2F42
 
 | |
2HDO
 
 | |
3PZC
 
 | | Crystal structure of class II aaRS homologue (Bll0957) complexed with Coenzyme A | | Descriptor: | ACETATE ION, Amino acid--[acyl-carrier-protein] ligase 1, COENZYME A, ... | | Authors: | Weygand-Durasevic, I, Luic, M, Mocibob, M, Ivic, N, Subasic, D. | | Deposit date: | 2010-12-14 | | Release date: | 2011-10-19 | | Last modified: | 2024-03-20 | | Method: | X-RAY DIFFRACTION (2.2 Å) | | Cite: | Substrate Recognition by Novel Family of Amino Acid:[Carrier Protein] Ligases
Croatica Chemica Acta, 84, 2011
|
|
4I7O
 
 | |
5RPE
 
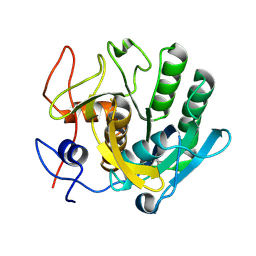 | | PanDDA analysis group deposition -- Proteinase K crystal structure Apo40 | | Descriptor: | Proteinase K | | Authors: | Lima, G.M.A, Talibov, V, Benz, L.S, Jagudin, E, Mueller, U. | | Deposit date: | 2020-09-23 | | Release date: | 2021-05-26 | | Last modified: | 2021-06-23 | | Method: | X-RAY DIFFRACTION (1.02 Å) | | Cite: | FragMAXapp: crystallographic fragment-screening data-analysis and project-management system.
Acta Crystallogr D Struct Biol, 77, 2021
|
|
3HAF
 
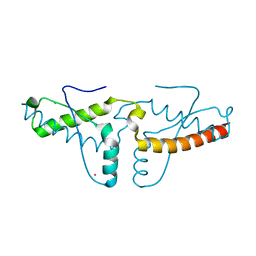 | | Human prion protein variant V129 domain swapped dimer | | Descriptor: | CADMIUM ION, CHLORIDE ION, Major prion protein | | Authors: | Lee, S, Antony, L, Hartmann, R, Knaus, K.J, Surewicz, K, Surewicz, W.K, Yee, V.C. | | Deposit date: | 2009-05-01 | | Release date: | 2010-01-12 | | Last modified: | 2017-11-01 | | Method: | X-RAY DIFFRACTION (2.26 Å) | | Cite: | Conformational diversity in prion protein variants influences intermolecular beta-sheet formation.
Embo J., 29, 2010
|
|
3HB7
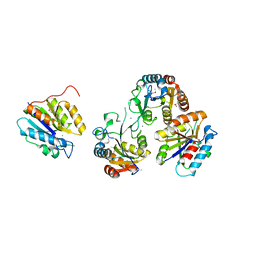 
 | | The Crystal Structure of an Isochorismatase-like Hydrolase from Alkaliphilus metalliredigens to 2.3A | | Descriptor: | AMMONIUM ION, Isochorismatase hydrolase, SODIUM ION | | Authors: | Stein, A.J, Xu, X, Cui, H, Ng, J, Edwards, A, Savchenko, A, Joachimiak, A, Midwest Center for Structural Genomics (MCSG) | | Deposit date: | 2009-05-04 | | Release date: | 2009-07-07 | | Last modified: | 2017-11-01 | | Method: | X-RAY DIFFRACTION (2.3 Å) | | Cite: | The Crystal Structure of an Isochorismatase-like Hydrolase from Alkaliphilus metalliredigens to 2.3A
To be Published
|
|
3PFT
 
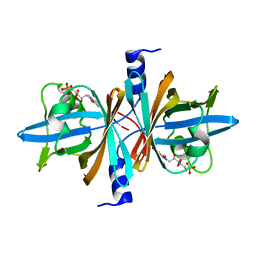 | | Crystal Structure of Untagged C54A Mutant Flavin Reductase (DszD) in Complex with FMN From Mycobacterium goodii | | Descriptor: | FLAVIN MONONUCLEOTIDE, Flavin reductase | | Authors: | Li, Q, Xu, P, Ma, C, Gu, L, Liu, X, Zhang, C, Li, N, Su, J, Li, B, Liu, S. | | Deposit date: | 2010-10-29 | | Release date: | 2011-11-02 | | Last modified: | 2024-03-20 | | Method: | X-RAY DIFFRACTION (1.601 Å) | | Cite: | The flavin reductase DSZD from a desulfurizing mycobacterium goodii strain: systemic manipulation and investigation based on the crystal structure
To be Published
|
|
4LR2
 
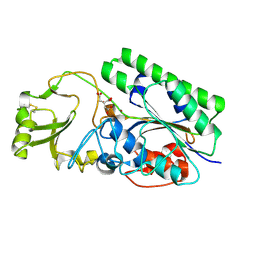 | | Crystal Structure of Human ENPP4 (apo) | | Descriptor: | 2-acetamido-2-deoxy-beta-D-glucopyranose, 2-acetamido-2-deoxy-beta-D-glucopyranose-(1-4)-2-acetamido-2-deoxy-beta-D-glucopyranose, Bis(5'-adenosyl)-triphosphatase ENPP4, ... | | Authors: | Albright, R.A, Braddock, D.T. | | Deposit date: | 2013-07-19 | | Release date: | 2013-12-18 | | Last modified: | 2023-09-20 | | Method: | X-RAY DIFFRACTION (1.5 Å) | | Cite: | Molecular basis of purinergic signal metabolism by ectonucleotide pyrophosphatase/phosphodiesterases 4 and 1 and implications in stroke.
J.Biol.Chem., 289, 2014
|
|
4LSL
 
 | | Crystal Structure of HIV-1 Reverse Transcriptase in Complex with (E)-3-(3-(4-chloro-2-(2-(2,4-dioxo-3,4-dihydropyrimidin-1(2H)-yl)ethoxy)phenoxy)phenyl)acrylonitrile (JLJ476), a non-nucleoside inhibitor | | Descriptor: | (2E)-3-(3-{4-chloro-2-[2-(2,4-dioxo-3,4-dihydropyrimidin-1(2H)-yl)ethoxy]phenoxy}phenyl)prop-2-enenitrile, HIV-1 reverse transcriptase, p51 subunit, ... | | Authors: | Frey, K.M, Anderson, K.S. | | Deposit date: | 2013-07-22 | | Release date: | 2013-12-25 | | Last modified: | 2023-09-20 | | Method: | X-RAY DIFFRACTION (2.69 Å) | | Cite: | Structure-Based Evaluation of C5 Derivatives in the Catechol Diether Series Targeting HIV-1 Reverse Transcriptase.
Chem.Biol.Drug Des., 83, 2014
|
|
3GVR
 
 | |
2QVU
 
 | | Porcine Liver Fructose-1,6-bisphosphatase cocrystallized with Fru-2,6-P2 and Mg2+, I(T)-state | | Descriptor: | 2,6-di-O-phosphono-beta-D-fructofuranose, Fructose-1,6-bisphosphatase 1, MAGNESIUM ION, ... | | Authors: | Hines, J.K, Chen, X, Nix, J.C, Fromm, H.J, Honzatko, R.B. | | Deposit date: | 2007-08-08 | | Release date: | 2007-10-23 | | Last modified: | 2023-08-30 | | Method: | X-RAY DIFFRACTION (1.5 Å) | | Cite: | Structures of mammalian and bacterial fructose-1,6-bisphosphatase reveal the basis for synergism in AMP/fructose 2,6-bisphosphate inhibition
J.Biol.Chem., 282, 2007
|
|
4HRC
 
 | | Crystal structure of yeast 20S proteasome in complex with epoxyketone carmaphycin analogue 3 | | Descriptor: | N-hexanoyl-L-valyl-N~1~-[(2R,3S,4S)-1,3-dihydroxy-2,6-dimethylheptan-4-yl]-N~5~,N~5~-dimethyl-L-glutamamide, Proteasome component C1, Proteasome component C11, ... | | Authors: | Trivella, D.B.B, Stein, M, Groll, M. | | Deposit date: | 2012-10-27 | | Release date: | 2014-01-29 | | Last modified: | 2014-07-02 | | Method: | X-RAY DIFFRACTION (2.8 Å) | | Cite: | Enzyme inhibition by hydroamination: design and mechanism of a hybrid carmaphycin-syringolin enone proteasome inhibitor.
Chem.Biol., 21, 2014
|
|
2GTX
 
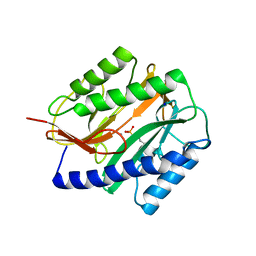 | | Structural Basis of Catalysis by Mononuclear Methionine Aminopeptidase | | Descriptor: | (1-AMINO-PENTYL)-PHOSPHONIC ACID, MANGANESE (II) ION, Methionine aminopeptidase, ... | | Authors: | Ye, Q.Z. | | Deposit date: | 2006-04-28 | | Release date: | 2006-07-04 | | Last modified: | 2023-08-30 | | Method: | X-RAY DIFFRACTION (2 Å) | | Cite: | Structural basis of catalysis by monometalated methionine aminopeptidase.
Proc.Natl.Acad.Sci.Usa, 103, 2006
|
|
4HV3
 
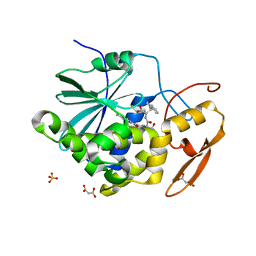 | | Structure of Ricin A chain bound with N-(N-(pterin-7-yl)carbonyl-L-serinyl)-L-tryptophan | | Descriptor: | (2S)-2-[[(2S)-2-[(2-azanyl-4-oxidanylidene-1H-pteridin-7-yl)carbonylamino]-3-oxidanyl-propanoyl]amino]-3-(1H-indol-3-yl)propanoic acid, MALONIC ACID, Ricin, ... | | Authors: | Robertus, J.D, Manzano, L.A, Jasheway, K.R, Monzingo, A.F, Saito, R, Pruet, J.M, Wiget, P.A, Anslyn, E.V. | | Deposit date: | 2012-11-05 | | Release date: | 2012-12-26 | | Last modified: | 2023-09-20 | | Method: | X-RAY DIFFRACTION (1.54 Å) | | Cite: | Peptide-conjugated pterins as inhibitors of ricin toxin A.
J.Med.Chem., 56, 2013
|
|
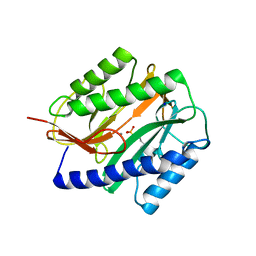
2GU4
 
 | |
3PIB
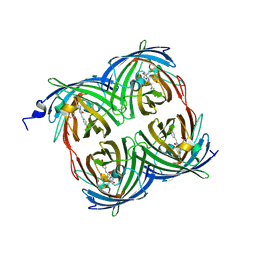 
 | |
2F9I
 
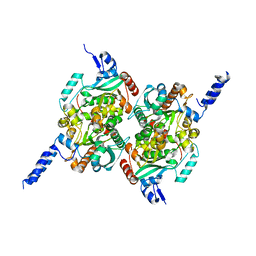 | | Crystal Structure of the carboxyltransferase subunit of ACC from Staphylococcus aureus | | Descriptor: | ZINC ION, acetyl-coenzyme A carboxylase carboxyl transferase subunit alpha, acetyl-coenzyme A carboxylase carboxyl transferase subunit beta | | Authors: | Bilder, P.W. | | Deposit date: | 2005-12-05 | | Release date: | 2006-12-05 | | Last modified: | 2023-08-30 | | Method: | X-RAY DIFFRACTION (1.98 Å) | | Cite: | The structure of the carboxyltransferase component of acetyl-coA carboxylase reveals a zinc-binding motif unique to the bacterial enzyme.
Biochemistry, 45, 2006
|
|
3PJ7
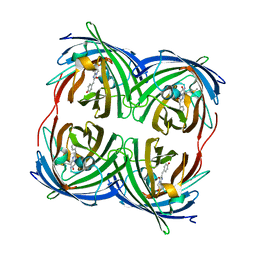 
 | |
4KPI
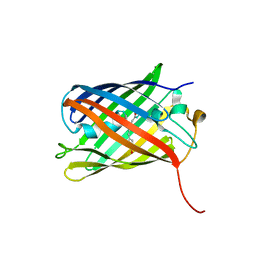 
 | |
5RTC
 
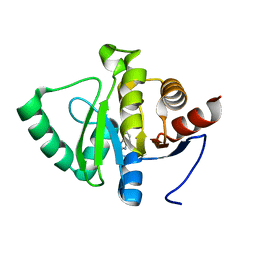 | | PanDDA analysis group deposition -- Crystal structure of SARS-CoV-2 NSP3 macrodomain in complex with ZINC000006490906 | | Descriptor: | 1H-benzimidazole-2-sulfonamide, Non-structural protein 3 | | Authors: | Correy, G.J, Young, I.D, Thompson, M.C, Fraser, J.S. | | Deposit date: | 2020-09-28 | | Release date: | 2020-12-16 | | Last modified: | 2024-05-22 | | Method: | X-RAY DIFFRACTION (1.06 Å) | | Cite: | Fragment binding to the Nsp3 macrodomain of SARS-CoV-2 identified through crystallographic screening and computational docking.
Sci Adv, 7, 2021
|
|
3GXD
 
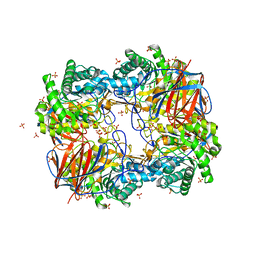 | | Crystal structure of Apo acid-beta-glucosidase pH 4.5 | | Descriptor: | 2-acetamido-2-deoxy-beta-D-glucopyranose, Glucosylceramidase, PHOSPHATE ION | | Authors: | Lieberman, R.L. | | Deposit date: | 2009-04-02 | | Release date: | 2009-05-05 | | Last modified: | 2023-09-06 | | Method: | X-RAY DIFFRACTION (2.5 Å) | | Cite: | Effects of pH and iminosugar pharmacological chaperones on lysosomal glycosidase structure and stability.
Biochemistry, 48, 2009
|
|
4KQ0
 
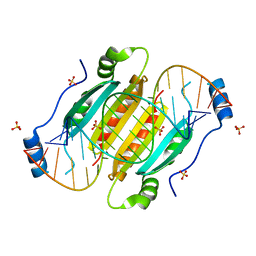 | | Crystal structure of double-helical CGG-repetitive RNA 19mer complexed with RSS p19 | | Descriptor: | 5'-R(P*GP*GP*CP*GP*GP*CP*GP*GP*CP*GP*GP*CP*GP*GP*CP*GP*GP*CP*C)-3', RNA silencing suppressor p19, SULFATE ION | | Authors: | Cabo, A, Katorcha, E, Tamjar, J, Popov, A.N, Malinina, L. | | Deposit date: | 2013-05-14 | | Release date: | 2014-05-14 | | Last modified: | 2024-02-28 | | Method: | X-RAY DIFFRACTION (2.1 Å) | | Cite: | Structural insights into CNG-repetitive RNAs associated with human Trinucleotide Repeat Expansion Diseases (TREDs)
To be Published
|
|
5RTR
 
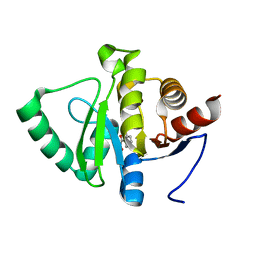 | | PanDDA analysis group deposition -- Crystal structure of SARS-CoV-2 NSP3 macrodomain in complex with ZINC000018169763 | | Descriptor: | Non-structural protein 3, SALICYLHYDROXAMIC ACID | | Authors: | Correy, G.J, Young, I.D, Thompson, M.C, Fraser, J.S. | | Deposit date: | 2020-09-28 | | Release date: | 2020-12-16 | | Last modified: | 2024-05-22 | | Method: | X-RAY DIFFRACTION (1 Å) | | Cite: | Fragment binding to the Nsp3 macrodomain of SARS-CoV-2 identified through crystallographic screening and computational docking.
Sci Adv, 7, 2021
|
|
2QY1
 
 | | pectate lyase A31G/R236F from Xanthomonas campestris | | Descriptor: | PHOSPHATE ION, Pectate lyase II | | Authors: | Garron, M.L, Shaya, D. | | Deposit date: | 2007-08-13 | | Release date: | 2008-02-26 | | Last modified: | 2021-10-20 | | Method: | X-RAY DIFFRACTION (1.9 Å) | | Cite: | Improvement of the thermostability and activity of a pectate lyase by single amino acid substitutions, using a strategy based on melting-temperature-guided sequence alignment.
Appl.Environ.Microbiol., 74, 2008
|
|
