8VWH
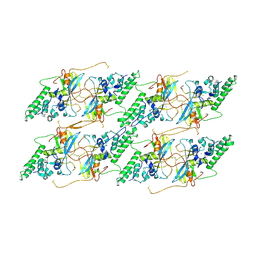 
 | | Structure of the baculovirus major nucleocapsid protein VP39 (localised reconstruction) | | Descriptor: | Major capsid protein, ZINC ION | | Authors: | Johnstone, B.A, Hardy, J.M, Ha, J.H, Venugopal, H, Coulibaly, F. | | Deposit date: | 2024-02-01 | | Release date: | 2024-11-06 | | Last modified: | 2025-05-21 | | Method: | ELECTRON MICROSCOPY (3.06 Å) | | Cite: | The nucleocapsid architecture and structural atlas of the prototype baculovirus define the hallmarks of a new viral realm.
Sci Adv, 10, 2024
|
|
7LXS
 
 | |
5ILB
 
 | | Crystal structure of protease domain of Deg2 linked with the PDZ domain of Deg9 | | Descriptor: | Protease Do-like 2, chloroplastic,Protease Do-like 9 | | Authors: | Ouyang, M, Liu, L, Li, X.Y, Zhao, S, Zhang, L.X. | | Deposit date: | 2016-03-04 | | Release date: | 2017-03-08 | | Last modified: | 2023-11-08 | | Method: | X-RAY DIFFRACTION (1.852 Å) | | Cite: | The crystal structure of Deg9 reveals a novel octameric-type HtrA protease
Nat Plants, 3, 2017
|
|
8VJD
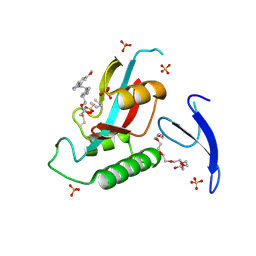 
 | | Human R14A Pin1 covalently bound to inhibitor 158F10 in P21 21 21 space group | | Descriptor: | 3,6,9,12,15,18-HEXAOXAICOSANE-1,20-DIOL, Inhibitor 158F10, Peptidyl-prolyl cis-trans isomerase NIMA-interacting 1, ... | | Authors: | Rodriguez, I, Blaha, G. | | Deposit date: | 2024-01-06 | | Release date: | 2024-11-06 | | Last modified: | 2025-03-05 | | Method: | X-RAY DIFFRACTION (1.57 Å) | | Cite: | Targeted degradation of Pin1 by protein-destabilizing compounds.
Proc.Natl.Acad.Sci.USA, 121, 2024
|
|
6VSE
 
 | |
8VIT
 
 | | Crystal structure of the N-terminal domain of fatty acid kinase A (FakA) from Staphylococcus aureus (Mg and ADP) | | Descriptor: | ADENOSINE-5'-DIPHOSPHATE, FATTY ACID KINASE A, MAGNESIUM ION, ... | | Authors: | Lovell, S, Kashipathy, M.M, Battaile, K.P, Bose, J.L. | | Deposit date: | 2024-01-05 | | Release date: | 2024-11-06 | | Last modified: | 2024-12-04 | | Method: | X-RAY DIFFRACTION (1.25 Å) | | Cite: | Molecular insights into the structure and function of the Staphylococcus aureus fatty acid kinase.
J.Biol.Chem., 300, 2024
|
|
5ILV
 
 | | Uninhibited ETV5 | | Descriptor: | ETS translocation variant 5 | | Authors: | Whitby, F.G, Currie, S.L. | | Deposit date: | 2016-03-04 | | Release date: | 2017-02-22 | | Last modified: | 2024-11-13 | | Method: | X-RAY DIFFRACTION (1.8 Å) | | Cite: | Structured and disordered regions cooperatively mediate DNA-binding autoinhibition of ETS factors ETV1, ETV4 and ETV5.
Nucleic Acids Res., 45, 2017
|
|
5VP1
 
 | | Discovery of Clinical Candidate N-{(1S)-1-[3-Fluoro-4-(trifluoromethoxy)phenyl]-2-methoxyethyl}-7-methoxy-2-oxo-2,3-dihydropyrido[2,3-b]pyrazine-4(1H)-carboxamide (TAK-915), A Highly Potent, Selective, and Brain-Penetrating Phosphodiesterase 2A Inhibitor for the Treatment of Cognitive Disorders | | Descriptor: | MAGNESIUM ION, N-{(1S)-2-hydroxy-2-methyl-1-[4-(trifluoromethoxy)phenyl]propyl}-6-methyl-5-(3-methyl-1H-1,2,4-triazol-1-yl)pyrazolo[1,5-a]pyrimidine-3-carboxamide, ZINC ION, ... | | Authors: | Hoffman, I.D. | | Deposit date: | 2017-05-03 | | Release date: | 2017-12-27 | | Last modified: | 2024-03-13 | | Method: | X-RAY DIFFRACTION (1.86 Å) | | Cite: | Discovery of a Novel Series of Pyrazolo[1,5-a]pyrimidine-Based Phosphodiesterase 2A Inhibitors Structurally Different from N-((1S)-1-(3-Fluoro-4-(trifluoromethoxy)phenyl)-2-methoxyethyl)-7-methoxy-2-oxo-2,3-dihydropyrido[2,3-b]pyrazine-4(1H)-carboxamide (TAK-915), for the Treatment of Cognitive Disorders.
Chem. Pharm. Bull., 65, 2017
|
|
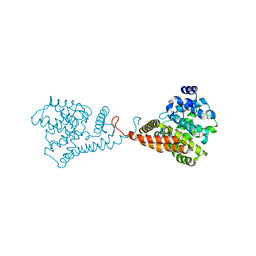
8VWI
 
 | | The base complex of the AcMNPV baculovirus nucleocapsid (Class 1, localised reconstruction) | | Descriptor: | 38K (AC98) protein in P143-LEF5 intergenic region, Capsid-associated protein VP80, Major capsid protein, ... | | Authors: | Johnstone, B.A, Koszalka, P, Ha, J, Venugopal, H, Coulibaly, F. | | Deposit date: | 2024-02-01 | | Release date: | 2024-11-06 | | Last modified: | 2024-11-27 | | Method: | ELECTRON MICROSCOPY (4.71 Å) | | Cite: | The baculovirus structure defines the hallmarks of a new viral realm
To Be Published
|
|
5IM2
 
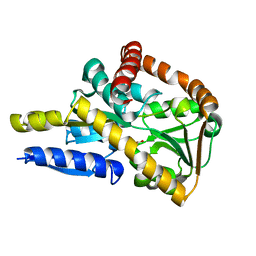 | | Crystal structure of a TRAP solute binding protein from Rhodoferax ferrireducens T118 (Rfer_2570, TARGET EFI-510210) in complex with copurified benzoate | | Descriptor: | BENZOIC ACID, Twin-arginine translocation pathway signal | | Authors: | Vetting, M.W, Al Obaidi, N.F, Morisco, L.L, Benach, J, Koss, J, Wasserman, S.R, Gerlt, J.A, Almo, S.C, Enzyme Function Initiative (EFI) | | Deposit date: | 2016-03-05 | | Release date: | 2016-03-23 | | Last modified: | 2023-09-27 | | Method: | X-RAY DIFFRACTION (1.7 Å) | | Cite: | Crystal structure of a TRAP solute binding protein from Rhodoferax ferrireducens T118 (Rfer_2570, TARGET EFI-510210) in complex with copurified benzoate
To be published
|
|
5VPC
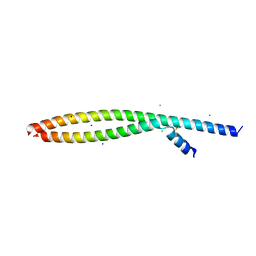 
 | | Transcription factor FosB/JunD bZIP domain in its oxidized form, type-II crystal | | Descriptor: | CHLORIDE ION, Protein fosB, SODIUM ION, ... | | Authors: | Yin, Z, Machius, M.C, Rudenko, G. | | Deposit date: | 2017-05-04 | | Release date: | 2017-09-06 | | Last modified: | 2024-10-30 | | Method: | X-RAY DIFFRACTION (2.498 Å) | | Cite: | Activator Protein-1: redox switch controlling structure and DNA-binding.
Nucleic Acids Res., 45, 2017
|
|
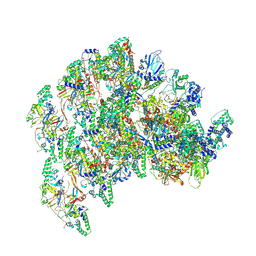
8VPQ
 
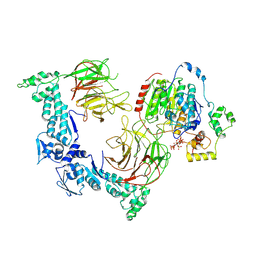 | | The structure of LSD1-CoREST-HDAC1 in complex with KBTBD4IPR310delinsTTYML | | Descriptor: | Histone deacetylase 1, INOSITOL HEXAKISPHOSPHATE, Isoform 2 of Kelch repeat and BTB domain-containing protein 4, ... | | Authors: | Xie, X, Liau, B, Zheng, N. | | Deposit date: | 2024-01-16 | | Release date: | 2024-11-27 | | Last modified: | 2025-06-11 | | Method: | ELECTRON MICROSCOPY (3.3 Å) | | Cite: | Converging mechanism of UM171 and KBTBD4 neomorphic cancer mutations.
Nature, 639, 2025
|
|
6VSO
 
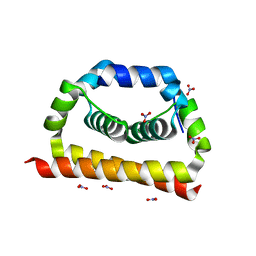 | | DengueV-2 Capsid Structure | | Descriptor: | Capsid premembrane protein, GLYCEROL, NITRATE ION | | Authors: | White, M, Xia, H, Shi, P. | | Deposit date: | 2020-02-11 | | Release date: | 2020-07-15 | | Last modified: | 2023-10-11 | | Method: | X-RAY DIFFRACTION (3.001 Å) | | Cite: | A cocrystal structure of dengue capsid protein in complex of inhibitor.
Proc.Natl.Acad.Sci.USA, 117, 2020
|
|
5IC5
 
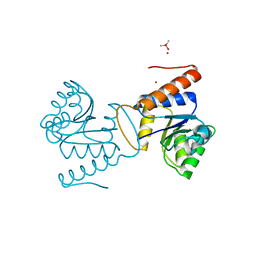 | | Bacteriophytochrome response regulator RtBRR | | Descriptor: | CACODYLATE ION, Candidate response regulator, CheY, ... | | Authors: | Baker, A.W, Satyshur, K.A, Forest, K.T. | | Deposit date: | 2016-02-22 | | Release date: | 2016-03-02 | | Last modified: | 2024-03-06 | | Method: | X-RAY DIFFRACTION (1.9 Å) | | Cite: | Arm-in-Arm Response Regulator Dimers Promote Intermolecular Signal Transduction.
J.Bacteriol., 198, 2016
|
|
5VPH
 
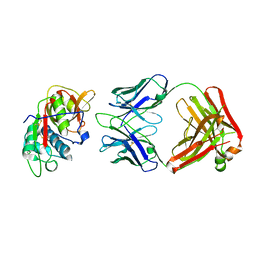 | | CRYSTAL STRUCTURE OF DER P 1 COMPLEXED WITH FAB 4C1 | | Descriptor: | 1,2-ETHANEDIOL, CALCIUM ION, Der p 1 allergen, ... | | Authors: | Chruszcz, M, Vailes, L.D, Chapman, M.D, Pomes, A, Minor, W. | | Deposit date: | 2017-05-05 | | Release date: | 2017-05-24 | | Last modified: | 2024-10-16 | | Method: | X-RAY DIFFRACTION (2.5 Å) | | Cite: | Molecular Determinants For Antibody Binding On Group 1 House Dust Mite Allergens.
J.Biol.Chem., 287, 2012
|
|
5VMK
 
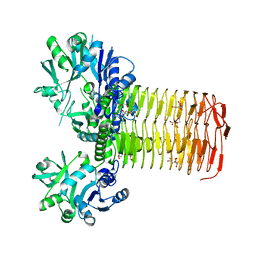 | |
8VJG
 
 | | Human R14A Pin1 covalently bound to inhibitor 164A10 | | Descriptor: | 2,3-DIHYDROXY-1,4-DITHIOBUTANE, 3,6,9,12,15,18-HEXAOXAICOSANE-1,20-DIOL, Nalpha-{(2S)-1-[(3S)-2-acetyl-2,3,4,9-tetrahydro-1H-pyrido[3,4-b]indole-3-carbonyl]piperidine-2-carbonyl}-5-fluoro-L-tryptophanamide, ... | | Authors: | Rodriguez, I, Blaha, G. | | Deposit date: | 2024-01-06 | | Release date: | 2024-11-06 | | Last modified: | 2025-03-05 | | Method: | X-RAY DIFFRACTION (1.58 Å) | | Cite: | Targeted degradation of Pin1 by protein-destabilizing compounds.
Proc.Natl.Acad.Sci.USA, 121, 2024
|
|
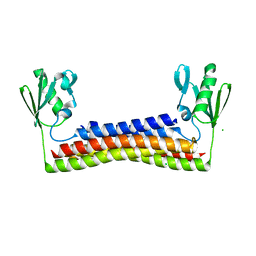
6VUD
 
 | |
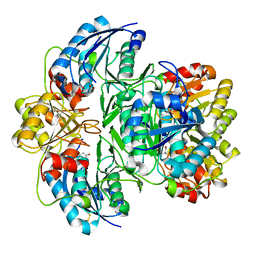
7M11
 
 | |
6VUY
 
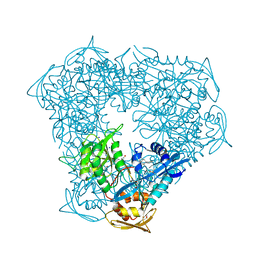 | | Crystal structure of Eis from Mycobacterium tuberculosis in complex with inhibitor SGT358 | | Descriptor: | (7S)-7-phenyl-2-{[3-(piperidin-1-yl)propyl]sulfanyl}-5,6,7,8-tetrahydro[1]benzothieno[2,3-d]pyrimidin-4-amine, DI(HYDROXYETHYL)ETHER, DIMETHYL SULFOXIDE, ... | | Authors: | Punetha, A, Hou, C, Ngo, H.X, Garneau-Tsodikova, S, Tsodikov, O.V. | | Deposit date: | 2020-02-16 | | Release date: | 2020-06-03 | | Last modified: | 2023-10-11 | | Method: | X-RAY DIFFRACTION (2.7 Å) | | Cite: | Structure-Guided Optimization of Inhibitors of Acetyltransferase Eis fromMycobacterium tuberculosis.
Acs Chem.Biol., 15, 2020
|
|
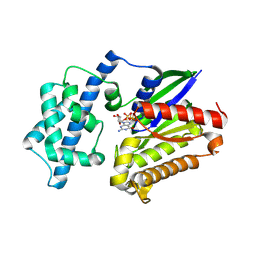
8VGA
 | |
5VPY
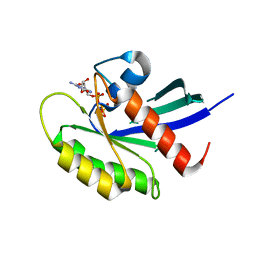 
 | | Crystal structure of human KRAS G12A mutant in complex with GppNHp | | Descriptor: | 2-amino-9-{5-O-[(S)-hydroxy{[(R)-hydroxy(phosphonoamino)phosphoryl]oxy}phosphoryl]-alpha-L-xylofuranosyl}-1,9-dihydro-6H-purin-6-one, GTPase KRas, MAGNESIUM ION, ... | | Authors: | Xu, S, Long, B, Boris, G, Ni, S, Kennedy, M.A. | | Deposit date: | 2017-05-06 | | Release date: | 2017-12-06 | | Last modified: | 2024-03-13 | | Method: | X-RAY DIFFRACTION (2 Å) | | Cite: | Structural insight into the rearrangement of the switch I region in GTP-bound G12A K-Ras.
Acta Crystallogr D Struct Biol, 73, 2017
|
|
8VRT
 
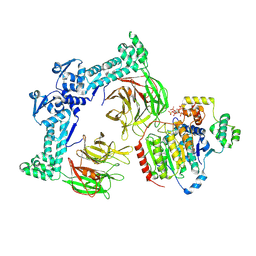 | | The structure of LSD1-CoREST-HDAC1 in complex with KBTBD4R313PRR mutant | | Descriptor: | Histone deacetylase 1, INOSITOL HEXAKISPHOSPHATE, Kelch repeat and BTB domain-containing protein 4, ... | | Authors: | Xie, X, Liau, B, Zheng, N. | | Deposit date: | 2024-01-22 | | Release date: | 2024-11-27 | | Last modified: | 2025-06-11 | | Method: | ELECTRON MICROSCOPY (3.42 Å) | | Cite: | Converging mechanism of UM171 and KBTBD4 neomorphic cancer mutations.
Nature, 639, 2025
|
|
8VWK
 
 | | Crystal Structure of a fatty acid decarboxylase from Kocuria marina in complex with myristic acid | | Descriptor: | Cytochrome P450, DI(HYDROXYETHYL)ETHER, GLYCEROL, ... | | Authors: | Generoso, W.C, Miyamoto, R.Y, Murakami, M.T, Zanphorlin, L.M. | | Deposit date: | 2024-02-01 | | Release date: | 2024-12-04 | | Last modified: | 2025-04-02 | | Method: | X-RAY DIFFRACTION (2.05 Å) | | Cite: | Coordinated conformational changes in P450 decarboxylases enable hydrocarbons production from renewable feedstocks.
Nat Commun, 16, 2025
|
|
5VQ2
 
 | | Crystal structure of human WT-KRAS in complex with GTP | | Descriptor: | GTPase KRas, GUANOSINE-5'-TRIPHOSPHATE, MAGNESIUM ION | | Authors: | Xu, S, Long, B, Boris, G, Ni, S, Kennedy, M.A. | | Deposit date: | 2017-05-07 | | Release date: | 2017-12-06 | | Last modified: | 2024-10-16 | | Method: | X-RAY DIFFRACTION (1.96 Å) | | Cite: | Structural insight into the rearrangement of the switch I region in GTP-bound G12A K-Ras.
Acta Crystallogr D Struct Biol, 73, 2017
|
|
