1FTF
 
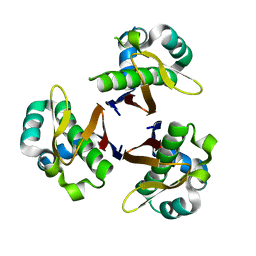 | | CRYSTAL STRUCTURE OF STREPTOCOCCUS PNEUMONIAE ACYL CARRIER PROTEIN SYNTHASE (NATIVE 2) | | Descriptor: | ACYL CARRIER PROTEIN SYNTHASE | | Authors: | Chirgadze, N, Briggs, S, McAllister, K, Fischl, A, Zhao, G. | | Deposit date: | 2000-09-12 | | Release date: | 2001-09-12 | | Last modified: | 2023-08-09 | | Method: | X-RAY DIFFRACTION (2.05 Å) | | Cite: | Crystal structure of Streptococcus pneumoniae acyl carrier protein synthase: an essential enzyme in bacterial fatty acid biosynthesis.
EMBO J., 19, 2000
|
|
1FTG
 
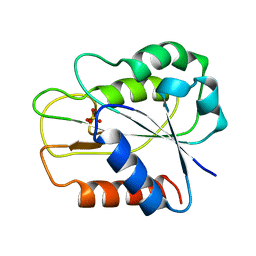 | |
1FTH
 
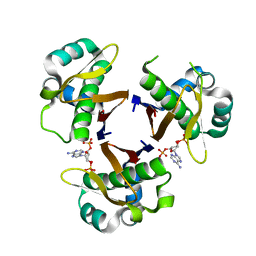 | | CRYSTAL STRUCTURE OF STREPTOCOCCUS PNEUMONIAE ACYL CARRIER PROTEIN SYNTHASE (3'5'-ADP COMPLEX) | | Descriptor: | ACYL CARRIER PROTEIN SYNTHASE, ADENOSINE-3'-5'-DIPHOSPHATE | | Authors: | Chirgadze, N, Briggs, S, McAllister, K, Fischl, A, Zhao, G. | | Deposit date: | 2000-09-12 | | Release date: | 2001-09-12 | | Last modified: | 2023-08-09 | | Method: | X-RAY DIFFRACTION (1.9 Å) | | Cite: | Crystal structure of Streptococcus pneumoniae acyl carrier protein synthase: an essential enzyme in bacterial fatty acid biosynthesis.
EMBO J., 19, 2000
|
|
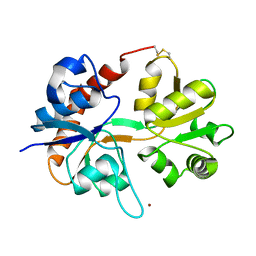
1FTJ
 
 | |
1FTK
 
 | |
1FTL
 
 | |
1FTM
 
 | |
1FTN
 
 | | CRYSTAL STRUCTURE OF THE HUMAN RHOA/GDP COMPLEX | | Descriptor: | GUANOSINE-5'-DIPHOSPHATE, MAGNESIUM ION, TRANSFORMING PROTEIN RHOA (H12) | | Authors: | Wei, Y, Zhang, Y, Derewenda, U, Liu, X, Minor, W, Nakamoto, R.K, Somlyo, A.V, Somlyo, A.P, Derewenda, Z.S. | | Deposit date: | 1997-03-13 | | Release date: | 1998-03-18 | | Last modified: | 2024-04-03 | | Method: | X-RAY DIFFRACTION (2.1 Å) | | Cite: | Crystal structure of RhoA-GDP and its functional implications.
Nat.Struct.Biol., 4, 1997
|
|
1FTO
 
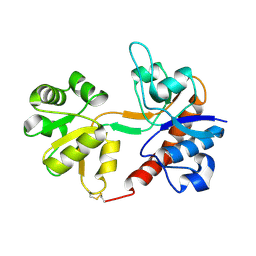 | |
1FTP
 
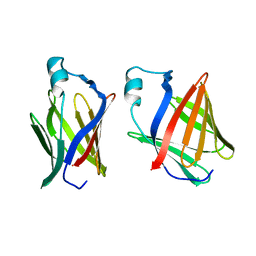 | |
1FTQ
 
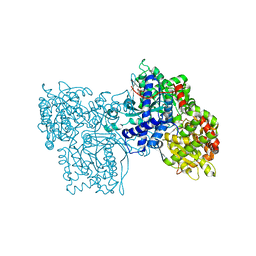 | | STRUCTURES OF GLYCOGEN PHOSPHORYLASE-INHIBITOR COMPLEXES AND THE IMPLICATIONS FOR STRUCTURE-BASED DRUG DESIGN | | Descriptor: | (5S,7R,8S,9S,10R)-3-amino-8,9,10-trihydroxy-7-(hydroxymethyl)-6-oxa-1,3-diazaspiro[4.5]decane-2,4-dione, GLYCOGEN PHOSPHORYLASE, INOSINIC ACID, ... | | Authors: | Watson, K.A, Tsitsanou, K.E, Gregoriou, M, Zographos, S.E, Skamnaki, V.T, Oikonomakos, N.G, Fleet, G.W, Johnson, L.N. | | Deposit date: | 2000-09-13 | | Release date: | 2000-10-04 | | Last modified: | 2023-08-09 | | Method: | X-RAY DIFFRACTION (2.35 Å) | | Cite: | Kinetic and crystallographic studies of glucopyranose spirohydantoin and glucopyranosylamine analogs inhibitors of glycogen phosphorylase.
Proteins, 61, 2005
|
|
1FTR
 
 | |
1FTS
 
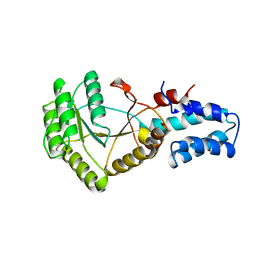 | |
1FTT
 
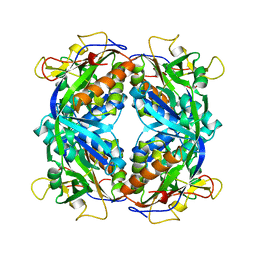
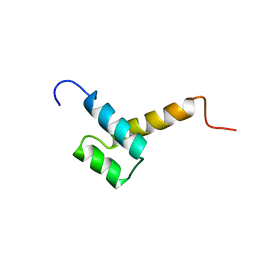 | | THYROID TRANSCRIPTION FACTOR 1 HOMEODOMAIN (RATTUS NORVEGICUS) | | Descriptor: | THYROID TRANSCRIPTION FACTOR 1 HOMEODOMAIN | | Authors: | Fogolari, F, Esposito, G, Damante, G, Formisano, S, Di Lauro, R, Viglino, P. | | Deposit date: | 1995-10-03 | | Release date: | 1996-01-29 | | Last modified: | 2024-05-22 | | Method: | SOLUTION NMR | | Cite: | Analysis of the solution structure of the homeodomain of rat thyroid transcription factor 1 by 1H-NMR spectroscopy and restrained molecular mechanics.
Eur.J.Biochem., 241, 1996
|
|
1FTW
 
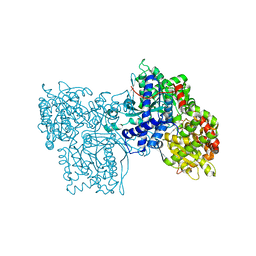 | | STRUCTURES OF GLYCOGEN PHOSPHORYLASE-INHIBITOR COMPLEXES AND THE IMPLICATIONS FOR STRUCTURE-BASED DRUG DESIGN | | Descriptor: | (5S,7R,8S,9S,10R)-3,8,9,10-tetrahydroxy-7-(hydroxymethyl)-6-oxa-1,3-diazaspiro[4.5]decane-2,4-dione, GLYCOGEN PHOSPHORYLASE, PYRIDOXAL-5'-PHOSPHATE | | Authors: | Watson, K.A, Tsitsanou, K.E, Gregoriou, M, Zographos, S.E, Skamnaki, V.T, Oikonomakos, N.G, Fleet, G.W, Johnson, L.N. | | Deposit date: | 2000-09-13 | | Release date: | 2000-10-04 | | Last modified: | 2023-08-09 | | Method: | X-RAY DIFFRACTION (2.36 Å) | | Cite: | Kinetic and crystallographic studies of glucopyranose spirohydantoin and glucopyranosylamine analogs inhibitors of glycogen phosphorylase.
Proteins, 61, 2005
|
|
1FTX
 
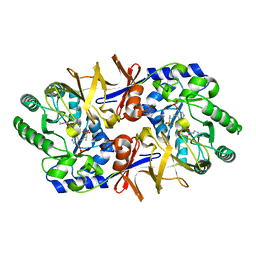 | |
1FTY
 
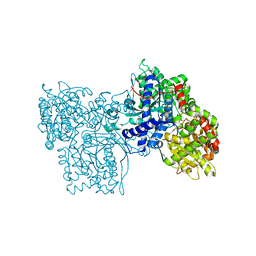 | | STRUCTURES OF GLYCOGEN PHOSPHORYLASE-INHIBITOR COMPLEXES AND THE IMPLICATIONS FOR STRUCTURE-BASED DRUG DESIGN | | Descriptor: | 8,9,10-TRIHYDROXY-7-HYDROXYMETHYL-3-METHYL-6-OXA-1,3-DIAZA-SPIRO[4.5]DECANE-2,4-DIONE, GLYCOGEN PHOSPHORYLASE, PYRIDOXAL-5'-PHOSPHATE | | Authors: | Watson, K.A, Tsitsanou, K.E, Gregoriou, M, Zographos, S.E, Skamnaki, V.T, Oikonomakos, N.G, Fleet, G.W, Johnson, L.N. | | Deposit date: | 2000-09-13 | | Release date: | 2000-10-04 | | Last modified: | 2023-08-09 | | Method: | X-RAY DIFFRACTION (2.38 Å) | | Cite: | Kinetic and crystallographic studies of glucopyranose spirohydantoin and glucopyranosylamine analogs inhibitors of glycogen phosphorylase.
Proteins, 61, 2005
|
|
1FTZ
 
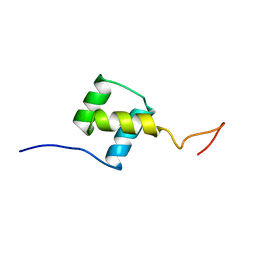 | | NUCLEAR MAGNETIC RESONANCE SOLUTION STRUCTURE OF THE FUSHI TARAZU HOMEODOMAIN FROM DROSOPHILA AND COMPARISON WITH THE ANTENNAPEDIA HOMEODOMAIN | | Descriptor: | FUSHI TARAZU PROTEIN | | Authors: | Qian, Y.Q, Furukubo-Tokunaga, K, Resendez-Perez, D, Muller, M, Gehring, W.J, Wuthrich, K. | | Deposit date: | 1994-01-07 | | Release date: | 1994-05-31 | | Last modified: | 2024-05-22 | | Method: | SOLUTION NMR | | Cite: | Nuclear magnetic resonance solution structure of the fushi tarazu homeodomain from Drosophila and comparison with the Antennapedia homeodomain.
J.Mol.Biol., 238, 1994
|
|
1FU0
 
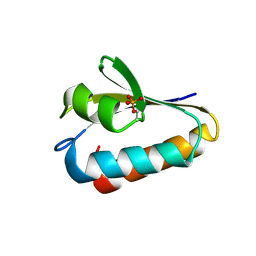 | | CRYSTAL STRUCTURE ANALYSIS OF THE PHOSPHO-SERINE 46 HPR FROM ENTEROCOCCUS FAECALIS | | Descriptor: | PHOSPHOCARRIER PROTEIN HPR | | Authors: | Audette, G.F, Engelmann, R, Hengstenberg, W, Deutscher, J, Hayakawa, K, Quail, J.W, Delbaere, L.T.J. | | Deposit date: | 2000-09-13 | | Release date: | 2000-11-22 | | Last modified: | 2022-12-21 | | Method: | X-RAY DIFFRACTION (1.9 Å) | | Cite: | The 1.9 A resolution structure of phospho-serine 46 HPr from Enterococcus faecalis.
J.Mol.Biol., 303, 2000
|
|
1FU1
 
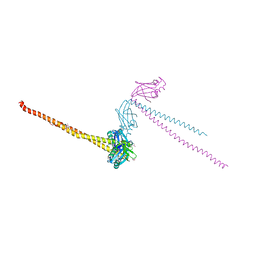 | | CRYSTAL STRUCTURE OF HUMAN XRCC4 | | Descriptor: | ACETIC ACID, DNA REPAIR PROTEIN XRCC4 | | Authors: | Junop, M, Modesti, M, Guarne, A, Gellert, M, Yang, W. | | Deposit date: | 2000-09-13 | | Release date: | 2000-12-11 | | Last modified: | 2021-11-03 | | Method: | X-RAY DIFFRACTION (2.7 Å) | | Cite: | Crystal structure of the Xrcc4 DNA repair protein and implications for end joining.
EMBO J., 19, 2000
|
|
1FU2
 
 | | FIRST PROTEIN STRUCTURE DETERMINED FROM X-RAY POWDER DIFFRACTION DATA | | Descriptor: | CHLORIDE ION, INSULIN, A CHAIN, ... | | Authors: | Von Dreele, R.B, Stephens, P.W, Blessing, R.H, Smith, G.D. | | Deposit date: | 2000-09-13 | | Release date: | 2000-10-16 | | Last modified: | 2018-10-03 | | Method: | POWDER DIFFRACTION | | Cite: | The first protein crystal structure determined from high-resolution X-ray powder diffraction data: a variant of T3R3 human insulin-zinc complex produced by grinding.
Acta Crystallogr.,Sect.D, 56, 2000
|
|
1FU3
 | |
1FU4
 
 | | STRUCTURES OF GLYCOGEN PHOSPHORYLASE-INHIBITOR COMPLEXES AND THE IMPLICATIONS FOR STRUCTURE-BASED DRUG DESIGN | | Descriptor: | GLYCOGEN PHOSPHORYLASE, N-[(5S,7R,8S,9S,10R)-8,9,10-trihydroxy-7-(hydroxymethyl)-2,4-dioxo-6-oxa-1,3-diazaspiro[4.5]dec-3-yl]acetamide, PYRIDOXAL-5'-PHOSPHATE | | Authors: | Watson, K.A, Tsitsanou, K.E, Gregoriou, M, Zographos, S.E, Skamnaki, V.T, Oikonomakos, N.G, Fleet, G.W, Johnson, L.N. | | Deposit date: | 2000-09-14 | | Release date: | 2000-10-04 | | Last modified: | 2023-08-09 | | Method: | X-RAY DIFFRACTION (2.36 Å) | | Cite: | Kinetic and crystallographic studies of glucopyranose spirohydantoin and glucopyranosylamine analogs inhibitors of glycogen phosphorylase.
Proteins, 61, 2005
|
|
1FU5
 
 | | NMR STRUCTURE OF THE N-SH2 DOMAIN OF THE P85 SUBUNIT OF PI3-KINASE COMPLEXED TO A DOUBLY PHOSPHORYLATED PEPTIDE DERIVED FROM POLYOMAVIRUS MIDDLE T ANTIGEN | | Descriptor: | DOUBLY PHOSPHORYLATED MIDDLE T ANTIGEN, PHOSPHATIDYLINOSITOL 3-KINASE REGULATORY ALPHA SUBUNIT | | Authors: | Weber, T, Schaffhausen, B, Liu, Y, Guenther, U.L. | | Deposit date: | 2000-09-14 | | Release date: | 2001-02-21 | | Last modified: | 2022-02-23 | | Method: | SOLUTION NMR | | Cite: | NMR structure of the N-SH2 of the p85 subunit of phosphoinositide 3-kinase complexed to a doubly phosphorylated peptide reveals a second phosphotyrosine binding site.
Biochemistry, 39, 2000
|
|
1FU6
 
 | |
