6F1Y
 
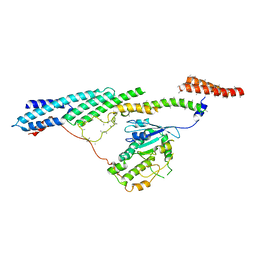 | | Dynein light intermediate chain region of the dynein tail/dynactin/BICDR1 complex | | Descriptor: | Cytoplasmic dynein 1 heavy chain 1,Dynein heavy chain, Cytoplasmic dynein 1 light intermediate chain 2 | | Authors: | Urnavicius, L, Lau, C.K, Elshenawy, M.M, Morales-Rios, E, Motz, C, Yildiz, A, Carter, A.P. | | Deposit date: | 2017-11-23 | | Release date: | 2018-01-17 | | Last modified: | 2024-05-15 | | Method: | ELECTRON MICROSCOPY (3.4 Å) | | Cite: | Cryo-EM shows how dynactin recruits two dyneins for faster movement.
Nature, 554, 2018
|
|
6F1V
 
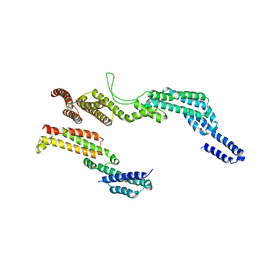 | | C terminal region of the dynein heavy chains in the dynein tail/dynactin/BICDR1 complex | | Descriptor: | Cytoplasmic dynein 1 heavy chain 1 | | Authors: | Urnavicius, L, Lau, C.K, Elshenawy, M.M, Morales-Rios, E, Motz, C, Yildiz, A, Carter, A.P. | | Deposit date: | 2017-11-23 | | Release date: | 2018-01-17 | | Last modified: | 2024-05-15 | | Method: | ELECTRON MICROSCOPY (3.4 Å) | | Cite: | Cryo-EM shows how dynactin recruits two dyneins for faster movement.
Nature, 554, 2018
|
|
4LA7
 
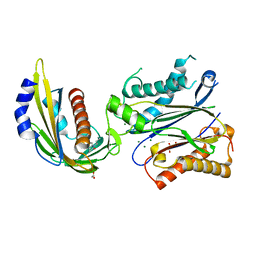 | | X-ray crystal structure of the PYL2-quinabactin-Hab1 ternary complex | | Descriptor: | ACETATE ION, Abscisic acid receptor PYL2, GLYCEROL, ... | | Authors: | Peterson, F.C, Volkman, B.F, Cutler, S.R. | | Deposit date: | 2013-06-19 | | Release date: | 2013-08-07 | | Last modified: | 2023-09-20 | | Method: | X-RAY DIFFRACTION (1.98 Å) | | Cite: | Activation of dimeric ABA receptors elicits guard cell closure, ABA-regulated gene expression, and drought tolerance.
Proc.Natl.Acad.Sci.USA, 110, 2013
|
|
6F1Z
 
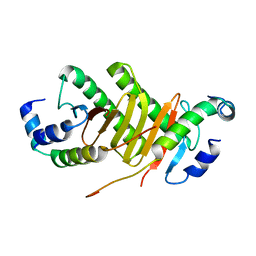 | | Roadblock-1 region of the dynein tail/dynactin/BICDR1 complex | | Descriptor: | Cytoplasmic dynein 1 intermediate chain 2, Dynein light chain roadblock-type 1 | | Authors: | Urnavicius, L, Lau, C.K, Elshenawy, M.M, Morales-Rios, E, Motz, C, Yildiz, A, Carter, A.P. | | Deposit date: | 2017-11-23 | | Release date: | 2018-01-17 | | Last modified: | 2024-05-15 | | Method: | ELECTRON MICROSCOPY (3.4 Å) | | Cite: | Cryo-EM shows how dynactin recruits two dyneins for faster movement.
Nature, 554, 2018
|
|
3MSY
 | |
1D5Q
 | | SOLUTION STRUCTURE OF A MINI-PROTEIN REPRODUCING THE CORE OF THE CD4 SURFACE INTERACTING WITH THE HIV-1 ENVELOPE GLYCOPROTEIN | | Descriptor: | CHIMERIC MINI-PROTEIN | | Authors: | Vita, C, Drakopoulou, E, Vizzanova, J, Rochette, S, Martin, L, Menez, A, Roumestand, C, Yang, Y.S, Ylisastigui, L, Benjouad, A, Gluckman, J.C. | | Deposit date: | 1999-10-11 | | Release date: | 2000-10-11 | | Last modified: | 2022-02-16 | | Method: | SOLUTION NMR | | Cite: | Rational engineering of a miniprotein that reproduces the core of the CD4 site interacting with HIV-1 envelope glycoprotein.
Proc.Natl.Acad.Sci.USA, 96, 1999
|
|
1YHT
 
 | | Crystal structure analysis of Dispersin B | | Descriptor: | ACETIC ACID, DspB, GLYCEROL | | Authors: | Ramasubbu, N, Thomas, L.M, Ragunath, C, Kaplan, J.B. | | Deposit date: | 2005-01-10 | | Release date: | 2006-01-10 | | Last modified: | 2024-02-14 | | Method: | X-RAY DIFFRACTION (2 Å) | | Cite: | Structural Analysis of Dispersin B, a Biofilm-releasing Glycoside Hydrolase from the Periodontopathogen Actinobacillus actinomycetemcomitans.
J.Mol.Biol., 349, 2005
|
|
3HWO
 
 | |
8BWC
 
 | | E. coli BAM complex (BamABCDE) wild-type | | Descriptor: | Outer membrane protein assembly factor BamA, Outer membrane protein assembly factor BamB, Outer membrane protein assembly factor BamC, ... | | Authors: | Machin, J.M, Radford, S.E, Ranson, N.A. | | Deposit date: | 2022-12-06 | | Release date: | 2023-05-24 | | Last modified: | 2023-09-06 | | Method: | ELECTRON MICROSCOPY (3.5 Å) | | Cite: | Darobactin B Stabilises a Lateral-Closed Conformation of the BAM Complex in E. coli Cells.
Angew.Chem.Int.Ed.Engl., 62, 2023
|
|
6ZP8
 
 | | Yeast 20S proteasome in complex with glidobactin-like natural product HB335 | | Descriptor: | (2~{S},3~{R})-~{N}-[(5~{S},8~{S},10~{S})-5-methyl-10-oxidanyl-2,7-bis(oxidanylidene)-1,6-diazacyclododec-8-yl]-3-oxidanyl-2-(3-phenylpropanoylamino)butanamide, CHLORIDE ION, MAGNESIUM ION, ... | | Authors: | Zhao, L, Le Chapelain, C, Brachmann, A.O, Kaiser, M, Groll, M, Bode, H.B. | | Deposit date: | 2020-07-08 | | Release date: | 2021-05-19 | | Last modified: | 2024-01-31 | | Method: | X-RAY DIFFRACTION (3 Å) | | Cite: | Activation, Structure, Biosynthesis and Bioactivity of Glidobactin-like Proteasome Inhibitors from Photorhabdus laumondii.
Chembiochem, 22, 2021
|
|
1X5R
 | |
7EXH
 
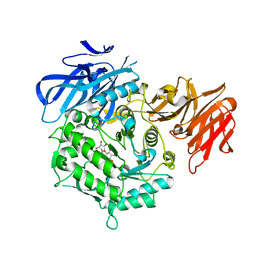 | | Crystal structure of D383A mutant from Arabidopsis thaliana complexed with Galactinol. | | Descriptor: | Probable galactinol--sucrose galactosyltransferase 6, galactinol | | Authors: | Chuankhayan, P, Guan, H.H, Lin, C.C, Chen, N.C, Huang, Y.C, Yoshimura, M, Nakagawa, A, Lee, R.H, Chen, C.J. | | Deposit date: | 2021-05-27 | | Release date: | 2022-11-30 | | Last modified: | 2023-11-29 | | Method: | X-RAY DIFFRACTION (2.63 Å) | | Cite: | Structural insight into the hydrolase and synthase activities of an alkaline alpha-galactosidase from Arabidopsis from complexes with substrate/product.
Acta Crystallogr D Struct Biol, 79, 2023
|
|
1X5O
 | | Solution structure of RRM domain in RNA binding motif, single-stranded interacting protein 1 | | Descriptor: | RNA binding motif, single-stranded interacting protein 1 | | Authors: | Dang, W, Muto, Y, Inoue, M, Kigawa, T, Shirouzu, M, Terada, T, Yokoyama, S, RIKEN Structural Genomics/Proteomics Initiative (RSGI) | | Deposit date: | 2005-05-16 | | Release date: | 2005-11-16 | | Last modified: | 2024-05-29 | | Method: | SOLUTION NMR | | Cite: | Solution structure of RRM domain in RNA binding motif, single-stranded interacting protein 1
To be Published
|
|
1X4E
 
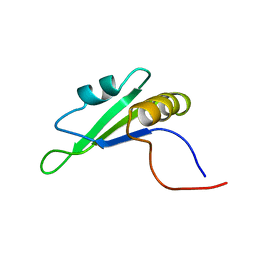 | | Solution structure of RRM domain in RNA binding motif, single-stranded interacting protein 2 | | Descriptor: | RNA binding motif, single-stranded interacting protein 2 | | Authors: | He, F, Muto, Y, Inoue, M, Kigawa, T, Shirouzu, M, Terada, T, Yokoyama, S, RIKEN Structural Genomics/Proteomics Initiative (RSGI) | | Deposit date: | 2005-05-14 | | Release date: | 2005-11-14 | | Last modified: | 2024-05-29 | | Method: | SOLUTION NMR | | Cite: | Solution structure of RRM domain in RNA binding motif, single-stranded interacting protein 2
To be Published
|
|
1CN2
 
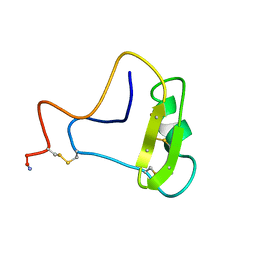 | | SOLUTION STRUCTURE OF TOXIN 2 FROM CENTRUROIDES NOXIUS HOFFMANN, A BETA SCORPION NEUROTOXIN ACTING ON SODIUM CHANNELS, NMR, 15 STRUCTURES | | Descriptor: | TOXIN 2 | | Authors: | Pintar, A, Possani, L.D, Delepierre, M. | | Deposit date: | 1998-06-21 | | Release date: | 1999-01-13 | | Last modified: | 2022-02-16 | | Method: | SOLUTION NMR | | Cite: | Solution structure of toxin 2 from centruroides noxius Hoffmann, a beta-scorpion neurotoxin acting on sodium channels.
J.Mol.Biol., 287, 1999
|
|
3GFV
 
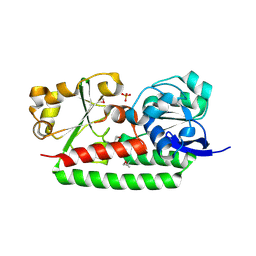 | | Crystal Structure of Petrobactin-binding Protein YclQ from Bacillu subtilis | | Descriptor: | ASPARAGINE, PHOSPHATE ION, Uncharacterized ABC transporter solute-binding protein yclQ | | Authors: | Kim, Y, Maltseva, N, Zawadzka, A.M, Raymond, K.N, Joachimiak, A, Midwest Center for Structural Genomics (MCSG) | | Deposit date: | 2009-02-27 | | Release date: | 2009-05-12 | | Last modified: | 2017-11-01 | | Method: | X-RAY DIFFRACTION (1.75 Å) | | Cite: | Characterization of a Bacillus subtilis transporter for petrobactin, an anthrax stealth siderophore.
Proc.Natl.Acad.Sci.USA, 106, 2009
|
|
1KPB
 
 | | PKCI-1-APO | | Descriptor: | HUMAN PROTEIN KINASE C INTERACTING PROTEIN 1 (ZINC PROTEIN) | | Authors: | Lima, C.D, Klein, M.G, Weinstein, I.B, Hendrickson, W.A. | | Deposit date: | 1996-01-06 | | Release date: | 1996-07-11 | | Last modified: | 2024-02-14 | | Method: | X-RAY DIFFRACTION (2 Å) | | Cite: | Three-dimensional structure of human protein kinase C interacting protein 1, a member of the HIT family of proteins.
Proc.Natl.Acad.Sci.USA, 93, 1996
|
|
1KPA
 
 | | PKCI-1-ZINC | | Descriptor: | HUMAN PROTEIN KINASE C INTERACTING PROTEIN 1 (ZINC PROTEIN) | | Authors: | Lima, C.D, Klein, M.G, Weinstein, I.B, Hendrickson, W.A. | | Deposit date: | 1996-01-06 | | Release date: | 1996-07-11 | | Last modified: | 2024-02-14 | | Method: | X-RAY DIFFRACTION (2 Å) | | Cite: | Three-dimensional structure of human protein kinase C interacting protein 1, a member of the HIT family of proteins.
Proc.Natl.Acad.Sci.USA, 93, 1996
|
|
1KPC
 
 | | PKCI-1-APO+ZINC | | Descriptor: | HUMAN PROTEIN KINASE C INTERACTING PROTEIN 1 (ZINC PROTEIN) | | Authors: | Lima, C.D, Klein, M.G, Weinstein, I.B, Hendrickson, W.A. | | Deposit date: | 1996-01-06 | | Release date: | 1996-07-11 | | Last modified: | 2024-02-14 | | Method: | X-RAY DIFFRACTION (2.2 Å) | | Cite: | Three-dimensional structure of human protein kinase C interacting protein 1, a member of the HIT family of proteins.
Proc.Natl.Acad.Sci.USA, 93, 1996
|
|
4AOM
 
 | | MTIP and MyoA complex | | Descriptor: | MYOSIN A TAIL DOMAIN INTERACTING PROTEIN, MYOSIN-A | | Authors: | Salgado, P.S, Douse, C.H, Simpson, P.J, Thomas, J.C, Holder, A.A, Tate, E.W, Cota, E. | | Deposit date: | 2012-03-29 | | Release date: | 2012-09-05 | | Last modified: | 2023-12-20 | | Method: | X-RAY DIFFRACTION (1.939 Å) | | Cite: | Regulation of the Plasmodium Motor Complex: Phosphorylation of Myosin a Tail Interacting Protein (Mtip) Loosens its Grip on Myoa
J.Biol.Chem., 287, 2012
|
|
1D2F
 
 | |
2X3J
 
 | | CO-COMPLEX STRUCTURE OF ACHROMOBACTIN SYNTHETASE PROTEIN D (ACSD) WITH ATP AND N-CITRYL-ETHYLENEDIAMINE FROM PECTOBACTERIUM CHRYSANTHEMI | | Descriptor: | (2S)-2-{2-[(2-AMINOETHYL)AMINO]-2-OXOETHYL}-2-HYDROXYBUTANEDIOIC ACID, ACSD, ADENOSINE-5'-TRIPHOSPHATE, ... | | Authors: | Schmelz, S, Challis, G.L, Naismith, J.H. | | Deposit date: | 2010-01-25 | | Release date: | 2011-01-19 | | Last modified: | 2023-12-20 | | Method: | X-RAY DIFFRACTION (2 Å) | | Cite: | Structural Basis for Acyl Acceptor Specificity in the Achromobactin Biosynthetic Enzyme Acsd.
J.Mol.Biol., 412, 2011
|
|
1X71
 
 | | Crystal structure of Siderocalin (NGAL, Lipocalin 2) complexed with TRENCAM-3,2-HOPO, a cepabactin analogue | | Descriptor: | 2,3-DIHYDROXYBENZAMIDE, FE (III) ION, Neutrophil gelatinase-associated lipocalin | | Authors: | Holmes, M.A, Paulsene, W, Jide, X, Ratledge, C, Strong, R.K. | | Deposit date: | 2004-08-12 | | Release date: | 2005-01-18 | | Last modified: | 2024-04-03 | | Method: | X-RAY DIFFRACTION (2.1 Å) | | Cite: | Siderocalin (Lcn 2) Also Binds Carboxymycobactins, Potentially Defending against Mycobacterial Infections through Iron Sequestration
Structure, 13, 2005
|
|
3CBC
 
 | | Crystal structure of Siderocalin (NGAL, Lipocalin 2) Y106F complexed with Ferric Enterobactin | | Descriptor: | 2-(2,3-DIHYDROXY-BENZOYLAMINO)-3-HYDROXY-PROPIONIC ACID, GLYCEROL, Neutrophil gelatinase-associated lipocalin, ... | | Authors: | Clifton, M.C, Pizaro, J.C, Strong, R.K. | | Deposit date: | 2008-02-21 | | Release date: | 2008-09-09 | | Last modified: | 2024-03-13 | | Method: | X-RAY DIFFRACTION (2.17 Å) | | Cite: | The siderocalin/enterobactin interaction: a link between mammalian immunity and bacterial iron transport.
J.Am.Chem.Soc., 130, 2008
|
|
2F42
 
 | |
