2KF4
 
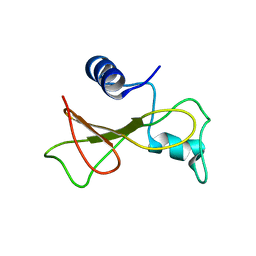 | | Barnase high pressure structure | | Descriptor: | Ribonuclease | | Authors: | Williamson, M.P, Wilton, D.J. | | Deposit date: | 2009-02-11 | | Release date: | 2009-12-08 | | Last modified: | 2024-05-08 | | Method: | SOLUTION NMR | | Cite: | Pressure-dependent structure changes in barnase on ligand binding reveal intermediate rate fluctuations.
Biophys.J., 97, 2009
|
|
1I3E
 
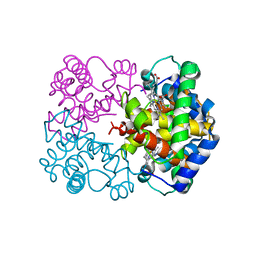 | | HUMAN AZIDO-MET HEMOGLOBIN BART'S (GAMMA4) | | Descriptor: | AZIDE ION, HEMOGLOBIN GAMMA CHAINS, PROTOPORPHYRIN IX CONTAINING FE | | Authors: | Kidd, R.D, Baker, H.M, Mathews, A.J, Brittain, T, Baker, E.N. | | Deposit date: | 2001-02-15 | | Release date: | 2001-09-12 | | Last modified: | 2024-04-03 | | Method: | X-RAY DIFFRACTION (1.86 Å) | | Cite: | Oligomerization and ligand binding in a homotetrameric hemoglobin: two high-resolution crystal structures of hemoglobin Bart's (gamma(4)), a marker for alpha-thalassemia.
Protein Sci., 10, 2001
|
|
5QCF
 
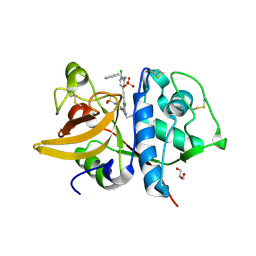 | | Crystal structure of human Cathepsin-S with bound ligand | | Descriptor: | Cathepsin S, GLYCEROL, N-benzyl-1-{2-chloro-5-[(2-chloro-5-{5-(methylsulfonyl)-1-[3-(morpholin-4-yl)propyl]-4,5,6,7-tetrahydro-1H-pyrazolo[4,3-c]pyridin-3-yl}phenyl)ethynyl]phenyl}methanamine, ... | | Authors: | Bembenek, S.D, Ameriks, M.K, Mirzadegan, T, Yang, H, Shao, C, Burley, S.K. | | Deposit date: | 2017-08-04 | | Release date: | 2017-12-20 | | Last modified: | 2024-10-23 | | Method: | X-RAY DIFFRACTION (2.1 Å) | | Cite: | Crystal structure of human Cathepsin-S with bound ligand
To be published
|
|
2KF5
 
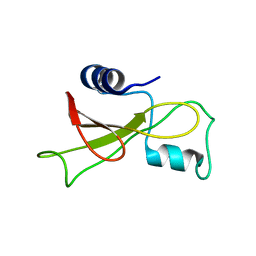 | |
5Y9W
 
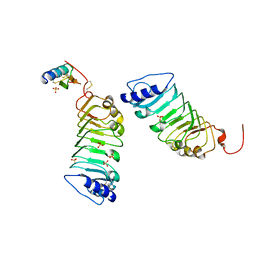 | | Crystal 1 for AtLURE1.2-AtPRK6LRR | | Descriptor: | Pollen receptor-like kinase 6, Protein LURE 1.2, SULFATE ION | | Authors: | Chai, J, Zhang, X. | | Deposit date: | 2017-08-28 | | Release date: | 2017-12-27 | | Last modified: | 2024-11-06 | | Method: | X-RAY DIFFRACTION (1.847 Å) | | Cite: | Structural basis for receptor recognition of pollen tube attraction peptides.
Nat Commun, 8, 2017
|
|
2KF3
 
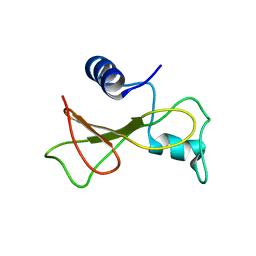 | |
6FBX
 
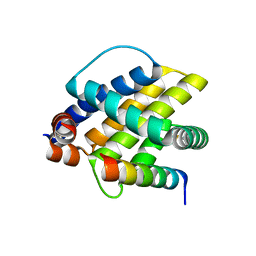 | |
5YAH
 
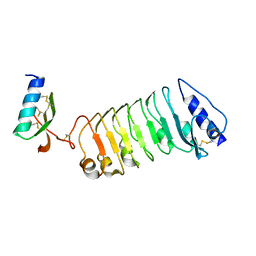 | | Crystal 2 for AtLURE1.2-AtPRK6LRR | | Descriptor: | Pollen receptor-like kinase 6, Protein LURE 1.2 | | Authors: | Chai, J, Zhang, X. | | Deposit date: | 2017-08-31 | | Release date: | 2018-04-11 | | Last modified: | 2024-11-20 | | Method: | X-RAY DIFFRACTION (2.104 Å) | | Cite: | Structural basis for receptor recognition of pollen tube attraction peptides.
Nat Commun, 8, 2017
|
|
5YEM
 
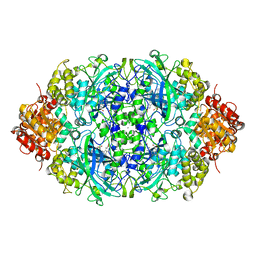 | | CATPO mutant - T188F | | Descriptor: | CALCIUM ION, Catalase, PROTOPORPHYRIN IX CONTAINING FE | | Authors: | Yuzugullu Karakus, Y, Balci, S, Goc, G, Pearson, A.R, Yorke, B. | | Deposit date: | 2017-09-18 | | Release date: | 2018-07-18 | | Last modified: | 2024-02-14 | | Method: | X-RAY DIFFRACTION (1.49 Å) | | Cite: | Investigation of how gate residues in the main channel affect the catalytic activity of Scytalidium thermophilum catalase.
Acta Crystallogr D Struct Biol, 80, 2024
|
|
7YY0
 
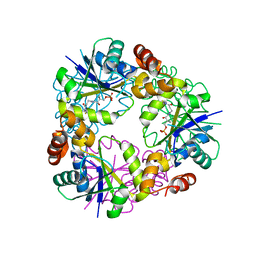 | |
7YY5
 
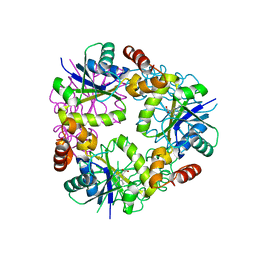 | |
7YY6
 
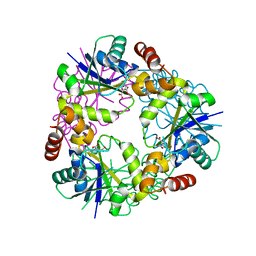 | |
7YY1
 
 | |
7YY3
 
 | |
7YY9
 
 | |
7YYB
 
 | |
7YYC
 
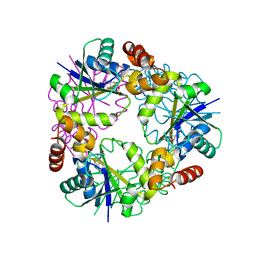 | |
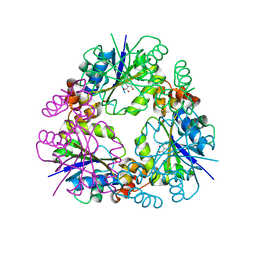
7YY7
 
 | |
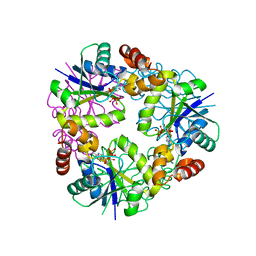
7YWM
 
 | |
6D2I
 
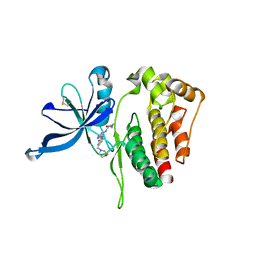 | | JAK2 Pseudokinase V617F in complex with AT9283 | | Descriptor: | 1-cyclopropyl-3-{3-[5-(morpholin-4-ylmethyl)-1H-benzimidazol-2-yl]-1H-pyrazol-4-yl}urea, Tyrosine-protein kinase | | Authors: | Li, Q, Li, K, Eck, M.J. | | Deposit date: | 2018-04-13 | | Release date: | 2019-03-20 | | Last modified: | 2024-10-30 | | Method: | X-RAY DIFFRACTION (3.192 Å) | | Cite: | Discovery and Structural Characterization of ATP-Site Ligands for the Wild-Type and V617F Mutant JAK2 Pseudokinase Domain.
ACS Chem. Biol., 14, 2019
|
|
7YY2
 
 | |
7YY8
 
 | |
7YY4
 
 | |
7YYA
 
 | |
2DJV
 
 | | Solution structures of the WHEP-TRS domain of human methionyl-tRNA synthetase | | Descriptor: | Methionyl-tRNA synthetase | | Authors: | Sato, M, Sasagawa, A, Tochio, N, Koshiba, S, Inoue, M, Kigawa, T, Yokoyama, S, RIKEN Structural Genomics/Proteomics Initiative (RSGI) | | Deposit date: | 2006-04-05 | | Release date: | 2006-10-05 | | Last modified: | 2024-05-29 | | Method: | SOLUTION NMR | | Cite: | Solution structures of the WHEP-TRS domain of human methionyl-tRNA synthetase
To be Published
|
|
