7JWF
 
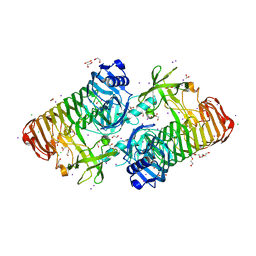 | | Crystal structure of PdGH110B D344N in complex with alpha-(1,3)-galactobiose | | Descriptor: | 1,2-ETHANEDIOL, 4-(2-HYDROXYETHYL)-1-PIPERAZINE ETHANESULFONIC ACID, ACETATE ION, ... | | Authors: | Hettle, A.G, Boraston, A.B. | | Deposit date: | 2020-08-25 | | Release date: | 2020-11-04 | | Last modified: | 2023-10-18 | | Method: | X-RAY DIFFRACTION (2.187 Å) | | Cite: | The structure of a family 110 glycoside hydrolase provides insight into the hydrolysis of alpha-1,3-galactosidic linkages in lambda-carrageenan and blood group antigens.
J.Biol.Chem., 295, 2020
|
|
7JW4
 
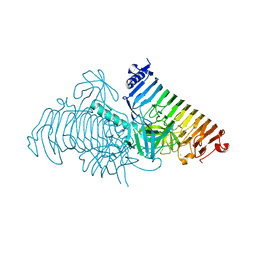 | | Crystal structure of PdGH110B in complex with D-galactose | | Descriptor: | CHLORIDE ION, Glycoside hydrolase family 110, NICKEL (II) ION, ... | | Authors: | Hettle, A.G, Boraston, A.B. | | Deposit date: | 2020-08-24 | | Release date: | 2020-11-04 | | Last modified: | 2024-04-03 | | Method: | X-RAY DIFFRACTION (2.342 Å) | | Cite: | The structure of a family 110 glycoside hydrolase provides insight into the hydrolysis of alpha-1,3-galactosidic linkages in lambda-carrageenan and blood group antigens.
J.Biol.Chem., 295, 2020
|
|
5KKT
 
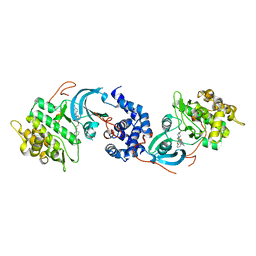 | | ROCK 1 bound to azaindole thiazole piperazine inhibitor | | Descriptor: | 2-[3-[3-(4-methylpiperazin-1-yl)propoxy]phenyl]-~{N}-[4-(1~{H}-pyrrolo[2,3-b]pyridin-3-yl)-1,3-thiazol-2-yl]ethanamide, Rho-associated protein kinase 1 | | Authors: | Jacobs, M.D. | | Deposit date: | 2016-06-22 | | Release date: | 2017-06-28 | | Last modified: | 2023-09-27 | | Method: | X-RAY DIFFRACTION (2.8 Å) | | Cite: | ROCK inhibitors 3: Design, synthesis and structure-activity relationships of 7-azaindole-based Rho kinase (ROCK) inhibitors
Bioorg.Med.Chem.Lett., 2018
|
|
5KKS
 
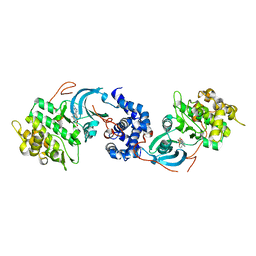 | | ROCK 1 bound to azaindole thiazole inhibitor | | Descriptor: | 2-[3-(methylsulfonylamino)phenyl]-~{N}-[4-(1~{H}-pyrrolo[2,3-b]pyridin-3-yl)-1,3-thiazol-2-yl]ethanamide, Rho-associated protein kinase 1 | | Authors: | Jacobs, M.D. | | Deposit date: | 2016-06-22 | | Release date: | 2017-06-28 | | Last modified: | 2023-09-27 | | Method: | X-RAY DIFFRACTION (3.3 Å) | | Cite: | ROCK inhibitors 3: Design, synthesis and structure-activity relationships of 7-azaindole-based Rho kinase (ROCK) inhibitors
Bioorg.Med.Chem.Lett., 2018
|
|
2UZJ
 
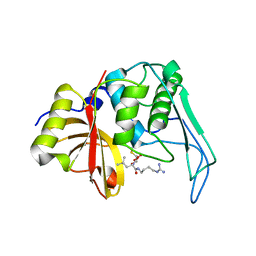 | | Crystal structure of the mature streptococcal cysteine protease, mSpeB | | Descriptor: | N-[N-[1-HYDROXYCARBOXYETHYL-CARBONYL]LEUCYLAMINO-BUTYL]-GUANIDINE, STREPTOPAIN | | Authors: | Olsen, J.G, Dagil, R, Niclasen, L.M, Soerensen, O.E, Kragelund, B.B. | | Deposit date: | 2008-09-16 | | Release date: | 2009-09-08 | | Last modified: | 2024-10-09 | | Method: | X-RAY DIFFRACTION (1.55 Å) | | Cite: | Structure of the Mature Streptococcal Cysteine Protease Exotoxin Mspeb in its Active Dimeric Form.
J.Mol.Biol., 393, 2009
|
|
9F2K
 
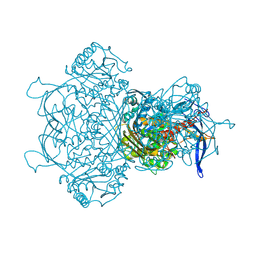 | | Myo-inositol-1-phosphate synthase from Thermochaetoides thermophila in complex with NAD | | Descriptor: | NICOTINAMIDE-ADENINE-DINUCLEOTIDE, inositol-3-phosphate synthase | | Authors: | Traeger, T.K, Kyrilis, F.L, Hamdi, F, Kastritis, P.L. | | Deposit date: | 2024-04-23 | | Release date: | 2024-08-14 | | Last modified: | 2024-08-28 | | Method: | ELECTRON MICROSCOPY (2.48 Å) | | Cite: | Disorder-to-order active site capping regulates the rate-limiting step of the inositol pathway.
Proc.Natl.Acad.Sci.USA, 121, 2024
|
|
5BUH
 
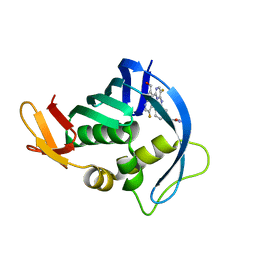 | | Influenza PB2 bound to a hydroxymethyl azaindole inhibitor | | Descriptor: | N-[(1R,3S)-3-({5-fluoro-2-[5-fluoro-2-(hydroxymethyl)-1H-pyrrolo[2,3-b]pyridin-3-yl]pyrimidin-4-yl}amino)cyclohexyl]pyrrolidine-1-carboxamide, Polymerase basic protein 2 | | Authors: | Jacobs, M.D. | | Deposit date: | 2015-06-03 | | Release date: | 2016-06-08 | | Last modified: | 2024-03-06 | | Method: | X-RAY DIFFRACTION (2.55 Å) | | Cite: | Novel 2-Substituted 7-Azaindole and 7-Azaindazole Analogues as Potential Antiviral Agents for the Treatment of Influenza.
ACS Med Chem Lett, 8, 2017
|
|
5VGP
 
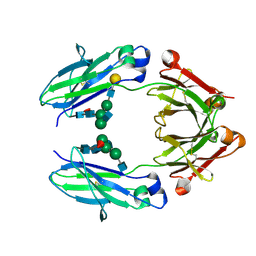 | | Fc fragment of human IgG1 antibody, from NIST mAb | | Descriptor: | 2-acetamido-2-deoxy-beta-D-glucopyranose-(1-2)-alpha-D-mannopyranose-(1-3)-[2-acetamido-2-deoxy-beta-D-glucopyranose-(1-2)-alpha-D-mannopyranose-(1-6)]beta-D-mannopyranose-(1-4)-2-acetamido-2-deoxy-beta-D-glucopyranose-(1-4)-[alpha-L-fucopyranose-(1-6)]2-acetamido-2-deoxy-beta-D-glucopyranose, Human Fc fragment with G1F/G0F glycan, beta-D-galactopyranose-(1-4)-2-acetamido-2-deoxy-beta-D-glucopyranose-(1-2)-alpha-D-mannopyranose-(1-6)-[2-acetamido-2-deoxy-beta-D-glucopyranose-(1-2)-alpha-D-mannopyranose-(1-3)]beta-D-mannopyranose-(1-4)-2-acetamido-2-deoxy-beta-D-glucopyranose-(1-4)-[alpha-L-fucopyranose-(1-6)]2-acetamido-2-deoxy-beta-D-glucopyranose | | Authors: | Gallagher, D.T, Galvin, C.V, Karageorgos, I, Marino, J.P. | | Deposit date: | 2017-04-11 | | Release date: | 2018-04-18 | | Last modified: | 2023-10-04 | | Method: | X-RAY DIFFRACTION (2.116 Å) | | Cite: | Structure of the Fc fragment of the NIST reference antibody RM8671.
Acta Crystallogr F Struct Biol Commun, 74, 2018
|
|
7KKE
 
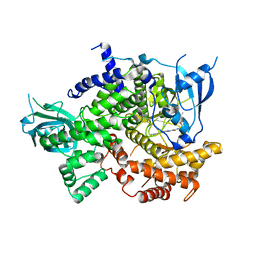 | | Phosphoinositide 3-Kinase gamma bound to a thiazole inhibitor | | Descriptor: | N-[2-(3,3-dimethylbutoxy)ethyl]-N'-{4-methyl-5-[(pyridin-4-yl)ethynyl]-1,3-thiazol-2-yl}urea, Phosphatidylinositol 4,5-bisphosphate 3-kinase catalytic subunit gamma isoform | | Authors: | Jacobs, M.D, Griffith, J.P. | | Deposit date: | 2020-10-27 | | Release date: | 2021-03-24 | | Last modified: | 2023-10-18 | | Method: | X-RAY DIFFRACTION (2.81 Å) | | Cite: | Discovery of a Novel Series of Potent and Selective Alkynylthiazole-Derived PI3K gamma Inhibitors.
Acs Med.Chem.Lett., 12, 2021
|
|
1FGB
 
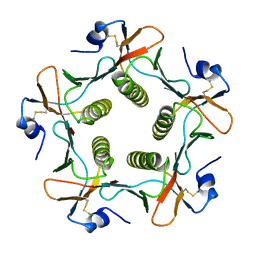 | | TOXIN | | Descriptor: | CHOLERA TOXIN B SUBUNIT PENTAMER | | Authors: | Zhang, R.-G, Westbrook, E. | | Deposit date: | 1996-02-21 | | Release date: | 1996-12-23 | | Last modified: | 2024-06-05 | | Method: | X-RAY DIFFRACTION (2.4 Å) | | Cite: | The 2.4 A crystal structure of cholera toxin B subunit pentamer: choleragenoid.
J.Mol.Biol., 251, 1995
|
|
7F2D
 
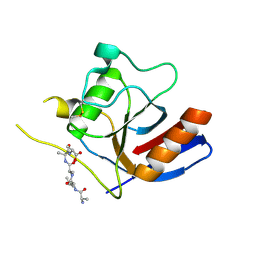 | |
7F2I
 
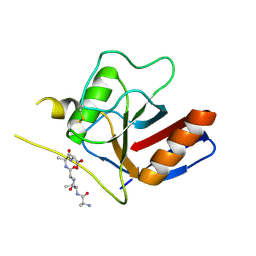 | |
5UZK
 | | Crystal Structure of PKA bound to an pyrrolo pyridine inhibitor | | Descriptor: | 2-{3-[3-(piperidin-4-yl)propoxy]phenyl}-N-[4-(1H-pyrrolo[2,3-b]pyridin-3-yl)-1,3-thiazol-2-yl]acetamide, cAMP-dependent protein kinase catalytic subunit alpha, cAMP-dependent protein kinase inhibitor alpha | | Authors: | Jacobs, M.D, Brown, K. | | Deposit date: | 2017-02-26 | | Release date: | 2018-03-07 | | Last modified: | 2023-10-04 | | Method: | X-RAY DIFFRACTION (2.3 Å) | | Cite: | ROCK inhibitors 3: Design, synthesis and structure-activity relationships of 7-azaindole-based Rho kinase (ROCK) inhibitors.
Bioorg. Med. Chem. Lett., 28, 2018
|
|
5UZJ
 
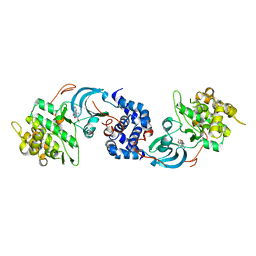 | | Crystal Structure of ROCK1 bound to an aminopyridine inhibitor | | Descriptor: | N-[4-(2-aminopyridin-4-yl)-1,3-thiazol-2-yl]-2-(3-methoxyphenyl)acetamide, Rho-associated protein kinase 1 | | Authors: | Jacobs, M.D. | | Deposit date: | 2017-02-26 | | Release date: | 2018-03-07 | | Last modified: | 2023-10-04 | | Method: | X-RAY DIFFRACTION (3.3 Å) | | Cite: | ROCK inhibitors 3: Design, synthesis and structure-activity relationships of 7-azaindole-based Rho kinase (ROCK) inhibitors.
Bioorg. Med. Chem. Lett., 28, 2018
|
|
5O5W
 
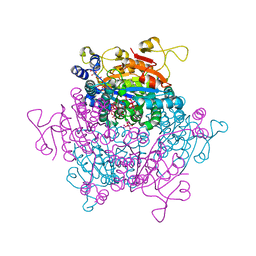 | | Molybdenum storage protein room-temperature structure determined by serial millisecond crystallography | | Descriptor: | (mu3-oxo)-tris(mu2-oxo)-nonakisoxo-trimolybdenum (VI), ADENOSINE-5'-TRIPHOSPHATE, MAGNESIUM ION, ... | | Authors: | Steffen, B, Weinert, T, Ermler, U, Standfuss, J. | | Deposit date: | 2017-06-02 | | Release date: | 2017-09-27 | | Last modified: | 2024-01-17 | | Method: | X-RAY DIFFRACTION (1.7 Å) | | Cite: | Serial millisecond crystallography for routine room-temperature structure determination at synchrotrons.
Nat Commun, 8, 2017
|
|
1DKZ
 
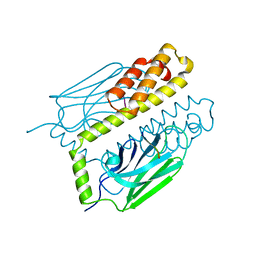 | | THE SUBSTRATE BINDING DOMAIN OF DNAK IN COMPLEX WITH A SUBSTRATE PEPTIDE, DETERMINED FROM TYPE 1 NATIVE CRYSTALS | | Descriptor: | SUBSTRATE BINDING DOMAIN OF DNAK, SUBSTRATE PEPTIDE (7 RESIDUES) | | Authors: | Zhu, X, Zhao, X, Burkholder, W.F, Gragerov, A, Ogata, C.M, Gottesman, M.E, Hendrickson, W.A. | | Deposit date: | 1996-06-03 | | Release date: | 1996-12-07 | | Last modified: | 2024-02-07 | | Method: | X-RAY DIFFRACTION (2 Å) | | Cite: | Structural analysis of substrate binding by the molecular chaperone DnaK.
Science, 272, 1996
|
|
1DKX
 | | THE SUBSTRATE BINDING DOMAIN OF DNAK IN COMPLEX WITH A SUBSTRATE PEPTIDE, DETERMINED FROM TYPE 1 SELENOMETHIONYL CRYSTALS | | Descriptor: | SUBSTRATE BINDING DOMAIN OF DNAK, SUBSTRATE PEPTIDE (7 RESIDUES) | | Authors: | Zhu, X, Zhao, X, Burkholder, W.F, Gragerov, A, Ogata, C.M, Gottesman, M.E, Hendrickson, W.A. | | Deposit date: | 1996-06-03 | | Release date: | 1996-12-07 | | Last modified: | 2024-02-07 | | Method: | X-RAY DIFFRACTION (2 Å) | | Cite: | Structural analysis of substrate binding by the molecular chaperone DnaK.
Science, 272, 1996
|
|
1DKY
 
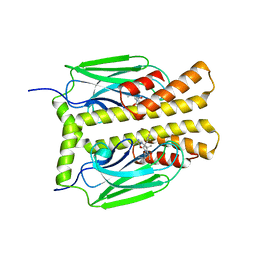 | | THE SUBSTRATE BINDING DOMAIN OF DNAK IN COMPLEX WITH A SUBSTRATE PEPTIDE, DETERMINED FROM TYPE 2 NATIVE CRYSTALS | | Descriptor: | DNAK, PEPTIDE SUBSTRATE | | Authors: | Zhu, X, Zhao, X, Burkholder, W.F, Gragerov, A, Ogata, C.M, Gottesman, M.E, Hendrickson, W.A. | | Deposit date: | 1996-06-03 | | Release date: | 1996-12-07 | | Last modified: | 2024-02-07 | | Method: | X-RAY DIFFRACTION (2.8 Å) | | Cite: | Structural analysis of substrate binding by the molecular chaperone DnaK.
Science, 272, 1996
|
|
2FYL
 
 | | Haddock model of the complex between double module of LRP, CR56, and first domain of receptor associated protein, RAP-d1. | | Descriptor: | Alpha-2-macroglobulin receptor-associated protein, CALCIUM ION, Low-density lipoprotein receptor-related protein 1 | | Authors: | Jensen, G.A, Andersen, O.M, Bonvin, A.M, Bjerrum-Bohr, I, Etzerodt, M, O'shea, C, Poulsen, F.M, Kragelund, B.B. | | Deposit date: | 2006-02-08 | | Release date: | 2006-10-10 | | Last modified: | 2022-03-09 | | Method: | SOLUTION NMR | | Cite: | Binding Site Structure of One LRP-RAP Complex:Implications for a Common Ligand-Receptor Binding Motif.
J.Mol.Biol., 362, 2006
|
|
3QK8
 
 | |
3QIV
 
 | |
3Q8N
 
 | |
3QHA
 
 | |
3QUA
 
 | |
3QUV
 
 | |
