9S8V
 
 | |
9QAS
 
 | | Human angiotensin-1 converting enzyme N-domain in complex with trandolaprilat | | Descriptor: | 1,2-ETHANEDIOL, 2-acetamido-2-deoxy-beta-D-glucopyranose-(1-4)-2-acetamido-2-deoxy-beta-D-glucopyranose, 2-acetamido-2-deoxy-beta-D-glucopyranose-(1-4)-[alpha-L-fucopyranose-(1-6)]2-acetamido-2-deoxy-beta-D-glucopyranose, ... | | Authors: | Gregory, K.S, Acharya, K.R. | | Deposit date: | 2025-02-28 | | Release date: | 2025-09-03 | | Method: | X-RAY DIFFRACTION (2.2 Å) | | Cite: | Molecular basis of domain-specific angiotensin I-converting enzyme inhibition by the antihypertensive drugs enalaprilat, ramiprilat, trandolaprilat, quinaprilat and perindoprilat.
Febs J., 2025
|
|
6HIJ
 
 | | Cryo-EM structure of the human ABCG2-MZ29-Fab complex with cholesterol and PE lipids docked | | Descriptor: | 1,2-dioleoyl-sn-glycero-3-phosphoethanolamine, ATP-binding cassette sub-family G member 2, CHOLESTEROL, ... | | Authors: | Jackson, S.M, Manolaridis, I, Kowal, J, Zechner, M, Taylor, N.M.I, Bause, M, Bauer, S, Bartholomaeus, R, Stahlberg, H, Bernhardt, G, Koenig, B, Buschauer, A, Altmann, K.H, Locher, K.P. | | Deposit date: | 2018-08-30 | | Release date: | 2018-09-19 | | Last modified: | 2025-10-01 | | Method: | ELECTRON MICROSCOPY (3.56 Å) | | Cite: | Structural basis of small-molecule inhibition of human multidrug transporter ABCG2.
Nat.Struct.Mol.Biol., 25, 2018
|
|
8TP4
 
 | | H2 hemagglutinin (A/Singapore/1/1957) in complex with RBS-targeting Fab 1-1-1E04 | | Descriptor: | 2-acetamido-2-deoxy-beta-D-glucopyranose, 2-acetamido-2-deoxy-beta-D-glucopyranose-(1-4)-2-acetamido-2-deoxy-beta-D-glucopyranose, Heavy chain of Fab 1-1-1E04, ... | | Authors: | Yang, Y.R, Han, J, Perrett, H.R, Ward, A.B. | | Deposit date: | 2023-08-04 | | Release date: | 2023-10-11 | | Last modified: | 2025-06-04 | | Method: | ELECTRON MICROSCOPY (3.3 Å) | | Cite: | Immune memory shapes human polyclonal antibody responses to H2N2 vaccination.
Cell Rep, 43, 2024
|
|
9US4
 
 | | Klebsiella pneumoniae maltohexaose-producing alpha-amylase in complex with acarbose | | Descriptor: | 4,6-dideoxy-4-{[(1S,4R,5S,6S)-4,5,6-trihydroxy-3-(hydroxymethyl)cyclohex-2-en-1-yl]amino}-alpha-D-glucopyranose-(1-4)-alpha-D-glucopyranose, 4,6-dideoxy-4-{[(1S,4R,5S,6S)-4,5,6-trihydroxy-3-(hydroxymethyl)cyclohex-2-en-1-yl]amino}-alpha-D-glucopyranose-(1-4)-alpha-D-glucopyranose-(1-4)-alpha-D-glucopyranose, 4,6-dideoxy-4-{[(1S,4R,5S,6S)-4,5,6-trihydroxy-3-(hydroxymethyl)cyclohex-2-en-1-yl]amino}-alpha-D-glucopyranose-(1-4)-alpha-D-glucopyranose-(1-4)-beta-D-glucopyranose, ... | | Authors: | Fujimoto, Z, Kishine, N, Momma, M. | | Deposit date: | 2025-05-01 | | Release date: | 2025-07-09 | | Last modified: | 2025-09-03 | | Method: | X-RAY DIFFRACTION (1.95 Å) | | Cite: | Crystal structure of Klebsiella pneumoniae maltohexaose-producing alpha-amylase.
J.Biochem., 178, 2025
|
|
9QAR
 
 | | Human angiotensin-1 converting enzyme N-domain in complex with ramiprilat | | Descriptor: | 1,2-ETHANEDIOL, 2-acetamido-2-deoxy-beta-D-glucopyranose, 2-acetamido-2-deoxy-beta-D-glucopyranose-(1-4)-2-acetamido-2-deoxy-beta-D-glucopyranose, ... | | Authors: | Gregory, K.S, Acharya, K.R. | | Deposit date: | 2025-02-28 | | Release date: | 2025-09-03 | | Method: | X-RAY DIFFRACTION (2 Å) | | Cite: | Molecular basis of domain-specific angiotensin I-converting enzyme inhibition by the antihypertensive drugs enalaprilat, ramiprilat, trandolaprilat, quinaprilat and perindoprilat.
Febs J., 2025
|
|
3HT5
 
 | |
8WTQ
 
 | | HUMAN SQUALENE SYNTHASE IN COMPLEX WITH {1-[2-Chloro-5-(2,2-dimethyl-propyl)-13-(2-methoxy-phenyl)-6-oxo-6,7,10,11-tetrahydro-5H,9H,13H-12-oxa-5,8-diaza-benzocycloundecene-8-carbonyl]-piperidin-4-yl}-acetic acid | | Descriptor: | 2-[1-[[(10~{S})-13-chloranyl-2-(2,2-dimethylpropyl)-10-(2-methoxyphenyl)-3-oxidanylidene-9-oxa-2,5-diazabicyclo[9.4.0]pentadeca-1(15),11,13-trien-5-yl]carbonyl]piperidin-4-yl]ethanoic acid, PHOSPHATE ION, Squalene synthase | | Authors: | Suzuki, M, Haginoya, N, Suzuki, M, Ishigai, Y, Terayama, K, Kanda, A, Sugita, K. | | Deposit date: | 2023-10-19 | | Release date: | 2024-05-22 | | Method: | X-RAY DIFFRACTION (2 Å) | | Cite: | Discovery of Novel 11-Membered Templates as Squalene Synthase Inhibitors.
J.Med.Chem., 67, 2024
|
|
6EDM
 
 | | Structure of apo-CDD-1 beta-lactamase | | Descriptor: | Beta-lactamase, SULFATE ION | | Authors: | Smith, C.A, Vakulenko, S.B. | | Deposit date: | 2018-08-09 | | Release date: | 2019-08-14 | | Last modified: | 2025-04-02 | | Method: | X-RAY DIFFRACTION (1.4 Å) | | Cite: | The crystal structures of CDD-1, the intrinsic class D beta-lactamase from the pathogenic Gram-positive bacterium Clostridioides difficile, and its complex with cefotaxime.
J.Struct.Biol., 208, 2019
|
|
5FWA
 
 | | Crystal Structure of Mus musculus Protein Arginine Methyltransferase 2 with CP1 | | Descriptor: | 1,2-ETHANEDIOL, 9-(6-carbamimidamido-5,6-dideoxy-beta-D-ribo-hexofuranosyl)-9H-purin-6-amine, CALCIUM ION, ... | | Authors: | Cura, V, Troffer-Charlier, N, Marechal, N, Bonnefond, L, Cavarelli, J. | | Deposit date: | 2016-02-16 | | Release date: | 2016-11-09 | | Last modified: | 2024-01-10 | | Method: | X-RAY DIFFRACTION (1.8 Å) | | Cite: | Structural studies of protein arginine methyltransferase 2 reveal its interactions with potential substrates and inhibitors.
FEBS J., 284, 2017
|
|
9MIB
 
 | | 206-9C09 Fab in complex with HIV-1 GT1.1 v4.1 SOSIP Env trimer and RM20A3 Fab | | Descriptor: | 2-acetamido-2-deoxy-beta-D-glucopyranose, 2-acetamido-2-deoxy-beta-D-glucopyranose-(1-4)-2-acetamido-2-deoxy-beta-D-glucopyranose, 206-9C09 heavy chain Fv, ... | | Authors: | Phulera, S, Ozorowski, G, Ward, A.B. | | Deposit date: | 2024-12-12 | | Release date: | 2025-05-21 | | Last modified: | 2025-08-13 | | Method: | ELECTRON MICROSCOPY (2.8 Å) | | Cite: | Precise targeting of HIV broadly neutralizing antibody precursors in humans.
Science, 389, 2025
|
|
5J1G
 | |
9MII
 
 | | 253-7A03 Fab in complex with HIV-1 BG505 SOSIP Env trimer and RM20A3 Fab | | Descriptor: | 2-acetamido-2-deoxy-beta-D-glucopyranose, 2-acetamido-2-deoxy-beta-D-glucopyranose-(1-4)-2-acetamido-2-deoxy-beta-D-glucopyranose, 253-7A03 heavy chain Fv, ... | | Authors: | Phulera, S, Ozorowski, G, Ward, A.B. | | Deposit date: | 2024-12-12 | | Release date: | 2025-05-21 | | Last modified: | 2025-08-13 | | Method: | ELECTRON MICROSCOPY (3.3 Å) | | Cite: | Precise targeting of HIV broadly neutralizing antibody precursors in humans.
Science, 389, 2025
|
|
8HU2
 
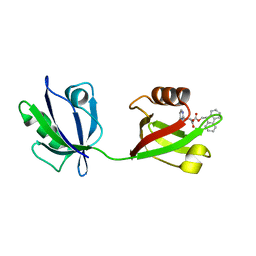 | | Rattus Syntenin-1 PDZ domain with inhibitor | | Descriptor: | (2~{S})-2-(9~{H}-fluoren-9-ylmethoxycarbonylamino)-3-(4-oxidanylidene-5~{H}-pyrimidin-2-yl)propanoic acid, Syntenin-1 | | Authors: | Heo, Y, Lee, J, Yun, J.H, Lee, W. | | Deposit date: | 2022-12-22 | | Release date: | 2024-01-17 | | Method: | X-RAY DIFFRACTION (1.6 Å) | | Cite: | Crystal structure of STNPDZ with inhibitor at 1.60 Angstroms resolution.
To Be Published
|
|
6TG0
 
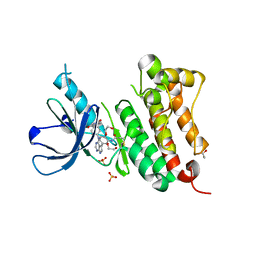 | | Crystal Structure of EGFR T790M/V948R in Complex with Covalent Pyrrolopyrimidine 21a | | Descriptor: | 1,2-ETHANEDIOL, Epidermal growth factor receptor, SULFATE ION, ... | | Authors: | Niggenaber, J, Mueller, M.P, Rauh, D. | | Deposit date: | 2019-11-14 | | Release date: | 2020-09-30 | | Last modified: | 2024-11-20 | | Method: | X-RAY DIFFRACTION (1.5 Å) | | Cite: | Targeting Her2-insYVMA with Covalent Inhibitors-A Focused Compound Screening and Structure-Based Design Approach.
J.Med.Chem., 63, 2020
|
|
9GGQ
 
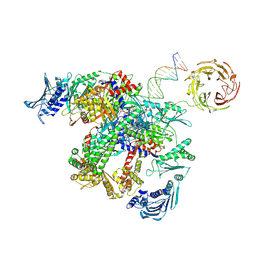 | | E.coli gyrase holocomplex with cleaved chirally wrapped 217 bp DNA fragment and moxifloxacin | | Descriptor: | 1-cyclopropyl-6-fluoro-8-methoxy-7-[(4aS,7aS)-octahydro-6H-pyrrolo[3,4-b]pyridin-6-yl]-4-oxo-1,4-dihydroquinoline-3-carboxylic acid, DNA gyrase subunit A, DNA gyrase subunit B, ... | | Authors: | Ghilarov, D, Heddle, J.G, Pabis, M. | | Deposit date: | 2024-08-13 | | Release date: | 2024-09-11 | | Last modified: | 2025-07-02 | | Method: | ELECTRON MICROSCOPY (2.6 Å) | | Cite: | Structural basis of chiral wrap and T-segment capture by Escherichia coli DNA gyrase.
Proc.Natl.Acad.Sci.USA, 121, 2024
|
|
9DWM
 
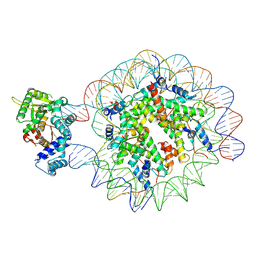 | |
7WBO
 
 | | Crystal structure of Sarcoplasmic Calcium-Binding Protein from Scylla paramamosain | | Descriptor: | 1,2-ETHANEDIOL, CALCIUM ION, SODIUM ION, ... | | Authors: | Chen, Y, Jin, T, Liu, G. | | Deposit date: | 2021-12-17 | | Release date: | 2022-12-21 | | Last modified: | 2023-11-29 | | Method: | X-RAY DIFFRACTION (1.6 Å) | | Cite: | Crystal Structure Analysis of Sarcoplasmic-Calcium-Binding Protein: An Allergen in Scylla paramamosain.
J.Agric.Food Chem., 71, 2023
|
|
6EQ4
 | | MTH1 in complex with fragment 8 | | Descriptor: | 4-(3-fluoranylpyridin-4-yl)-1~{H}-pyrrolo[2,3-b]pyridine, 7,8-dihydro-8-oxoguanine triphosphatase, SULFATE ION | | Authors: | Wiedmer, L, Sledz, P, Caflisch, A. | | Deposit date: | 2017-10-12 | | Release date: | 2018-10-31 | | Last modified: | 2024-05-08 | | Method: | X-RAY DIFFRACTION (1.4 Å) | | Cite: | Ligand retargeting by binding site analogy.
Eur.J.Med.Chem., 175, 2019
|
|
8CE3
 
 | | Crystal structure of MGAT5 (alpha-1,6-mannosylglycoprotein 6-beta-N-acetylglucosaminyltransferase V) luminal domain with a Lys329-Ile345 loop truncation, in complex with 3D fragment 2548 | | Descriptor: | (3~{S})-1-ethanoyl-3-(4-methylphenyl)piperidine-3-carboxylic acid, 1,2-ETHANEDIOL, Alpha-1,6-mannosylglycoprotein 6-beta-N-acetylglucosaminyltransferase A, ... | | Authors: | Gilio, A.K, Darby, J.F, Wu, L, Davies, G.J. | | Deposit date: | 2023-02-01 | | Release date: | 2024-02-21 | | Last modified: | 2025-09-03 | | Method: | X-RAY DIFFRACTION (1.89 Å) | | Cite: | Design, Modular Synthesis and Screening of 58 Shape-Diverse 3-D Fragments
Chemrxiv, 2025
|
|
9DWF
 
 | |
6I12
 
 | | CRYSTAL STRUCTURE OF FASCIN IN COMPLEX WITH COMPOUND 5 | | Descriptor: | 2-[(4-chlorophenyl)methyl]-~{N}-(1-methylpyrazol-4-yl)-1-oxidanylidene-isoquinoline-4-carboxamide, ACETATE ION, Fascin | | Authors: | Schuettelkopf, A.W. | | Deposit date: | 2018-10-27 | | Release date: | 2019-02-27 | | Last modified: | 2024-01-24 | | Method: | X-RAY DIFFRACTION (1.65 Å) | | Cite: | Structure-based design, synthesis and biological evaluation of a novel series of isoquinolone and pyrazolo[4,3-c]pyridine inhibitors of fascin 1 as potential anti-metastatic agents.
Bioorg.Med.Chem.Lett., 29, 2019
|
|
7QOA
 
 | | Structure of CodB, a cytosine transporter in an outward-facing conformation | | Descriptor: | 2,3-dihydroxypropyl (9Z)-octadec-9-enoate, 6-AMINOPYRIMIDIN-2(1H)-ONE, Cytosine permease, ... | | Authors: | Hatton, C.E, Cameron, A.D. | | Deposit date: | 2021-12-23 | | Release date: | 2022-07-13 | | Last modified: | 2024-01-31 | | Method: | X-RAY DIFFRACTION (2.4 Å) | | Cite: | Structure of cytosine transport protein CodB provides insight into nucleobase-cation symporter 1 mechanism.
Embo J., 41, 2022
|
|
5CQR
 
 | | Dimerization of Elp1 is essential for Elongator complex assembly | | Descriptor: | Elongator complex protein 1 | | Authors: | Lin, Z, Xu, H, Li, F, Diao, W, Long, J, Shen, Y. | | Deposit date: | 2015-07-22 | | Release date: | 2015-08-19 | | Last modified: | 2024-03-20 | | Method: | X-RAY DIFFRACTION (3.015 Å) | | Cite: | Dimerization of elongator protein 1 is essential for Elongator complex assembly.
Proc.Natl.Acad.Sci.USA, 112, 2015
|
|
9DWH
 
 | | DNA Polymerase Beta bound to a nucleosome containing a 1-nt gap at SHL-4.5 (State 2, composite) | | Descriptor: | 601 I strand (damaged strand 1), 601 J strand (non-damaged strand), 601 K strand (damaged strand 2), ... | | Authors: | Weaver, T.M, Ryan, B.J, Freudenthal, B.D. | | Deposit date: | 2024-10-09 | | Release date: | 2025-01-15 | | Method: | ELECTRON MICROSCOPY (3.3 Å) | | Cite: | DNA Polymerase Beta bound to a nucleosome containing a 1-nt gap at SHL-4.5 (State 2, composite)
To Be Published
|
|
