1E4G
 
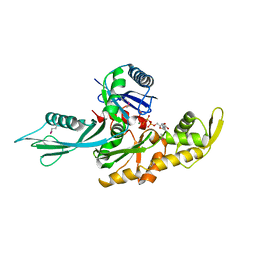 | | FtsA (ATP-bound form) from Thermotoga maritima | | Descriptor: | ADENOSINE-5'-TRIPHOSPHATE, CELL DIVISION PROTEIN FTSA, MAGNESIUM ION | | Authors: | van den Ent, F, Lowe, J. | | Deposit date: | 2000-07-03 | | Release date: | 2000-10-18 | | Last modified: | 2024-10-23 | | Method: | X-RAY DIFFRACTION (2.6 Å) | | Cite: | Crystal Structure of the Cell Division Protein Ftsa from Thermotoga Maritima
Embo J., 19, 2000
|
|
5LG8
 
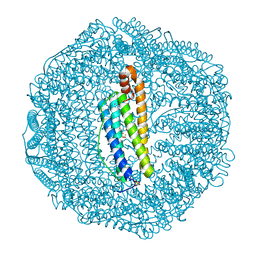 | | Human L-type ferritin iron loaded for 60 minutes | | Descriptor: | CADMIUM ION, FE (III) ION, Ferritin light chain, ... | | Authors: | Pozzi, C, Di Pisa, F, Mangani, S. | | Deposit date: | 2016-07-06 | | Release date: | 2017-02-22 | | Last modified: | 2024-01-10 | | Method: | X-RAY DIFFRACTION (1.98 Å) | | Cite: | Chemistry at the protein-mineral interface in L-ferritin assists the assembly of a functional ( mu (3)-oxo)Tris[( mu (2)-peroxo)] triiron(III) cluster.
Proc. Natl. Acad. Sci. U.S.A., 114, 2017
|
|
1DZJ
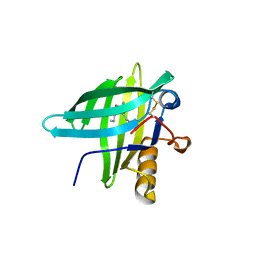 
 | | Porcine Odorant Binding Protein Complexed with 2-amino-4-butyl-5-propylselenazole | | Descriptor: | 4-butyl-5-propyl-1,3-selenazol-2-amine, ODORANT-BINDING PROTEIN | | Authors: | Vincent, F, Spinelli, S, Cambillau, C, Tegoni, M. | | Deposit date: | 2000-03-01 | | Release date: | 2000-12-06 | | Last modified: | 2024-10-23 | | Method: | X-RAY DIFFRACTION (2 Å) | | Cite: | Complexes of Porcine Odorant Binding Protein with Odorant Molecules Belonging to Different Chemical Classes
J.Mol.Biol., 300, 2000
|
|
5LGB
 
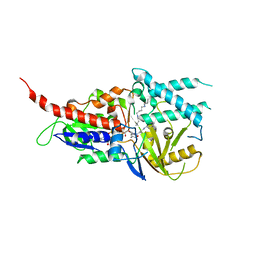 | | Crystal structure of murine N1-acetylpolyamine oxidase in complex with MDL72527 | | Descriptor: | FAD-MDL72527 adduct, N,N'-BIS(2,3-BUTADIENYL)-1,4-BUTANE-DIAMINE, Peroxisomal N(1)-acetyl-spermine/spermidine oxidase,Peroxisomal N(1)-acetyl-spermine/spermidine oxidase | | Authors: | Sjogren, T, Aagaard, A, Wassvik, C, Snijder, A, Barlind, L. | | Deposit date: | 2016-07-06 | | Release date: | 2017-08-16 | | Last modified: | 2024-01-10 | | Method: | X-RAY DIFFRACTION (1.8 Å) | | Cite: | The Structure of Murine N(1)-Acetylspermine Oxidase Reveals Molecular Details of Vertebrate Polyamine Catabolism.
Biochemistry, 56, 2017
|
|
5LDD
 
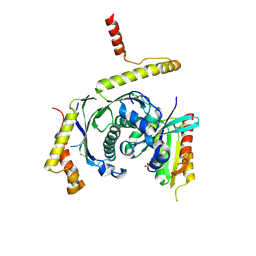 | |
1E1G
 
 | | Human prion protein variant M166V | | Descriptor: | PRION PROTEIN | | Authors: | Calzolai, L, Lysek, D.A, Guntert, P, Von Schroetter, C, Zahn, R, Riek, R, Wuthrich, K. | | Deposit date: | 2000-05-04 | | Release date: | 2000-07-20 | | Last modified: | 2024-11-06 | | Method: | SOLUTION NMR | | Cite: | NMR Structures of Three Single-Residue Variants of the Human Prion Protein
Proc.Natl.Acad.Sci.USA, 97, 2000
|
|
1E1W
 
 | | Human prion protein variant R220K | | Descriptor: | PRION PROTEIN | | Authors: | Calzolai, L, Lysek, D.A, Guntert, P, Von Schroetter, C, Zahn, R, Riek, R, Wuthrich, K. | | Deposit date: | 2000-05-11 | | Release date: | 2000-07-20 | | Last modified: | 2024-11-20 | | Method: | SOLUTION NMR | | Cite: | NMR Structures of Three Single-Residue Variants of the Human Prion Protein
Proc.Natl.Acad.Sci.USA, 97, 2000
|
|
5LE0
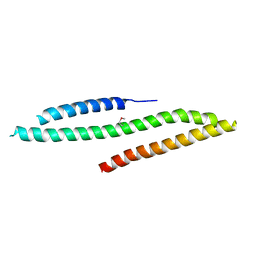 
 | | MICAL1 Cterminal domain | | Descriptor: | Protein-methionine sulfoxide oxidase MICAL1 | | Authors: | Hammich, H, Pylypenko, O, Houdusse, A. | | Deposit date: | 2016-06-29 | | Release date: | 2017-03-01 | | Last modified: | 2024-11-13 | | Method: | X-RAY DIFFRACTION (3.3 Å) | | Cite: | Oxidation of F-actin controls the terminal steps of cytokinesis.
Nat Commun, 8, 2017
|
|
1E02
 
 | | Porcine Odorant Binding Protein Complexed with undecanal | | Descriptor: | ODORANT-BINDING PROTEIN, UNDECANAL | | Authors: | Vincent, F, Spinelli, S, Cambillau, C, Tegoni, M. | | Deposit date: | 2000-03-09 | | Release date: | 2000-12-06 | | Last modified: | 2024-11-06 | | Method: | X-RAY DIFFRACTION (2.15 Å) | | Cite: | Complexes of Porcine Odorant Binding Protein with Odorant Molecules Belonging to Different Chemical Classes
J.Mol.Biol., 300, 2000
|
|
1E1U
 
 | | Human prion protein variant R220K | | Descriptor: | PRION PROTEIN | | Authors: | Calzolai, L, Lysek, D.A, Guntert, P, Von Schroetter, C, Zahn, R, Riek, R, Wuthrich, K. | | Deposit date: | 2000-05-11 | | Release date: | 2000-07-20 | | Last modified: | 2024-11-20 | | Method: | SOLUTION NMR | | Cite: | NMR Structures of Three Single-Residue Variants of the Human Prion Protein
Proc.Natl.Acad.Sci.USA, 97, 2000
|
|
1DZP
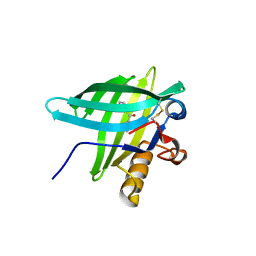 
 | | Porcine Odorant Binding Protein Complexed with diphenylmethanone | | Descriptor: | DIPHENYLMETHANONE, ODORANT-BINDING PROTEIN | | Authors: | Vincent, F, Spinelli, S, Cambillau, C, Tegoni, M. | | Deposit date: | 2000-03-06 | | Release date: | 2000-12-06 | | Last modified: | 2024-11-13 | | Method: | X-RAY DIFFRACTION (1.83 Å) | | Cite: | Complexes of Porcine Odorant Binding Protein with Odorant Molecules Belonging to Different Chemical Classes
J.Mol.Biol., 300, 2000
|
|
5LGQ
 
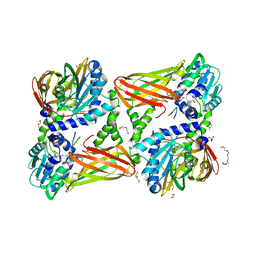 | | Crystal structure of mouse CARM1 in complex with ligand P2C3s | | Descriptor: | (2~{R},3~{R},4~{S},5~{R})-2-(6-aminopurin-9-yl)-5-propyl-oxolane-3,4-diol, 1,2-DIMETHOXYETHANE, 1,2-ETHANEDIOL, ... | | Authors: | Marechal, N, Troffer-Charlier, N, Cura, V, Bonnefond, L, Cavarelli, J. | | Deposit date: | 2016-07-08 | | Release date: | 2017-03-22 | | Last modified: | 2024-11-13 | | Method: | X-RAY DIFFRACTION (2.11 Å) | | Cite: | Transition state mimics are valuable mechanistic probes for structural studies with the arginine methyltransferase CARM1.
Proc. Natl. Acad. Sci. U.S.A., 114, 2017
|
|
5LHT
 
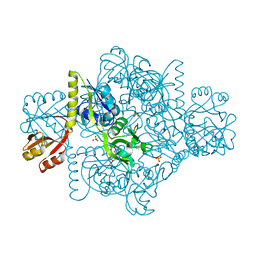 | | ATP Phosphoribosyltransferase from Mycobacterium tuberculosis in complex with the allosteric activator 3-(2-Thienyl)-L-alanine | | Descriptor: | ATP phosphoribosyltransferase, BETA(2-THIENYL)ALANINE, GLYCEROL, ... | | Authors: | de Chiara, C, Pisco, J.P, de Carvalho, L.P, Smerdon, S.J, Walker, P.A, Ogrodowicz, R. | | Deposit date: | 2016-07-12 | | Release date: | 2017-07-05 | | Last modified: | 2024-10-16 | | Method: | X-RAY DIFFRACTION (2.0601 Å) | | Cite: | Uncoupling conformational states from activity in an allosteric enzyme.
Nat Commun, 8, 2017
|
|
1E06
 
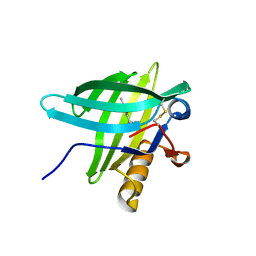 | | Porcine Odorant Binding Protein Complexed with 5-methyl-2-(1-methylethyl)phenol | | Descriptor: | 5-METHYL-2-(1-METHYLETHYL)PHENOL, ODORANT-BINDING PROTEIN | | Authors: | Vincent, F, Spinelli, S, Cambillau, C, Tegoni, M. | | Deposit date: | 2000-03-10 | | Release date: | 2000-12-06 | | Last modified: | 2024-10-09 | | Method: | X-RAY DIFFRACTION (2.12 Å) | | Cite: | Complexes of Porcine Odorant Binding Protein with Odorant Molecules Belonging to Different Chemical Classes
J.Mol.Biol., 300, 2000
|
|
1DWZ
 
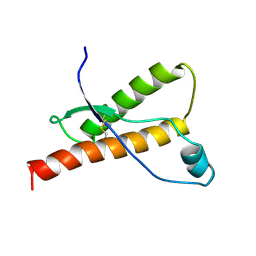 | | Bovine prion protein fragment 121-230 | | Descriptor: | PRION PROTEIN | | Authors: | Lopez-Garcia, F, Zahn, R, Riek, R, Billeter, M, Wuthrich, K. | | Deposit date: | 1999-12-15 | | Release date: | 2000-07-20 | | Last modified: | 2024-10-23 | | Method: | SOLUTION NMR | | Cite: | NMR Structure of the Bovine Prion Protein
Proc.Natl.Acad.Sci.USA, 97, 2000
|
|
5LE4
 
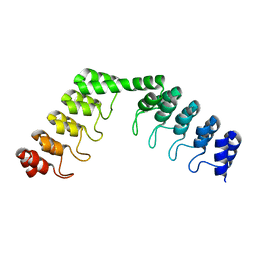 | | Crystal structure of DARPin-DARPin rigid fusion, variant DD_D12_11_D12 | | Descriptor: | DD_D12_11_D12 | | Authors: | Batyuk, A, Wu, Y, Mittl, P.R, Plueckthun, A. | | Deposit date: | 2016-06-29 | | Release date: | 2017-08-02 | | Last modified: | 2024-01-10 | | Method: | X-RAY DIFFRACTION (2.35 Å) | | Cite: | Rigidly connected multispecific artificial binders with adjustable geometries.
Sci Rep, 7, 2017
|
|
5LEA
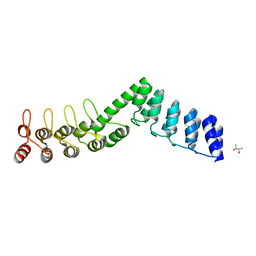 
 | | Crystal structure of DARPin-DARPin rigid fusion, variant DD_Off7_12_3G124 | | Descriptor: | (4S)-2-METHYL-2,4-PENTANEDIOL, DD_Off7_12_3G124 | | Authors: | Batyuk, A, Wu, Y, Mittl, P.R, Plueckthun, A. | | Deposit date: | 2016-06-29 | | Release date: | 2017-08-02 | | Last modified: | 2024-01-10 | | Method: | X-RAY DIFFRACTION (2.4 Å) | | Cite: | Rigidly connected multispecific artificial binders with adjustable geometries.
Sci Rep, 7, 2017
|
|
5LED
 
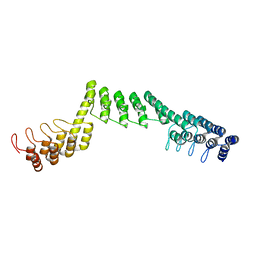 | | Crystal structure of DARPin-DARPin rigid fusion, variant DDD_D12_12_D12_12_D12 | | Descriptor: | DDD_D12_12_D12_12_D12 | | Authors: | Batyuk, A, Wu, Y, Mittl, P.R, Plueckthun, A. | | Deposit date: | 2016-06-29 | | Release date: | 2017-08-02 | | Last modified: | 2024-01-10 | | Method: | X-RAY DIFFRACTION (2.6 Å) | | Cite: | Rigidly connected multispecific artificial binders with adjustable geometries.
Sci Rep, 7, 2017
|
|
1E0C
 
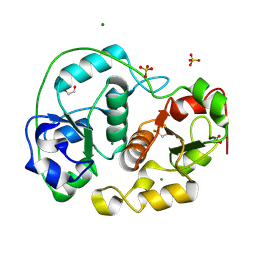 | | SULFURTRANSFERASE FROM AZOTOBACTER VINELANDII | | Descriptor: | 1,2-ETHANEDIOL, MAGNESIUM ION, SULFATE ION, ... | | Authors: | Bordo, D, Deriu, D, Colnaghi, R, Carpen, A, Pagani, S, Bolognesi, M. | | Deposit date: | 2000-03-23 | | Release date: | 2000-05-08 | | Last modified: | 2024-11-13 | | Method: | X-RAY DIFFRACTION (1.8 Å) | | Cite: | The Crystal Structure of a Sulfurtransferase from Azotobacter Vinelandii Highlights the Evolutionary Relationship between the Rhodanese and Phosphatase Enzyme Families
J.Mol.Biol., 298, 2000
|
|
8IM7
 
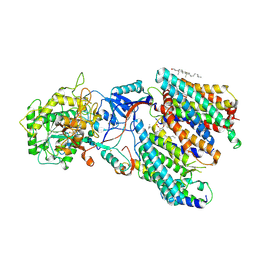 | | Human gamma-secretase treated with ganglioside GM1 | | Descriptor: | 1,2-DIACYL-SN-GLYCERO-3-PHOSPHOCHOLINE, 2-acetamido-2-deoxy-beta-D-glucopyranose, 2-acetamido-2-deoxy-beta-D-glucopyranose-(1-4)-2-acetamido-2-deoxy-beta-D-glucopyranose, ... | | Authors: | Zhou, R, Yang, G, Shi, Y. | | Deposit date: | 2023-03-06 | | Release date: | 2024-01-17 | | Last modified: | 2025-07-23 | | Method: | ELECTRON MICROSCOPY (3.4 Å) | | Cite: | Preferential Regulation of Gamma-Secretase-Mediated Cleavage of APP by Ganglioside GM1 Reveals a Potential Therapeutic Target for Alzheimer's Disease.
Adv Sci, 10, 2023
|
|
5LI3
 
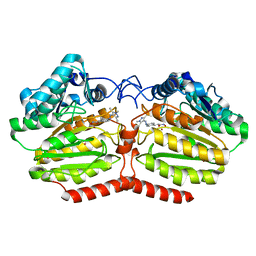 | | Crystal structure of HDAC-like protein from P. aeruginosa in complex with a photo-switchable inhibitor. | | Descriptor: | (2E)-N-hydroxy-3-{4-[(E)-(1,3,5-trimethyl-1H-pyrazol-4-yl)diazenyl]phenyl}prop-2-enamide, Acetoin utilization protein, POTASSIUM ION, ... | | Authors: | Kraemer, A, Meyer-Almes, F.J, Yildiz, O. | | Deposit date: | 2016-07-14 | | Release date: | 2016-11-23 | | Last modified: | 2024-01-10 | | Method: | X-RAY DIFFRACTION (2.4 Å) | | Cite: | Toward Photopharmacological Antimicrobial Chemotherapy Using Photoswitchable Amidohydrolase Inhibitors.
ACS Infect Dis, 3, 2017
|
|
1E0P
 
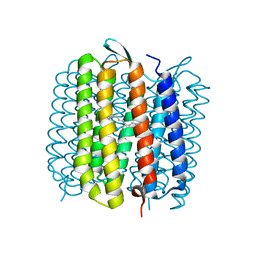 | | L intermediate of bacteriorhodopsin | | Descriptor: | BACTERIORHODOPSIN, GROUND STATE, RETINAL | | Authors: | Royant, A, Edman, K, Ursby, T, Pebay-Peyroula, E, Landau, E.M, Neutze, R. | | Deposit date: | 2000-04-04 | | Release date: | 2000-08-19 | | Last modified: | 2024-11-13 | | Method: | X-RAY DIFFRACTION (2.1 Å) | | Cite: | Helix Deformation is Coupled to Vectorial Proton Transport in Bacteriorhodopsin'S Photocycle
Nature, 406, 2000
|
|
5LIF
 
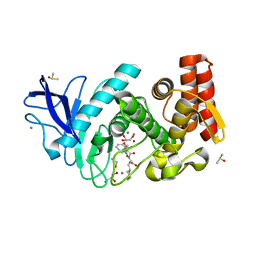 | | Thermolysin in complex with inhibitor | | Descriptor: | (2~{S})-3-cyclohexyl-2-[[(2~{S})-4-methyl-2-[[oxidanyl(phenylmethoxycarbonylaminomethyl)phosphoryl]amino]pentanoyl]amino]propanoic acid, CALCIUM ION, DIMETHYL SULFOXIDE, ... | | Authors: | Krimmer, S.G, Cramer, J, Heine, A, Klebe, G. | | Deposit date: | 2016-07-14 | | Release date: | 2016-12-21 | | Last modified: | 2024-01-10 | | Method: | X-RAY DIFFRACTION (1.31 Å) | | Cite: | Elucidating the Origin of Long Residence Time Binding for Inhibitors of the Metalloprotease Thermolysin.
ACS Chem. Biol., 12, 2017
|
|
1E1P
 
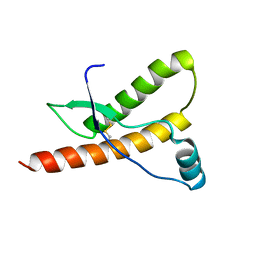 | | Human prion protein variant S170N | | Descriptor: | PRION PROTEIN | | Authors: | Calzolai, L, Lysek, D.A, Guntert, P, Von Schroetter, C, Zahn, R, Riek, R, Wuthrich, K. | | Deposit date: | 2000-05-09 | | Release date: | 2000-07-20 | | Last modified: | 2024-11-13 | | Method: | SOLUTION NMR | | Cite: | NMR Structures of Three Single-Residue Variants of the Human Prion Protein
Proc.Natl.Acad.Sci.USA, 97, 2000
|
|
1E10
 
 | | [2Fe-2S]-Ferredoxin from Halobacterium salinarum | | Descriptor: | ACETYL GROUP, FE2/S2 (INORGANIC) CLUSTER, FERREDOXIN | | Authors: | Marg, B.-L, Schweimer, K, Oesterhelt, D, Roesch, P, Sticht, H. | | Deposit date: | 2000-04-12 | | Release date: | 2001-04-09 | | Last modified: | 2025-04-09 | | Method: | SOLUTION NMR | | Cite: | A Two-Alpha-Helix Extra Domain Mediates the Halophilic Character of a Plant-Type Ferredoxin from Halophilic Archaea.
Biochemistry, 44, 2005
|
|
