 2004-08-17 | | Last modified: | 2024-05-22 | | Method: | SOLUTION NMR | | Cite: | Structural characterization of the RNase E S1 domain and identification of its oligonucleotide-binding and dimerization interfaces.
J.Mol.Biol., 341, 2004
|
|
1SN8
 
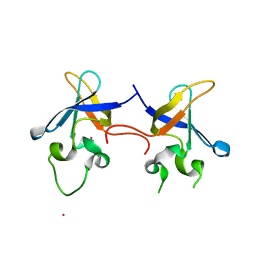 | | Crystal structure of the S1 domain of RNase E from E. coli (Pb derivative) | | Descriptor: | LEAD (II) ION, Ribonuclease E | | Authors: | Schubert, M, Edge, R.E, Lario, P, Cook, M.A, Strynadka, N.C.J, Mackie, G.A, McIntosh, L.P. | | Deposit date: | 2004-03-10 | | Release date: | 2004-08-17 | | Last modified: | 2024-02-14 | | Method: | X-RAY DIFFRACTION (2 Å) | | Cite: | Structural characterization of the RNase E S1 domain and identification of its oligonucleotide-binding and dimerization interfaces.
J.Mol.Biol., 341, 2004
|
|
1SMX
 
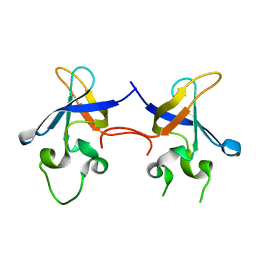 | | Crystal structure of the S1 domain of RNase E from E. coli (native) | | Descriptor: | Ribonuclease E | | Authors: | Schubert, M, Edge, R.E, Lario, P, Cook, M.A, Strynadka, N.C.J, Mackie, G.A, McIntosh, L.P. | | Deposit date: | 2004-03-09 | | Release date: | 2004-08-17 | | Last modified: | 2024-04-03 | | Method: | X-RAY DIFFRACTION (1.8 Å) | | Cite: | Structural characterization of the RNase E S1 domain and identification of its oligonucleotide-binding and dimerization interfaces.
J.Mol.Biol., 341, 2004
|
|
1SRO
 
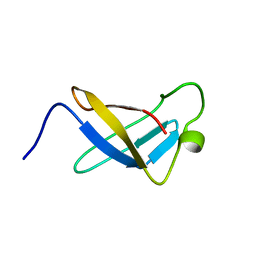 | | S1 RNA BINDING DOMAIN, NMR, 20 STRUCTURES | | Descriptor: | PNPASE | | Authors: | Bycroft, M. | | Deposit date: | 1996-11-27 | | Release date: | 1997-04-01 | | Last modified: | 2024-05-22 | | Method: | SOLUTION NMR | | Cite: | The solution structure of the S1 RNA binding domain: a member of an ancient nucleic acid-binding fold.
Cell(Cambridge,Mass.), 88, 1997
|
|
1LUZ
 
 | |
2CQO
 
 | | Solution structure of the S1 RNA binding domain of human hypothetical protein FLJ11067 | | Descriptor: | Nucleolar protein of 40 kDa | | Authors: | Suzuki, S, Muto, Y, Inoue, M, Kigawa, T, Terada, T, Shirouzu, M, Yokoyama, S, RIKEN Structural Genomics/Proteomics Initiative (RSGI) | | Deposit date: | 2005-05-20 | | Release date: | 2005-11-20 | | Last modified: | 2024-05-29 | | Method: | SOLUTION NMR | | Cite: | Solution structure of the S1 RNA binding domain of human hypothetical protein FLJ11067
To be Published
|
|
2EQS
 
 | | Solution structure of the S1 RNA binding domain of human ATP-dependent RNA helicase DHX8 | | Descriptor: | ATP-dependent RNA helicase DHX8 | | Authors: | Suzuki, S, Muto, Y, Inoue, M, Kigawa, T, Terada, T, Shirouzu, M, Yokoyama, S, RIKEN Structural Genomics/Proteomics Initiative (RSGI) | | Deposit date: | 2007-03-30 | | Release date: | 2007-10-02 | | Last modified: | 2024-05-29 | | Method: | SOLUTION NMR | | Cite: | Solution structure of the S1 RNA binding domain of human ATP-dependent RNA helicase DHX8
To be Published
|
|
4NNG
 
 | |
4NNH
 
 | |
4NNK
 
 | |
4NNI
 
 | | Structural basis for targeting the ribosomal protein S1 of Mycobacterium tuberculosis by pyrazinamide | | Descriptor: | 30S ribosomal protein S1, PYRAZINE-2-CARBOXYLIC ACID | | Authors: | Yang, J, Liu, Y, Cai, Q, Lin, D. | | Deposit date: | 2013-11-18 | | Release date: | 2014-12-24 | | Last modified: | 2024-03-20 | | Method: | X-RAY DIFFRACTION (2.64 Å) | | Cite: | Structural basis for targeting the ribosomal protein S1 of Mycobacterium tuberculosis by pyrazinamide.
Mol.Microbiol., 95, 2015
|
|
2KHI
 
 | | NMR structure of the domain 4 of the E. coli ribosomal protein S1 | | Descriptor: | 30S ribosomal protein S1 | | Authors: | Salah, P, Bisaglia, M, Aliprandi, P, Uzan, M, Sizun, C, Bontems, F. | | Deposit date: | 2009-04-07 | | Release date: | 2009-10-20 | | Last modified: | 2024-05-29 | | Method: | SOLUTION NMR | | Cite: | Probing the relationship between Gram-negative and Gram-positive S1 proteins by sequence analysis
Nucleic Acids Res., 37, 2009
|
|
2KHJ
 
 | | NMR structure of the domain 6 of the E. coli ribosomal protein S1 | | Descriptor: | 30S ribosomal protein S1 | | Authors: | Salah, P, Bisaglia, M, Aliprandi, P, Uzan, M, Sizun, C, Bontems, F. | | Deposit date: | 2009-04-07 | | Release date: | 2009-10-20 | | Last modified: | 2024-05-29 | | Method: | SOLUTION NMR | | Cite: | Probing the relationship between Gram-negative and Gram-positive S1 proteins by sequence analysis
Nucleic Acids Res., 37, 2009
|
|
2K52
 
 | | Structure of uncharacterized protein MJ1198 from Methanocaldococcus jannaschii. Northeast Structural Genomics Target MjR117B | | Descriptor: | Uncharacterized protein MJ1198 | | Authors: | Rossi, P, Maglaqui, M, Foote, E.L, Hamilton, K, Ciccosanti, C, Xiao, R, Nair, R, Swapna, G, Everett, J.K, Acton, T.B, Rost, B, Montelione, G.T, Northeast Structural Genomics Consortium (NESG) | | Deposit date: | 2008-06-24 | | Release date: | 2008-07-15 | | Last modified: | 2024-05-01 | | Method: | SOLUTION NMR | | Cite: | Structure of uncharacterized protein MJ1198 from
Methanocaldococcus jannaschii. Northeast Structural Genomics Target MjR117B
To be Published
|
|
2K4K
 
 | | Solution structure of GSP13 from Bacillus subtilis | | Descriptor: | General stress protein 13 | | Authors: | Yu, W, Yu, B, Hu, J, Jin, C, Xia, B. | | Deposit date: | 2008-06-13 | | Release date: | 2009-05-12 | | Last modified: | 2024-05-29 | | Method: | SOLUTION NMR | | Cite: | Solution structure of GSP13 from Bacillus subtilis exhibits an S1 domain related to cold shock proteins.
J.Biomol.Nmr, 43, 2009
|
|
5IE8
 
 | |
5XQ5
 
 | |
7A05
 
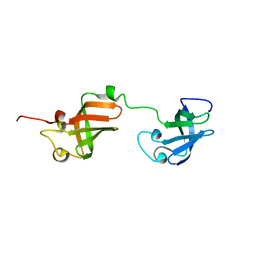 | | NMR structure of D3-D4 domains of Vibrio vulnificus ribosomal protein S1 | | Descriptor: | 30S ribosomal protein S1 | | Authors: | Qureshi, N.S, Matzel, T, Cetiner, E.C, Schnieders, S, Jonker, H.R.A, Schwalbe, H, Fuertig, B. | | Deposit date: | 2020-08-06 | | Release date: | 2021-06-23 | | Last modified: | 2024-01-17 | | Method: | SOLUTION NMR | | Cite: | NMR structure of the Vibrio vulnificus ribosomal protein S1 domains D3 and D4 provides insights into molecular recognition of single-stranded RNAs.
Nucleic Acids Res., 49, 2021
|
|
1GO3
 | | Structure of an archeal homolog of the eukaryotic RNA polymerase II RPB4/RPB7 complex | | Descriptor: | DNA-DIRECTED RNA POLYMERASE SUBUNIT E, DNA-DIRECTED RNA POLYMERASE SUBUNIT F | | Authors: | Todone, F, Brick, P, Werner, F, Weinzierl, R.O.J, Onesti, S. | | Deposit date: | 2001-10-17 | | Release date: | 2001-12-07 | | Last modified: | 2024-05-08 | | Method: | X-RAY DIFFRACTION (1.75 Å) | | Cite: | Structure of an Archaeal Homolog of the Eukaryotic RNA Polymerase II Rpb4/Rpb7 Complex
Mol.Cell, 8, 2001
|
|
1HH2
 
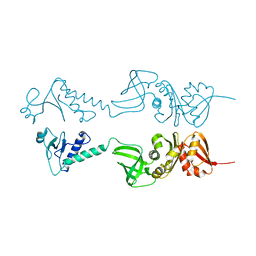 | | Crystal structure of NusA from Thermotoga maritima | | Descriptor: | N UTILIZATION SUBSTANCE PROTEIN A | | Authors: | Worbs, M, Bourenkov, G.P, Bartunik, H.D, Huber, R, Wahl, M.C. | | Deposit date: | 2000-12-18 | | Release date: | 2001-10-19 | | Last modified: | 2024-05-08 | | Method: | X-RAY DIFFRACTION (2.1 Å) | | Cite: | An Extended RNA Binding Surface Through Arrayed S1 and Kh Domains in Transcription Factor Nusa
Mol.Cell, 7, 2001
|
|
1L2F
 
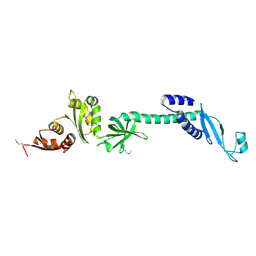 | | Crystal structure of NusA from Thermotoga maritima: a structure-based role of the N-terminal domain | | Descriptor: | N utilization substance protein A | | Authors: | Shin, D.H, Nguyen, H.H, Jancarik, J, Yokota, H, Kim, R, Kim, S.H, Berkeley Structural Genomics Center (BSGC) | | Deposit date: | 2002-02-20 | | Release date: | 2003-09-23 | | Last modified: | 2024-10-30 | | Method: | X-RAY DIFFRACTION (2.5 Å) | | Cite: | Crystal structure of NusA from Thermotoga maritima and functional implication of the N-terminal domain.
Biochemistry, 42, 2003
|
|
1KL9
 
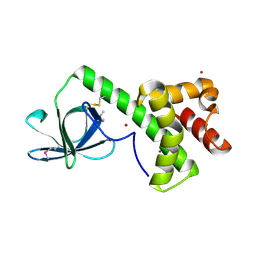 | |
4QJF
 
 | |
1Q8K
 
 | |
2OCE
 
 | |
