6BZK
 
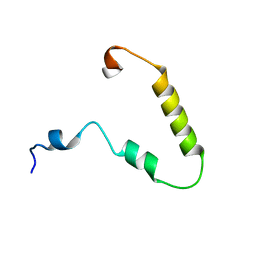 | | Solution structure of KTI55 | | Descriptor: | M protein | | Authors: | Castellino, F.J, Qiu, C, Yuan, Y. | | Deposit date: | 2017-12-24 | | Release date: | 2019-01-16 | | Last modified: | 2024-05-01 | | Method: | SOLUTION NMR | | Cite: | Contributions of different modules of the plasminogen-binding Streptococcus pyogenes M-protein that mediate its functional dimerization.
J.Struct.Biol., 204, 2018
|
|
2P64
 
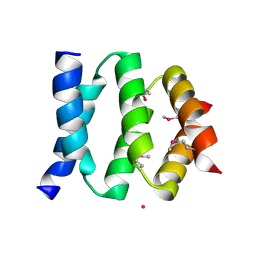 | | D domain of b-TrCP | | Descriptor: | CADMIUM ION, F-box/WD repeat protein 1A | | Authors: | Neculai, D, Orlicky, S, Ceccarelli, D. | | Deposit date: | 2007-03-16 | | Release date: | 2007-06-19 | | Last modified: | 2011-07-13 | | Method: | X-RAY DIFFRACTION (2.5 Å) | | Cite: | Suprafacial orientation of the SCFCdc4 dimer accommodates multiple geometries for substrate ubiquitination.
Cell(Cambridge,Mass.), 129, 2007
|
|
6BZL
 
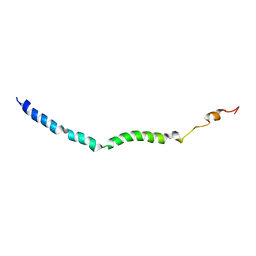 | | Solution structure of VEK75 | | Descriptor: | M protein | | Authors: | Qiu, C, Yuan, Y, Castellino, F.J. | | Deposit date: | 2017-12-24 | | Release date: | 2019-01-16 | | Last modified: | 2024-05-01 | | Method: | SOLUTION NMR | | Cite: | Contributions of different modules of the plasminogen-binding Streptococcus pyogenes M-protein that mediate its functional dimerization.
J.Struct.Biol., 204, 2018
|
|
3SSD
 
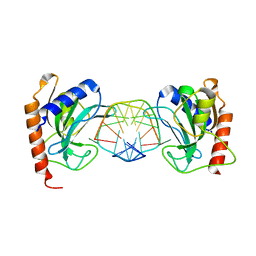 | | DNA binding domain of restriction endonuclease bound to DNA | | Descriptor: | 5-methylcytosine-specific restriction enzyme B, DNA (5'-D(*A*GP*CP*TP*AP*CP*CP*GP*GP*TP*CP*TP*C)-3'), DNA (5'-D(*T*GP*AP*GP*AP*(5CM)P*CP*GP*GP*TP*AP*GP*C)-3') | | Authors: | Sukackaite, R, Grazulis, S, Siksnys, V. | | Deposit date: | 2011-07-08 | | Release date: | 2012-05-23 | | Last modified: | 2024-02-28 | | Method: | X-RAY DIFFRACTION (2.2 Å) | | Cite: | The recognition domain of the methyl-specific endonuclease McrBC flips out 5-methylcytosine.
Nucleic Acids Res., 40, 2012
|
|
6BQ0
 
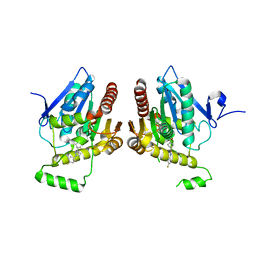 | | Structure of human monoacylglycerol lipase bound to a covalent inhibitor | | Descriptor: | 1-({(1R,5S,6r)-6-[1-(4-fluorophenyl)-1H-pyrazol-3-yl]-3-azabicyclo[3.1.0]hexane-3-carbonyl}oxy)pyrrolidine-2,5-dione, Monoglyceride lipase | | Authors: | Pandit, J. | | Deposit date: | 2017-11-27 | | Release date: | 2018-03-14 | | Last modified: | 2024-10-09 | | Method: | X-RAY DIFFRACTION (2 Å) | | Cite: | Discovery of Trifluoromethyl Glycol Carbamates as Potent and Selective Covalent Monoacylglycerol Lipase (MAGL) Inhibitors for Treatment of Neuroinflammation.
J. Med. Chem., 61, 2018
|
|
2WFK
 
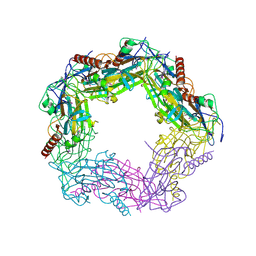 | | Calcium bound LipL32 | | Descriptor: | CALCIUM ION, LIPL32 | | Authors: | Tung, J.-Y, Yang, C.-W, Sun, Y.-J. | | Deposit date: | 2009-04-07 | | Release date: | 2009-11-24 | | Last modified: | 2011-07-13 | | Method: | X-RAY DIFFRACTION (2.3 Å) | | Cite: | Calcium Binds to Lipl32, a Lipoprotein from Pathogenic Leptospira, and Modulates Fibronectin Binding.
J.Biol.Chem., 285, 2010
|
|
6BZJ
 
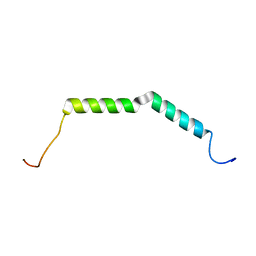 | | Solution structure of AGL55 | | Descriptor: | M protein | | Authors: | Qiu, C, Yuan, Y, Castellino, F.J. | | Deposit date: | 2017-12-24 | | Release date: | 2019-01-16 | | Last modified: | 2024-05-01 | | Method: | SOLUTION NMR | | Cite: | Contributions of different modules of the plasminogen-binding Streptococcus pyogenes M-protein that mediate its functional dimerization.
J.Struct.Biol., 204, 2018
|
|
3SSE
 
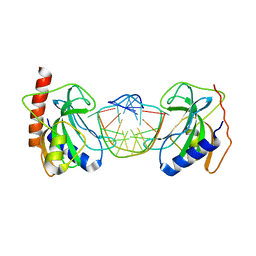 | | DNA binding domain of restriction endonuclease bound to DNA | | Descriptor: | 5-methylcytosine-specific restriction enzyme B, DNA (5'-D(*A*GP*CP*TP*AP*CP*CP*GP*GP*TP*CP*TP*C)-3'), DNA (5'-D(*T*GP*AP*GP*AP*CP*CP*GP*GP*TP*AP*GP*C)-3') | | Authors: | Sukackaite, R, Grazulis, S, Siksnys, V. | | Deposit date: | 2011-07-08 | | Release date: | 2012-05-23 | | Last modified: | 2024-02-28 | | Method: | X-RAY DIFFRACTION (2.7 Å) | | Cite: | The recognition domain of the methyl-specific endonuclease McrBC flips out 5-methylcytosine.
Nucleic Acids Res., 40, 2012
|
|
2OKU
 
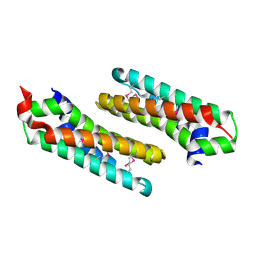 | |
3S2Z
 
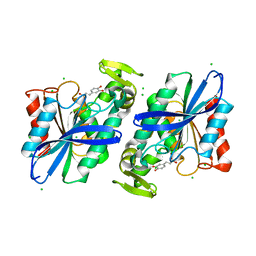 | | Crystal structure of the Lactobacillus johnsonii cinnamoyl esterase LJ0536 S106A mutant in complex with caffeic acid | | Descriptor: | CAFFEIC ACID, CHLORIDE ION, Cinnamoyl esterase | | Authors: | Stogios, P.J, Lai, K.K, Vu, C, Xu, X, Cui, H, Molloy, S, Gonzalez, C.F, Yakunin, A, Savchenko, A. | | Deposit date: | 2011-05-17 | | Release date: | 2011-08-31 | | Last modified: | 2023-09-13 | | Method: | X-RAY DIFFRACTION (1.76 Å) | | Cite: | An inserted alpha/beta subdomain shapes the catalytic pocket of Lactobacillus johnsonii cinnamoyl esterase.
Plos One, 6, 2011
|
|
3BT4
 
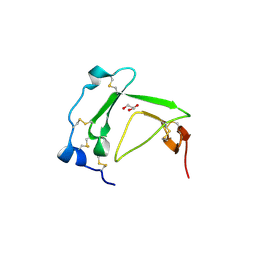 | | Crystal Structure Analysis of AmFPI-1, fungal protease inhibitor from Antheraea mylitta | | Descriptor: | Fungal protease inhibitor-1, GLYCEROL | | Authors: | Roy, S, Aravind, P, Madhurantakam, C, Ghosh, A.K, Sankarananarayanan, R, Das, A.K. | | Deposit date: | 2007-12-27 | | Release date: | 2008-12-30 | | Last modified: | 2017-10-25 | | Method: | X-RAY DIFFRACTION (2.1 Å) | | Cite: | Crystal structure of a fungal protease inhibitor from Antheraea mylitta
J.Struct.Biol., 166, 2009
|
|
3SSC
 
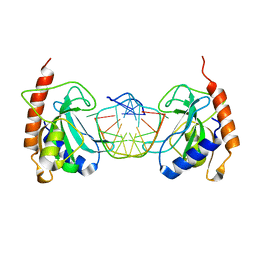 | | DNA binding domain of restriction endonuclease bound to DNA | | Descriptor: | 5-methylcytosine-specific restriction enzyme B, DNA (5'-D(*AP*GP*CP*TP*AP*(5CM)P*CP*GP*GP*TP*CP*TP*C)-3'), DNA (5'-D(*TP*GP*AP*GP*AP*(5CM)P*CP*GP*GP*TP*AP*GP*C)-3') | | Authors: | Sukackaite, R, Grazulis, S, Siksnys, V. | | Deposit date: | 2011-07-08 | | Release date: | 2012-05-23 | | Last modified: | 2024-02-28 | | Method: | X-RAY DIFFRACTION (2.1 Å) | | Cite: | The recognition domain of the methyl-specific endonuclease McrBC flips out 5-methylcytosine.
Nucleic Acids Res., 40, 2012
|
|
5G5M
 
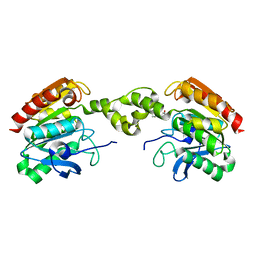 | |
5G5C
 
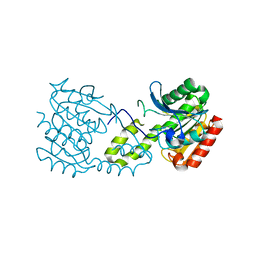 | |
2WP0
 
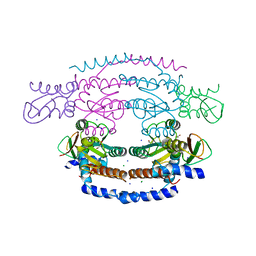 | | Crystal structure of a HobA-DnaA (domain I-II) complex from Helicobacter pylori. | | Descriptor: | ACETATE ION, CHLORIDE ION, CHROMOSOMAL REPLICATION INITIATOR PROTEIN DNAA, ... | | Authors: | Natrajan, G, Noirot-Gros, M.-F, Zawilak-Pawlik, A, Kapp, U, Terradot, L. | | Deposit date: | 2009-07-31 | | Release date: | 2009-11-17 | | Last modified: | 2023-12-20 | | Method: | X-RAY DIFFRACTION (2.67 Å) | | Cite: | The Structure of a Dnaa/Hoba Complex from Helicobacter Pylori Provides Insight Into Regulation of DNA Replication in Bacteria.
Proc.Natl.Acad.Sci.USA, 106, 2009
|
|
5G59
 
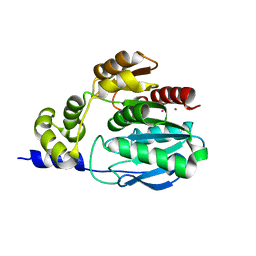 | |
4ZXF
 
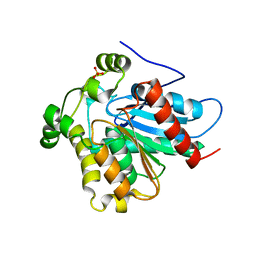 | | Crystal Structure of a Soluble Variant of Monoglyceride Lipase from Saccharomyces Cerevisiae in Complex with a Substrate Analog | | Descriptor: | 1-{3-[(R)-hydroxy(octadecyloxy)phosphoryl]propyl}triaza-1,2-dien-2-ium, Monoglyceride lipase, NITRATE ION, ... | | Authors: | Aschauer, P, Lichtenegger, J, Rengachari, S, Gruber, K, Oberer, M. | | Deposit date: | 2015-05-20 | | Release date: | 2016-05-25 | | Last modified: | 2024-01-10 | | Method: | X-RAY DIFFRACTION (2.5 Å) | | Cite: | Crystal structure of the Saccharomyces cerevisiae monoglyceride lipase Yju3p.
Biochim.Biophys.Acta, 1861, 2016
|
|
3QM1
 
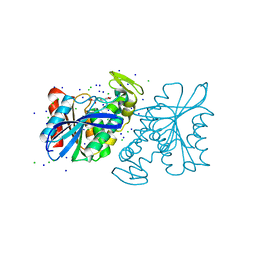 | | CRYSTAL STRUCTURE OF THE LACTOBACILLUS JOHNSONII CINNAMOYL ESTERASE LJ0536 S106A MUTANT IN COMPLEX WITH ETHYLFERULATE, Form II | | Descriptor: | CHLORIDE ION, Cinnamoyl esterase, SODIUM ION, ... | | Authors: | Stogios, P.J, Lai, K.K, Vu, C, Xu, X, Cui, H, Molloy, S, Gonzalez, C.F, Yakunin, A, Savchenko, A. | | Deposit date: | 2011-02-03 | | Release date: | 2011-08-31 | | Last modified: | 2023-09-13 | | Method: | X-RAY DIFFRACTION (1.817 Å) | | Cite: | An Inserted alpha/beta Subdomain Shapes the Catalytic Pocket of Lactobacillus johnsonii Cinnamoyl Esterase
Plos One, 6, 2011
|
|
5HLJ
 
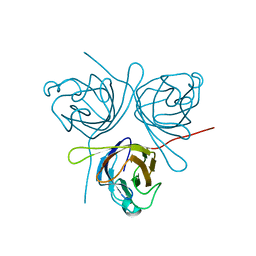 | | Crystal Structure of Major Envelope Protein VP24 from White Spot Syndrome Virus | | Descriptor: | VP24 | | Authors: | Sun, L.F, Su, Y.T, Zhao, Y.H, Fu, Z.Q, Wu, Y.K. | | Deposit date: | 2016-01-15 | | Release date: | 2016-09-14 | | Last modified: | 2024-04-03 | | Method: | X-RAY DIFFRACTION (2.406 Å) | | Cite: | Crystal Structure of Major Envelope Protein VP24 from White Spot Syndrome Virus
Sci Rep, 6, 2016
|
|
5LCN
 
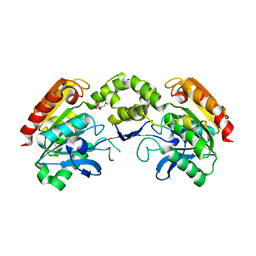 | |
5LE0
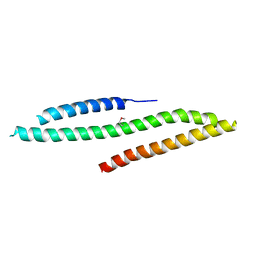 
 | | MICAL1 Cterminal domain | | Descriptor: | Protein-methionine sulfoxide oxidase MICAL1 | | Authors: | Hammich, H, Pylypenko, O, Houdusse, A. | | Deposit date: | 2016-06-29 | | Release date: | 2017-03-01 | | Last modified: | 2017-03-08 | | Method: | X-RAY DIFFRACTION (3.3 Å) | | Cite: | Oxidation of F-actin controls the terminal steps of cytokinesis.
Nat Commun, 8, 2017
|
|
5LUK
 
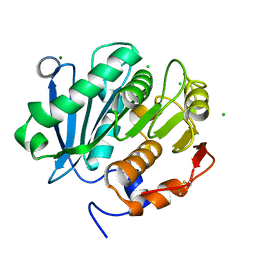 | | Structure of a double variant of cutinase 2 from Thermobifida cellulosilytica | | Descriptor: | CHLORIDE ION, Cutinase 2, MAGNESIUM ION | | Authors: | Hromic, A, Lyskowski, A, Gruber, K. | | Deposit date: | 2016-09-09 | | Release date: | 2017-07-12 | | Last modified: | 2024-01-17 | | Method: | X-RAY DIFFRACTION (1.45 Å) | | Cite: | Small cause, large effect: Structural characterization of cutinases from Thermobifida cellulosilytica.
Biotechnol. Bioeng., 114, 2017
|
|
7AC1
 
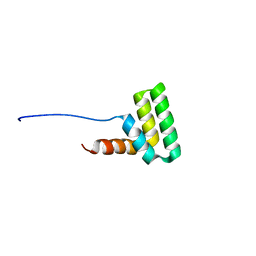 | | Solution structure of the TAF4-RST domain | | Descriptor: | Transcription initiation factor TFIID subunit 4 | | Authors: | Staby, L, Davidsen, R, Bugge, K, Skriver, K, Kragelund, B.B. | | Deposit date: | 2020-09-09 | | Release date: | 2021-10-06 | | Last modified: | 2024-06-19 | | Method: | SOLUTION NMR | | Cite: | alpha alpha-hub coregulator structure and flexibility determine transcription factor binding and selection in regulatory interactomes.
J.Biol.Chem., 298, 2022
|
|
5NNT
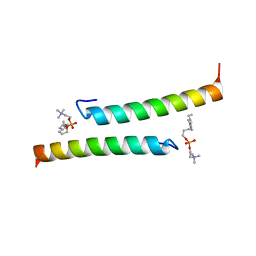 
 | | The dimeric structure of LL-37 crystallized in DPC | | Descriptor: | Cathelicidin antimicrobial peptide, dodecyl 2-(trimethylammonio)ethyl phosphate | | Authors: | Zeth, K, Sancho-Vaello, E. | | Deposit date: | 2017-04-10 | | Release date: | 2018-01-24 | | Last modified: | 2024-05-08 | | Method: | X-RAY DIFFRACTION (2.209 Å) | | Cite: | Structural remodeling and oligomerization of human cathelicidin on membranes suggest fibril-like structures as active species.
Sci Rep, 7, 2017
|
|
5LUJ
 
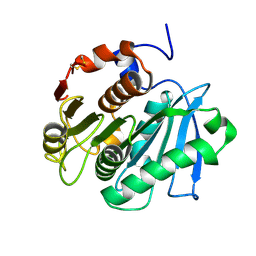 | |
