5IUK
 
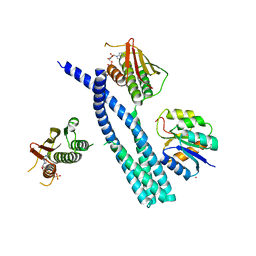 | | Crystal structure of the DesK-DesR complex in the phosphotransfer state with high Mg2+ (150 mM) | | Descriptor: | MAGNESIUM ION, PHOSPHOMETHYLPHOSPHONIC ACID ADENYLATE ESTER, POTASSIUM ION, ... | | Authors: | Trajtenberg, F, Imelio, J.A, Larrieux, N, Buschiazzo, A. | | Deposit date: | 2016-03-18 | | Release date: | 2016-12-21 | | Last modified: | 2024-03-06 | | Method: | X-RAY DIFFRACTION (2.9 Å) | | Cite: | Regulation of signaling directionality revealed by 3D snapshots of a kinase:regulator complex in action.
Elife, 5, 2016
|
|
3UR1
 
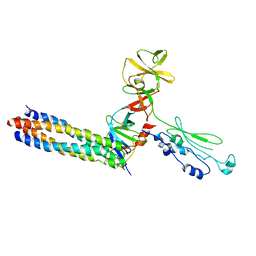 | |
5J5P
 
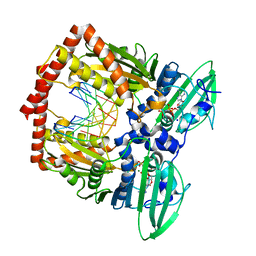 | | AMP-PNP-stabilized ATPase domain of topoisomerase IV from Streptococcus pneumoniae, complex type I | | Descriptor: | DNA (5'-D(*GP*CP*GP*CP*GP*C)-3'), DNA topoisomerase 4 subunit B, MAGNESIUM ION, ... | | Authors: | Laponogov, I, Pan, X.-S, Skamrova, G, Umrekar, T, Fisher, L.M, Sanderson, M.R. | | Deposit date: | 2016-04-03 | | Release date: | 2017-07-26 | | Last modified: | 2024-01-10 | | Method: | X-RAY DIFFRACTION (1.97 Å) | | Cite: | Trapping of the transport-segment DNA by the ATPase domains of a type II topoisomerase.
Nat Commun, 9, 2018
|
|
6ENH
 
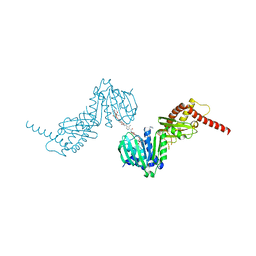 | |
9QQ6
 
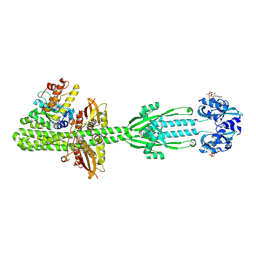 | | Structure of the Azotobacter vinelandii NifL-NifA complex | | Descriptor: | ADENOSINE-5'-DIPHOSPHATE, FLAVIN-ADENINE DINUCLEOTIDE, Nif-specific regulatory protein, ... | | Authors: | Bueno Batista, M, Richardson, J, Webster, M.W, Ghilarov, D, Peters, J.W, Lawson, D.M, Dixon, R. | | Deposit date: | 2025-03-31 | | Release date: | 2025-06-18 | | Method: | ELECTRON MICROSCOPY (6.45 Å) | | Cite: | Structural and functional analysis of the NifL-NifA complex for engineered control of nitrogen fixation in Proteobacteria
To Be Published
|
|
9BQD
 
 | | Human Topoisomerase 2 Beta ATPase domain bound to topobexin and non-hydrolyzable ATP analog AMPPNP | | Descriptor: | 8-(3,4-dihydroquinoline-1(2H)-carbonyl)-7-[2-(4-methylpiperazin-1-yl)ethoxy]-4-propyl-2H-1-benzopyran-2-one, DNA topoisomerase 2-beta, MAGNESIUM ION, ... | | Authors: | Cong, A.T.Q, Austin, C.A, Kubes, J, Karabanovich, G, Melnikova, I, Lencova, O, Kollarova, P, Piskackova, H.B, Kerestes, V, Applova, L, Arrouye, L, Alvey, J.A, Paluncic, J, Witter, T.L, Jirkovska, A, Kunes, J, Sterbova, P, Sterba, M, Simunek, T, Roh, J, Schellenberg, M.J. | | Deposit date: | 2024-05-09 | | Release date: | 2025-06-25 | | Method: | X-RAY DIFFRACTION (1.5 Å) | | Cite: | Topobexin targets the Topoisomerase II ATPase domain for beta isoform-selective inhibition and anthracycline cardioprotection.
Nat Commun, 16, 2025
|
|
9BQ6
 
 | | Human Topoisomerase 2 Alpha ATPase domain bound to non-hydrolyzable ATP analog AMPPNP | | Descriptor: | DNA topoisomerase 2-alpha, MAGNESIUM ION, PHOSPHOAMINOPHOSPHONIC ACID-ADENYLATE ESTER | | Authors: | Paluncic, J, Witter, T.L, Kubes, J, Karabanovich, G, Cong, A.T.Q, Melnikova, I, Lencova, O, Kollarova, P, Piskackova, H.B, Kerestes, V, Applova, L, Arrouye, L, Alvey, J.A, Jirkovska, A, Kunes, J, Austin, C.A, Sterbova, P, Sterba, M, Simunek, T, Roh, J, Schellenberg, M.J. | | Deposit date: | 2024-05-09 | | Release date: | 2025-06-25 | | Method: | X-RAY DIFFRACTION (1.9 Å) | | Cite: | Topobexin targets the Topoisomerase II ATPase domain for beta isoform-selective inhibition and anthracycline cardioprotection.
Nat Commun, 16, 2025
|
|
9BQ8
 
 | | Human Topoisomerase 2 Beta ATPase domain bound to non-hydrolyzable ATP analog AMPPNP | | Descriptor: | DNA topoisomerase 2-beta, MAGNESIUM ION, PHOSPHOAMINOPHOSPHONIC ACID-ADENYLATE ESTER | | Authors: | Cong, A.T.Q, Austin, C.A, Kubes, J, Karabanovich, G, Melnikova, I, Lencova, O, Kollarova, P, Piskackova, H.B, Kerestes, V, Applova, L, Arrouye, L, Alvey, J.A, Paluncic, J, Witter, T.L, Jirkovska, A, Kunes, J, Sterbova, P, Sterba, M, Simunek, T, Roh, J, Schellenberg, M.J. | | Deposit date: | 2024-05-09 | | Release date: | 2025-06-25 | | Method: | X-RAY DIFFRACTION (1.25 Å) | | Cite: | Topobexin targets the Topoisomerase II ATPase domain for beta isoform-selective inhibition and anthracycline cardioprotection.
Nat Commun, 16, 2025
|
|
9BQC
 
 | | Human Topoisomerase 2 Beta ATPase domain bound to obex 5c and non-hydrolyzable ATP analog AMPPNP | | Descriptor: | 8-(3,4-dihydroquinoline-1(2H)-carbonyl)-7-(2-hydroxyethoxy)-4-propyl-2H-1-benzopyran-2-one, DNA topoisomerase 2-beta, MAGNESIUM ION, ... | | Authors: | Cong, A.T.Q, Austin, C.A, Kubes, J, Karabanovich, G, Melnikova, I, Lencova, O, Kollarova, P, Piskackova, H.B, Kerestes, V, Applova, L, Arrouye, L, Alvey, J.A, Paluncic, J, Witter, T.L, Jirkovska, A, Kunes, J, Sterbova, P, Sterba, M, Simunek, T, Roh, J, Schellenberg, M.J. | | Deposit date: | 2024-05-09 | | Release date: | 2025-06-25 | | Method: | X-RAY DIFFRACTION (1.45 Å) | | Cite: | Topobexin targets the Topoisomerase II ATPase domain for beta isoform-selective inhibition and anthracycline cardioprotection.
Nat Commun, 16, 2025
|
|
9BQA
 
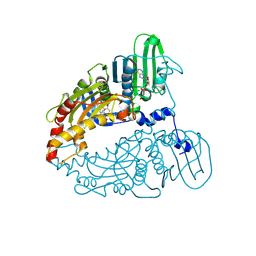 | | Human Topoisomerase 2 Beta ATPase domain bound to BNS22 and non-hydrolyzable ATP analog AMPPNP | | Descriptor: | 8-(3,4-dihydroquinoline-1(2H)-carbonyl)-5,7-dimethoxy-4-propyl-2H-1-benzopyran-2-one, DNA topoisomerase 2-beta, GLYCEROL, ... | | Authors: | Cong, A.T.Q, Austin, C.A, Kubes, J, Karabanovich, G, Melnikova, I, Lencova, O, Kollarova, P, Piskackova, H.B, Kerestes, V, Applova, L, Arrouye, L, Alvey, J.A, Paluncic, J, Witter, T.L, Jirkovska, A, Kunes, J, Sterbova, P, Sterba, M, Simunek, T, Roh, J, Schellenberg, M.J. | | Deposit date: | 2024-05-09 | | Release date: | 2025-06-25 | | Method: | X-RAY DIFFRACTION (1.9 Å) | | Cite: | Topobexin targets the Topoisomerase II ATPase domain for beta isoform-selective inhibition and anthracycline cardioprotection.
Nat Commun, 16, 2025
|
|
9BQB
 | | Human Topoisomerase 2 Alpha ATPase domain bound to topobexin and non-hydrolyzable ATP analog AMPPNP | | Descriptor: | 8-(3,4-dihydroquinoline-1(2H)-carbonyl)-7-[2-(4-methylpiperazin-1-yl)ethoxy]-4-propyl-2H-1-benzopyran-2-one, CHLORIDE ION, DNA topoisomerase 2-alpha, ... | | Authors: | Cong, A.T.Q, Austin, C.A, Kubes, J, Karabanovich, G, Melnikova, I, Lencova, O, Kollarova, P, Piskackova, H.B, Kerestes, V, Applova, L, Arrouye, L, Alvey, J.A, Paluncic, J, Witter, T.L, Jirkovska, A, Kunes, J, Sterbova, P, Sterba, M, Simunek, T, Roh, J, Schellenberg, M.J. | | Deposit date: | 2024-05-09 | | Release date: | 2025-06-25 | | Method: | X-RAY DIFFRACTION (1.5 Å) | | Cite: | Topobexin targets the Topoisomerase II ATPase domain for beta isoform-selective inhibition and anthracycline cardioprotection.
Nat Commun, 16, 2025
|
|
9BQ7
 
 | | Human Topoisomerase 2 Alpha ATPase domain bound to BNS22 and non-hydrolyzable ATP analog AMPPNP | | Descriptor: | 8-(3,4-dihydroquinoline-1(2H)-carbonyl)-5,7-dimethoxy-4-propyl-2H-1-benzopyran-2-one, CHLORIDE ION, DNA topoisomerase 2-alpha, ... | | Authors: | Cong, A.T.Q, Austin, C.A, Kubes, J, Karabanovich, G, Melnikova, I, Lencova, O, Kollarova, P, Piskackova, H.B, Kerestes, V, Applova, L, Arrouye, L, Alvey, J.A, Paluncic, J, Witter, T.L, Jirkovska, A, Kunes, J, Sterbova, P, Sterba, M, Simunek, T, Roh, J, Schellenberg, M.J. | | Deposit date: | 2024-05-09 | | Release date: | 2025-06-25 | | Method: | X-RAY DIFFRACTION (2.05 Å) | | Cite: | Topobexin targets the Topoisomerase II ATPase domain for beta isoform-selective inhibition and anthracycline cardioprotection.
Nat Commun, 16, 2025
|
|
9BQ9
 
 | | Human Topoisomerase 2 Alpha ATPase domain bound to obex 5c and non-hydrolyzable ATP analog AMPPNP | | Descriptor: | 8-(3,4-dihydroquinoline-1(2H)-carbonyl)-7-(2-hydroxyethoxy)-4-propyl-2H-1-benzopyran-2-one, CHLORIDE ION, DNA topoisomerase 2-alpha, ... | | Authors: | Cong, A.T.Q, Kubes, J, Karabanovich, G, Melnikova, I, Lencova, O, Kollarova, P, Piskackova, H.B, Kerestes, V, Applova, L, Arrouye, L, Alvey, J.A, Paluncic, J, Witter, T.L, Jirkovska, A, Kunes, J, Sterbova, P, Austin, C.A, Sterba, M, Simunek, T, Roh, J, Schellenberg, M.J. | | Deposit date: | 2024-05-09 | | Release date: | 2025-06-25 | | Method: | X-RAY DIFFRACTION (1.95 Å) | | Cite: | Topobexin targets the Topoisomerase II ATPase domain for beta isoform-selective inhibition and anthracycline cardioprotection.
Nat Commun, 16, 2025
|
|
8F5Z
 
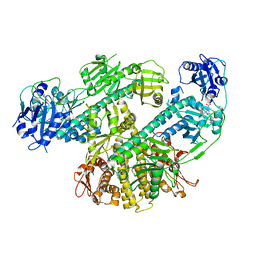 | | Composite map of CryoEM structure of Arabidopsis thaliana phytochrome A | | Descriptor: | 3-[5-[[(3~{R},4~{R})-3-ethyl-4-methyl-5-oxidanylidene-3,4-dihydropyrrol-2-yl]methyl]-2-[[5-[(4-ethyl-3-methyl-5-oxidanylidene-pyrrol-2-yl)methyl]-3-(3-hydroxy-3-oxopropyl)-4-methyl-1~{H}-pyrrol-2-yl]methyl]-4-methyl-1~{H}-pyrrol-3-yl]propanoic acid, Phytochrome A | | Authors: | Li, H, Li, H. | | Deposit date: | 2022-11-15 | | Release date: | 2023-06-28 | | Last modified: | 2024-10-23 | | Method: | ELECTRON MICROSCOPY (3.2 Å) | | Cite: | The structure of Arabidopsis phytochrome A reveals topological and functional diversification among the plant photoreceptor isoforms.
Nat.Plants, 9, 2023
|
|
8YB4
 | | Pfr conformer of Arabidopsis thaliana phytochrome B in complex with phytochrome-interacting factor 6 | | Descriptor: | 3-[5-[[(3~{R},4~{R})-3-ethyl-4-methyl-5-oxidanylidene-3,4-dihydropyrrol-2-yl]methyl]-2-[[5-[(4-ethyl-3-methyl-5-oxidanylidene-pyrrol-2-yl)methyl]-3-(3-hydroxy-3-oxopropyl)-4-methyl-1~{H}-pyrrol-2-yl]methyl]-4-methyl-1~{H}-pyrrol-3-yl]propanoic acid, phytochrome B, phytochrome-interacting factor 6 | | Authors: | Wang, Z, Wang, W, Zhao, D, Song, Y, Xu, B, Zhao, J, Wang, J. | | Deposit date: | 2024-02-11 | | Release date: | 2024-10-02 | | Last modified: | 2024-11-20 | | Method: | ELECTRON MICROSCOPY (3.1 Å) | | Cite: | Light-induced remodeling of phytochrome B enables signal transduction by phytochrome-interacting factor.
Cell, 187, 2024
|
|
8YO4
 
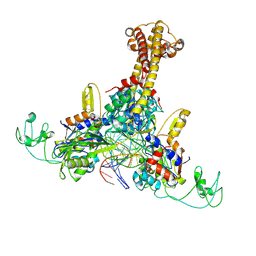 | | structure of phage T4 topoisomerase II central domain bound with DNA | | Descriptor: | DNA (5'-D(P*AP*TP*AP*TP*AP*TP*GP*TP*GP*TP*AP*TP*AP*TP*AP*TP*AP*CP*AP*CP*AP*CP*AP*T)-3'), DNA (5'-D(P*TP*GP*TP*GP*TP*GP*TP*AP*TP*AP*TP*AP*TP*AP*CP*AP*CP*AP*TP*AP*TP*AP*TP*A)-3'), DNA topoisomerase medium subunit, ... | | Authors: | Chen, Y.T, Xin, Y.H, Xian, R.Q. | | Deposit date: | 2024-03-12 | | Release date: | 2024-09-25 | | Method: | ELECTRON MICROSCOPY (3.2 Å) | | Cite: | structure of phage T4 topoisomerase II central domain bound with DNA
To be published
|
|
8YO5
 
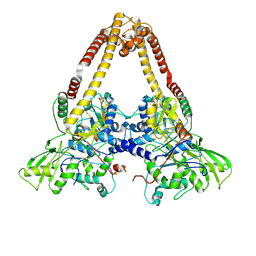 | |
8YON
 
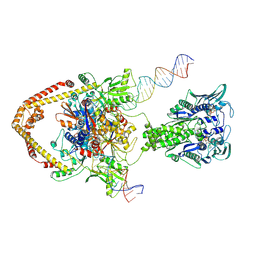 | |
8YO3
 
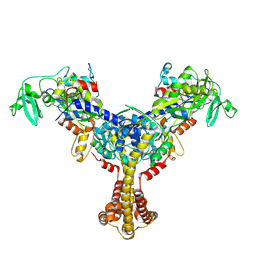 | |
8YOD
 
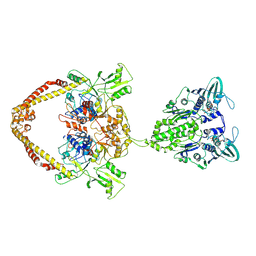 | |
8YO1
 
 | |
8YO7
 
 | |
8YLU
 
 | | structure of phage T6 topoisomerase II central domain bound with DNA | | Descriptor: | DNA (5'-D(P*TP*AP*TP*AP*TP*GP*TP*GP*TP*AP*TP*AP*TP*AP*TP*AP*CP*AP*CP*AP*CP*A)-3'), DNA topoisomerase (ATP-hydrolyzing), DNA topoisomerase medium subunit, ... | | Authors: | Chen, Y.T, Xin, Y.H. | | Deposit date: | 2024-03-06 | | Release date: | 2024-09-25 | | Method: | ELECTRON MICROSCOPY (2.8 Å) | | Cite: | structure of phage T6 topoisomerase II central domain bound with DNA
To be published
|
|
9EUY
 
 | | Cryo-EM structure of the full-length Pseudomonas aeruginosa bacteriophytochrome in its Pfr state | | Descriptor: | 3-[2-[(Z)-[3-(2-carboxyethyl)-5-[(Z)-(4-ethenyl-3-methyl-5-oxidanylidene-pyrrol-2-ylidene)methyl]-4-methyl-pyrrol-1-ium -2-ylidene]methyl]-5-[(Z)-[(3E)-3-ethylidene-4-methyl-5-oxidanylidene-pyrrolidin-2-ylidene]methyl]-4-methyl-1H-pyrrol-3- yl]propanoic acid, Bacteriophytochrome | | Authors: | Bodizs, S, Westenhoff, S. | | Deposit date: | 2024-03-28 | | Release date: | 2024-09-04 | | Last modified: | 2024-11-20 | | Method: | ELECTRON MICROSCOPY (2.95 Å) | | Cite: | Cryo-EM structures of a bathy phytochrome histidine kinase reveal a unique light-dependent activation mechanism.
Structure, 32, 2024
|
|
9EUT
 
 | | Cryo-EM structure of the full-length Pseudomonas aeruginosa bacteriophytochrome in its Pr state | | Descriptor: | 3-[2-[(Z)-[3-(2-carboxyethyl)-5-[(Z)-(4-ethenyl-3-methyl-5-oxidanylidene-pyrrol-2-ylidene)methyl]-4-methyl-pyrrol-1-ium -2-ylidene]methyl]-5-[(Z)-[(3E)-3-ethylidene-4-methyl-5-oxidanylidene-pyrrolidin-2-ylidene]methyl]-4-methyl-1H-pyrrol-3- yl]propanoic acid, Bacteriophytochrome | | Authors: | Bodizs, S, Westenhoff, S. | | Deposit date: | 2024-03-28 | | Release date: | 2024-09-04 | | Last modified: | 2024-11-20 | | Method: | ELECTRON MICROSCOPY (2.95 Å) | | Cite: | Cryo-EM structures of a bathy phytochrome histidine kinase reveal a unique light-dependent activation mechanism.
Structure, 32, 2024
|
|
