1Y6P
 
 | | Crystal structure of disulfide engineered porcine pancratic phospholipase a2 to group-x isozyme | | Descriptor: | CALCIUM ION, CHLORIDE ION, Phospholipase A2, ... | | Authors: | Yu, B.Z, Pan, Y.H, Jassen, M.J.W, Bahnson, B.J, Jain, M.K. | | Deposit date: | 2004-12-06 | | Release date: | 2005-03-22 | | Last modified: | 2024-11-13 | | Method: | X-RAY DIFFRACTION (2.25 Å) | | Cite: | Kinetic and structural properties of disulfide engineered phospholipase a(2): insight into the role of disulfide bonding patterns.
Biochemistry, 44, 2005
|
|
2QEV
 
 | | Crystal Structure of Anopheles gambiae D7r4 | | Descriptor: | D7R4 Protein | | Authors: | Andersen, J.F, Mans, B.J, Calvo, E, Ribeiro, J.M. | | Deposit date: | 2007-06-26 | | Release date: | 2007-10-09 | | Last modified: | 2024-10-30 | | Method: | X-RAY DIFFRACTION (1.998 Å) | | Cite: | The Crystal Structure of D7r4, a Salivary Biogenic Amine-binding Protein from the Malaria Mosquito Anopheles gambiae
J.Biol.Chem., 282, 2007
|
|
2A85
 
 | | Crystal Structure of the G81A mutant of the Active Chimera of (S)-Mandelate Dehydrogenase in complex with its substrate 2-hydroxyoctanoate | | Descriptor: | (2S)-2-HYDROXYOCTANOIC ACID, 2-(N-MORPHOLINO)-ETHANESULFONIC ACID, FLAVIN MONONUCLEOTIDE, ... | | Authors: | Sukumar, N, Xu, Y, Mitra, B, Mathews, F.S. | | Deposit date: | 2005-07-07 | | Release date: | 2006-07-11 | | Last modified: | 2023-08-23 | | Method: | X-RAY DIFFRACTION (2.5 Å) | | Cite: | Structures of the G81A mutant form of the active chimera of (S)-mandelate dehydrogenase and its complex with two of its substrates.
Acta Crystallogr.,Sect.D, 65, 2009
|
|
1EL5
 
 | | COMPLEX OF MONOMERIC SARCOSINE OXIDASE WITH THE INHIBITOR DIMETHYLGLYCINE | | Descriptor: | CHLORIDE ION, FLAVIN-ADENINE DINUCLEOTIDE, N,N-DIMETHYLGLYCINE, ... | | Authors: | Wagner, M.A, Trickey, P, Chen, Z.-W, Mathews, F.S, Jorns, M.S. | | Deposit date: | 2000-03-13 | | Release date: | 2000-12-06 | | Last modified: | 2025-03-26 | | Method: | X-RAY DIFFRACTION (1.8 Å) | | Cite: | Monomeric sarcosine oxidase: 1. Flavin reactivity and active site binding determinants.
Biochemistry, 39, 2000
|
|
3WSY
 
 | | SorLA Vps10p domain in complex with its own propeptide fragment | | Descriptor: | 2-acetamido-2-deoxy-beta-D-glucopyranose, Sortilin-related receptor, peptide from Sortilin-related receptor | | Authors: | Kitago, Y, Nakata, Z, Nagae, M, Nogi, T, Takagi, J. | | Deposit date: | 2014-03-30 | | Release date: | 2015-02-04 | | Last modified: | 2024-10-30 | | Method: | X-RAY DIFFRACTION (3.11 Å) | | Cite: | Structural basis for amyloidogenic peptide recognition by sorLA.
Nat.Struct.Mol.Biol., 22, 2015
|
|
3ZGZ
 
 | | Ternary complex of E. coli leucyl-tRNA synthetase, tRNA(leu) and toxic moiety from agrocin 84 (TM84) in aminoacylation-like conformation | | Descriptor: | LEUCINE--TRNA LIGASE, MAGNESIUM ION, TRNA-LEU UAA ISOACCEPTOR, ... | | Authors: | Chopra, S, Palencia, A, Virus, C, Tripathy, A, Temple, B.R, Velazquez-Campoy, A, Cusack, S, Reader, J.S. | | Deposit date: | 2012-12-19 | | Release date: | 2013-01-30 | | Last modified: | 2023-12-20 | | Method: | X-RAY DIFFRACTION (2.4 Å) | | Cite: | Plant Tumour Biocontrol Agent Employs a tRNA-Dependent Mechanism to Inhibit Leucyl-tRNA Synthetase
Nat.Commun., 4, 2013
|
|
6BI7
 
 | | Crystal structure of Rev7-WT/Rev3 as a monomer under high-salt conditions | | Descriptor: | DNA polymerase zeta catalytic subunit, Mitotic spindle assembly checkpoint protein MAD2B | | Authors: | Rizzo, A.A, Korzhnev, D.M, Hao, B, Li, Y. | | Deposit date: | 2017-11-01 | | Release date: | 2018-08-01 | | Last modified: | 2023-10-04 | | Method: | X-RAY DIFFRACTION (2.8 Å) | | Cite: | Rev7 dimerization is important for assembly and function of the Rev1/Pol zeta translesion synthesis complex.
Proc. Natl. Acad. Sci. U.S.A., 115, 2018
|
|
1P0E
 
 | | CRYSTAL STRUCTURE OF ZYMOMONAS MOBILIS tRNA-GUANINE TRANSGLYCOSYLASE (TGT) COCRYSTALLISED WITH PREQ1 AT PH 5.5 | | Descriptor: | 7-DEAZA-7-AMINOMETHYL-GUANINE, Queuine tRNA-ribosyltransferase, ZINC ION | | Authors: | Brenk, R, Stubbs, M.T, Heine, A, Reuter, K, Klebe, G. | | Deposit date: | 2003-04-10 | | Release date: | 2003-09-30 | | Last modified: | 2023-08-16 | | Method: | X-RAY DIFFRACTION (2.4 Å) | | Cite: | Flexible adaptations in the structure of the tRNA-modifying enzyme tRNA-guanine transglycosylase
and their implications for substrate selectivity, reaction mechanism and structure-based drug design
Chembiochem, 4, 2003
|
|
7F6Q
 
 | | Crystal structure of metal-citrate-binding mutant (S79A) protein (MctA) of ABC transporter in apo state | | Descriptor: | 1,2-ETHANEDIOL, 1,3-PROPANDIOL, ACETATE ION, ... | | Authors: | Kanaujia, S.P, Mandal, S.K. | | Deposit date: | 2021-06-25 | | Release date: | 2021-12-01 | | Last modified: | 2023-11-29 | | Method: | X-RAY DIFFRACTION (1.63 Å) | | Cite: | Structural and thermodynamic insights into a novel Mg 2+ -citrate-binding protein from the ABC transporter superfamily.
Acta Crystallogr D Struct Biol, 77, 2021
|
|
7F6S
 
 | |
7F6T
 
 | |
7F6F
 
 | | Crystal structure of metal-citrate-binding protein (MctA) of ABC transporter endogenously bound to Mg2+-citrate complex (Form II) | | Descriptor: | 1,2-ETHANEDIOL, CARBON DIOXIDE, CITRIC ACID, ... | | Authors: | Kanaujia, S.P, Mandal, S.K, Gogoi, P. | | Deposit date: | 2021-06-25 | | Release date: | 2021-12-01 | | Last modified: | 2023-11-29 | | Method: | X-RAY DIFFRACTION (2.1 Å) | | Cite: | Structural and thermodynamic insights into a novel Mg 2+ -citrate-binding protein from the ABC transporter superfamily.
Acta Crystallogr D Struct Biol, 77, 2021
|
|
7F6P
 
 | | Crystal structure of metal-citrate-binding mutant (D28A) protein (MctA) of ABC transporter endogenously bound to citrate | | Descriptor: | 1,2-ETHANEDIOL, 1,3-PROPANDIOL, CARBON DIOXIDE, ... | | Authors: | Kanaujia, S.P, Mandal, S.K. | | Deposit date: | 2021-06-25 | | Release date: | 2021-12-01 | | Last modified: | 2023-11-29 | | Method: | X-RAY DIFFRACTION (2.15 Å) | | Cite: | Structural and thermodynamic insights into a novel Mg 2+ -citrate-binding protein from the ABC transporter superfamily.
Acta Crystallogr D Struct Biol, 77, 2021
|
|
7F6R
 
 | |
7F6E
 
 | | Crystal structure of metal-citrate-binding protein (MctA) of ABC transporter endogenously bound to Mg2+-citrate complex (Form I) | | Descriptor: | (2S)-2-HYDROXYPROPANOIC ACID, 1,2-ETHANEDIOL, CARBON DIOXIDE, ... | | Authors: | Kanaujia, S.P, Mandal, S.K, Gogoi, P. | | Deposit date: | 2021-06-25 | | Release date: | 2021-12-01 | | Last modified: | 2023-11-29 | | Method: | X-RAY DIFFRACTION (1.77 Å) | | Cite: | Structural and thermodynamic insights into a novel Mg 2+ -citrate-binding protein from the ABC transporter superfamily.
Acta Crystallogr D Struct Biol, 77, 2021
|
|
7F6O
 
 | |
7F6K
 
 | | Crystal structure of metal-citrate-binding protein (MctA) of ABC transporter endogenously bound to citrate | | Descriptor: | 1,2-ETHANEDIOL, 2-(N-MORPHOLINO)-ETHANESULFONIC ACID, ACETATE ION, ... | | Authors: | Kanaujia, S.P, Mandal, S.K, Gogoi, P. | | Deposit date: | 2021-06-25 | | Release date: | 2021-12-01 | | Last modified: | 2023-11-29 | | Method: | X-RAY DIFFRACTION (2.1 Å) | | Cite: | Structural and thermodynamic insights into a novel Mg 2+ -citrate-binding protein from the ABC transporter superfamily.
Acta Crystallogr D Struct Biol, 77, 2021
|
|
7F6N
 
 | |
7F6U
 
 | | Crystal structure of metal-citrate-binding mutant (Y221A) protein (MctA) of ABC transporter in apo state | | Descriptor: | 1,2-ETHANEDIOL, 2-(hydroxymethyl)butane-1,4-diol, 2-HYDROXY BUTANE-1,4-DIOL, ... | | Authors: | Kanaujia, S.P, Mandal, S.K. | | Deposit date: | 2021-06-25 | | Release date: | 2021-12-01 | | Last modified: | 2023-11-29 | | Method: | X-RAY DIFFRACTION (1.75 Å) | | Cite: | Structural and thermodynamic insights into a novel Mg 2+ -citrate-binding protein from the ABC transporter superfamily.
Acta Crystallogr D Struct Biol, 77, 2021
|
|
1OZQ
 
 | | CRYSTAL STRUCTURE OF THE MUTATED TRNA-GUANINE TRANSGLYCOSYLASE (TGT)Y106F COMPLEXED WITH PREQ1 | | Descriptor: | 7-DEAZA-7-AMINOMETHYL-GUANINE, Queuine tRNA-ribosyltransferase, ZINC ION | | Authors: | Brenk, R, Stubbs, M.T, Heine, A, Reuter, K, Klebe, G. | | Deposit date: | 2003-04-09 | | Release date: | 2003-09-30 | | Last modified: | 2023-08-16 | | Method: | X-RAY DIFFRACTION (1.9 Å) | | Cite: | Flexible adaptations in the structure of the tRNA-modifying enzyme tRNA-guanine transglycosylase
and their implications for substrate selectivity, reaction mechanism and structure-based drug design
Chembiochem, 4, 2003
|
|
1P0B
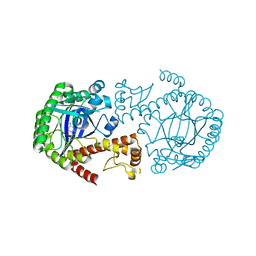 
 | | Crystal Structure Of tRNA-Guanine Transglycosylase (TGT) From Zymomonas mobilis Complexed With Archaeosine Precursor, Preq0 | | Descriptor: | 2-AMINO-4-OXO-4,7-DIHYDRO-3H-PYRROLO[2,3-D]PYRIMIDINE-5-CARBONITRILE, Queuine tRNA-ribosyltransferase, ZINC ION | | Authors: | Brenk, R, Stubbs, M.T, Heine, A, Reuter, K, Klebe, G. | | Deposit date: | 2003-04-10 | | Release date: | 2003-09-30 | | Last modified: | 2023-08-16 | | Method: | X-RAY DIFFRACTION (1.7 Å) | | Cite: | Flexible adaptations in the structure of the tRNA-modifying enzyme tRNA-guanine transglycosylase
and their implications for substrate selectivity, reaction mechanism and structure-based drug design
Chembiochem, 4, 2003
|
|
1OZM
 
 | | Y106F mutant of Z. mobilis TGT | | Descriptor: | Queuine tRNA-ribosyltransferase, ZINC ION | | Authors: | Brenk, R, Stubbs, M.T, Heine, A, Reuter, K, Klebe, G. | | Deposit date: | 2003-04-09 | | Release date: | 2003-09-30 | | Last modified: | 2023-08-16 | | Method: | X-RAY DIFFRACTION (1.95 Å) | | Cite: | Flexible adaptations in the structure of the tRNA-modifying enzyme
tRNA-guanine transglycosylase and their implications for substrate selectivity,
reaction mechanism and structure-based drug design
Chembiochem, 4, 2003
|
|
4CQN
 
 | | Crystal structure of the E.coli LeuRS-tRNA complex with the non- cognate isoleucyl adenylate analogue | | Descriptor: | ESCHERICHIA COLI TRNA-LEU UAA ISOACCEPTOR, LEUCINE--TRNA LIGASE, MAGNESIUM ION, ... | | Authors: | Palencia, A, Cusack, S, Cvetesic, N, Haslaz, I, Gruic-Sovulj, I. | | Deposit date: | 2014-02-20 | | Release date: | 2014-07-02 | | Last modified: | 2023-12-20 | | Method: | X-RAY DIFFRACTION (2.5 Å) | | Cite: | The Physiological Target for Leurs Translational Quality Control is Norvaline
Embo J., 33, 2014
|
|
2ZBG
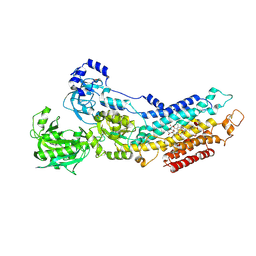 
 | | Calcium pump crystal structure with bound AlF4 and TG in the absence of calcium | | Descriptor: | MAGNESIUM ION, OCTANOIC ACID [3S-[3ALPHA, 3ABETA, ... | | Authors: | Toyoshima, C, Ogawa, H, Norimatsu, Y. | | Deposit date: | 2007-10-20 | | Release date: | 2007-12-11 | | Last modified: | 2024-10-16 | | Method: | X-RAY DIFFRACTION (2.55 Å) | | Cite: | How processing of aspartylphosphate is coupled to lumenal gating of the ion pathway in the calcium pump
Proc.Natl.Acad.Sci.Usa, 104, 2007
|
|
1Y6O
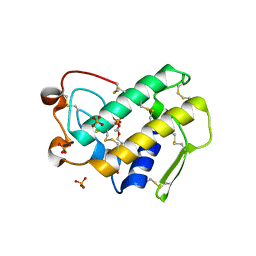 
 | | Crystal structure of disulfide engineered porcine pancreatic phospholipase A2 to group-X isozyme in complex with inhibitor MJ33 and phosphate ions | | Descriptor: | 1-HEXADECYL-3-TRIFLUOROETHYL-SN-GLYCERO-2-PHOSPHATE METHANE, CALCIUM ION, PHOSPHATE ION, ... | | Authors: | Yu, B.Z, Pan, Y.H, Jassen, M.J.W, Bahnson, B.J, Jain, M.K. | | Deposit date: | 2004-12-06 | | Release date: | 2005-03-22 | | Last modified: | 2024-10-30 | | Method: | X-RAY DIFFRACTION (2 Å) | | Cite: | Kinetic and structural properties of disulfide engineered phospholipase a(2): insight into the role of disulfide bonding patterns.
Biochemistry, 44, 2005
|
|
