4AKJ
 
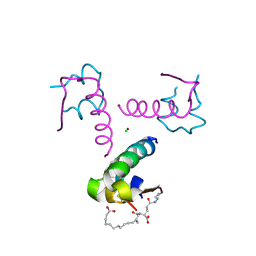 | | Ligand controlled assembly of hexamers, dihexamers, and linear multihexamer structures by an engineered acylated insulin | | Descriptor: | CHLORIDE ION, INSULIN A CHAIN, INSULIN B CHAIN, ... | | Authors: | Steensgaard, D.B, Schluckebier, G, Strauss, H.M, Norrman, M, Thomsen, J.K, Friderichsen, A.V, Havelund, S, Jonassen, I. | | Deposit date: | 2012-02-23 | | Release date: | 2013-01-09 | | Last modified: | 2024-10-23 | | Method: | X-RAY DIFFRACTION (2.01 Å) | | Cite: | Ligand Controlled Assembly of Hexamers, Dihexamers, and Linear Multihexamer Structures by the Engineered Acylated Insulin Degludec.
Biochemistry, 52, 2013
|
|
5JX2
 
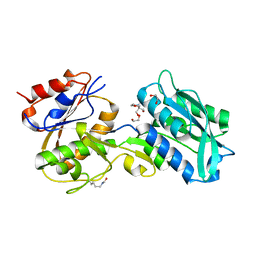 | | Crystal structure of MglB-2 (Tp0684) from Treponema pallidum | | Descriptor: | 1,2-ETHANEDIOL, 2-[BIS-(2-HYDROXY-ETHYL)-AMINO]-2-HYDROXYMETHYL-PROPANE-1,3-DIOL, CALCIUM ION, ... | | Authors: | Brautigam, C.A, Deka, R.K, Norgard, M.V. | | Deposit date: | 2016-05-12 | | Release date: | 2016-08-31 | | Last modified: | 2024-11-06 | | Method: | X-RAY DIFFRACTION (2.0504 Å) | | Cite: | The Tp0684 (MglB-2) Lipoprotein of Treponema pallidum: A Glucose-Binding Protein with Divergent Topology.
Plos One, 11, 2016
|
|
1R4P
 | | Shiga toxin type 2 | | Descriptor: | 1,2-ETHANEDIOL, 3-PYRIDINIUM-1-YLPROPANE-1-SULFONATE, FORMIC ACID, ... | | Authors: | Fraser, M.E, Fujinaga, M, Cherney, M.M, Melton-Celsa, A.R, Twiddy, E.M, O'Brien, A.D, James, M.N.G. | | Deposit date: | 2003-10-07 | | Release date: | 2004-05-11 | | Last modified: | 2024-11-13 | | Method: | X-RAY DIFFRACTION (1.77 Å) | | Cite: | Structure of Shiga Toxin Type 2 (Stx2) from Escherichia coli O157:H7.
J.Biol.Chem., 279, 2004
|
|
7U6V
 
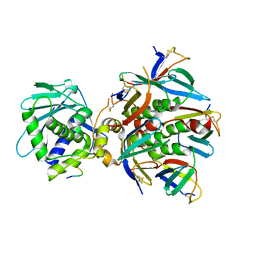 | | Cryo-EM structure of Shiga toxin 2 in complex with the native ribosomal P-stalk | | Descriptor: | C-terminal domain (CTD) from the Ribosomal P-stalk, Shiga toxin 2a subunit A (Stx2A), Shiga toxin 2a subunit B (Stx2B) | | Authors: | Kulczyk, A.W. | | Deposit date: | 2022-03-06 | | Release date: | 2023-01-11 | | Last modified: | 2024-11-13 | | Method: | ELECTRON MICROSCOPY (4.1 Å) | | Cite: | Cryo-EM structure of Shiga toxin 2 in complex with the native ribosomal P-stalk reveals residues involved in the binding interaction.
J.Biol.Chem., 299, 2022
|
|
2BLH
 
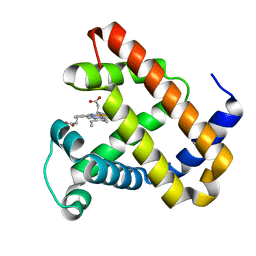 | | Ligand Migration and Protein Fluctuations in Myoglobin Mutant L29W | | Descriptor: | MYOGLOBIN, PROTOPORPHYRIN IX CONTAINING FE | | Authors: | Nienhaus, K, Ostermann, A, Nienhaus, G.U, Parak, F.G, Schmidt, M. | | Deposit date: | 2005-03-04 | | Release date: | 2005-04-06 | | Last modified: | 2024-05-08 | | Method: | X-RAY DIFFRACTION (1.77 Å) | | Cite: | Ligand Migration and Protein Fluctuations in Myoglobin Mutant L29W
Biochemistry, 44, 2005
|
|
1BV3
 
 | | HUMAN CARBONIC ANHYDRASE II COMPLEXED WITH UREA | | Descriptor: | 4-(HYDROXYMERCURY)BENZOIC ACID, PROTEIN (CARBONIC ANHYDRASE II), UREA, ... | | Authors: | Briganti, F, Mangani, S, Scozzafava, A, Vernaglione, G, Supuran, C.T. | | Deposit date: | 1998-09-22 | | Release date: | 1999-09-28 | | Last modified: | 2024-02-07 | | Method: | X-RAY DIFFRACTION (1.85 Å) | | Cite: | Carbonic anhydrase catalyzes cyanamide hydration to urea: is it mimicking the physiological reaction?
J.Biol.Inorg.Chem., 4, 1999
|
|
2RSE
 
 | | NMR structure of FKBP12-mTOR FRB domain-rapamycin complex structure determined based on PCS | | Descriptor: | Peptidyl-prolyl cis-trans isomerase FKBP1A, Serine/threonine-protein kinase mTOR, TERBIUM(III) ION | | Authors: | Kobashigawa, Y, Ushio, M, Saio, T, Inagaki, F. | | Deposit date: | 2012-01-25 | | Release date: | 2012-05-30 | | Last modified: | 2024-05-15 | | Method: | SOLUTION NMR | | Cite: | Convenient method for resolving degeneracies due to symmetry of the magnetic susceptibility tensor and its application to pseudo contact shift-based protein-protein complex structure determination.
J.Biomol.Nmr, 53, 2012
|
|
8GF3
 
 | | Crystallographic structure from BlMan5_7 | | Descriptor: | 1,2-ETHANEDIOL, 2-AMINO-2-HYDROXYMETHYL-PROPANE-1,3-DIOL, GH5 Mannanase, ... | | Authors: | Briganti, L, Araujo, E.A, Polikarpov, I. | | Deposit date: | 2023-03-07 | | Release date: | 2024-04-17 | | Last modified: | 2024-07-31 | | Method: | X-RAY DIFFRACTION (1.3 Å) | | Cite: | Unravelling biochemical and structural features of Bacillus licheniformis GH5 mannanase using site-directed mutagenesis and high-resolution protein crystallography studies.
Int.J.Biol.Macromol., 274, 2024
|
|
8GDV
 | |
8E4T
 
 | | Crystal structure of the kinase domain of RTKC8 from the choanoflagellate Monosiga brevicollis | | Descriptor: | PHOSPHATE ION, RTKC8 Kinase domain, STAUROSPORINE | | Authors: | Bajaj, T, Gee, C.L, Kuriyan, J. | | Deposit date: | 2022-08-18 | | Release date: | 2023-06-21 | | Last modified: | 2023-10-25 | | Method: | X-RAY DIFFRACTION (1.95 Å) | | Cite: | Crystal structure of the kinase domain of a receptor tyrosine kinase from a choanoflagellate, Monosiga brevicollis.
Plos One, 18, 2023
|
|
6QVS
 
 | | Unliganded structure of the human wild type Beta-galactoside alpha-2,6-sialyltransferase 1 (ST6Gal1) | | Descriptor: | Beta-galactoside alpha-2,6-sialyltransferase 1, DI(HYDROXYETHYL)ETHER, GLYCEROL | | Authors: | Harrus, D, Glumoff, T. | | Deposit date: | 2019-03-04 | | Release date: | 2020-06-10 | | Last modified: | 2024-11-13 | | Method: | X-RAY DIFFRACTION (1.6 Å) | | Cite: | Unliganded and CMP-Neu5Ac bound structures of human alpha-2,6-sialyltransferase ST6Gal I at high resolution.
J.Struct.Biol., 212, 2020
|
|
6J2R
 
 | | Crystal structure of Striga hermonthica HTL8 (ShHTL8) | | Descriptor: | GLYCEROL, Hyposensitive to light 8 | | Authors: | Zhang, Y.Y, Xi, Z. | | Deposit date: | 2019-01-02 | | Release date: | 2020-01-15 | | Last modified: | 2023-11-22 | | Method: | X-RAY DIFFRACTION (1.4 Å) | | Cite: | Crystal structure and biochemical characterization of Striga hermonthica HYPO-SENSITIVE TO LIGHT 8 (ShHTL8) in strigolactone signaling pathway.
Biochem.Biophys.Res.Commun., 523, 2020
|
|
1YEJ
 
 | | CATALYTIC ANTIBODY COMPLEX | | Descriptor: | 6-{4-[HYDROXY-(4-NITRO-PHENOXY)-PHOSPHORYL]-BUTYRYLAMINO}-HEXANOIC ACID, PROTEIN (IG ANTIBODY D2.3 (HEAVY CHAIN)), PROTEIN (IG ANTIBODY D2.3 (LIGHT CHAIN)), ... | | Authors: | Gigant, B, Knossow, M. | | Deposit date: | 1998-08-13 | | Release date: | 1999-01-13 | | Last modified: | 2024-10-30 | | Method: | X-RAY DIFFRACTION (1.85 Å) | | Cite: | Crossreactivity, efficiency and catalytic specificity of an esterase-like antibody.
J.Mol.Biol., 284, 1998
|
|
1YEI
 
 | | CATALYTIC ANTIBODY D2.3 COMPLEX | | Descriptor: | PARA-NITROPHENYLPHOSPHONOBUTANOYL-GLYCINE, PROTEIN (IG ANTIBODY D2.3 (HEAVY CHAIN)), PROTEIN (IG ANTIBODY D2.3 (LIGHT CHAIN)), ... | | Authors: | Gigant, B, Knossow, M. | | Deposit date: | 1998-08-13 | | Release date: | 1999-01-13 | | Last modified: | 2024-10-09 | | Method: | X-RAY DIFFRACTION (1.9 Å) | | Cite: | Crossreactivity, efficiency and catalytic specificity of an esterase-like antibody.
J.Mol.Biol., 284, 1998
|
|
1YEK
 
 | | CATALYTIC ANTIBODY D2.3 COMPLEX | | Descriptor: | P-NITROPHENOL, PROTEIN (IG ANTIBODY D2.3 (HEAVY CHAIN)), PROTEIN (IG ANTIBODY D2.3 (LIGHT CHAIN)), ... | | Authors: | Gigant, B, Knossow, M. | | Deposit date: | 1998-08-13 | | Release date: | 1999-01-13 | | Last modified: | 2024-10-09 | | Method: | X-RAY DIFFRACTION (2.1 Å) | | Cite: | Crossreactivity, efficiency and catalytic specificity of an esterase-like antibody.
J.Mol.Biol., 284, 1998
|
|
2WUU
 
 | | Structure of N-myristoyltransferase from L. donovani | | Descriptor: | 2-oxopentadecyl-CoA, N-MYRISTOYLTRANSFERASE | | Authors: | Brannigan, J.A, Smith, B.A, Yu, Z, Hodgkinson, M.R, Leatherbarrow, R.J, Tate, E.W, Brzozowski, A.M, Smith, D.F, Wilkinson, A.J. | | Deposit date: | 2009-10-09 | | Release date: | 2009-12-01 | | Last modified: | 2023-12-20 | | Method: | X-RAY DIFFRACTION (1.42 Å) | | Cite: | N-Myristoyltransferase from Leishmania Donovani: Structural and Functional Characterisation of a Potential Drug Target for Visceral Leishmaniasis.
J.Mol.Biol., 396, 2010
|
|
5YCL
 
 | |
6U3U
 
 | | Crystal Structure of Shiga Toxin 2K | | Descriptor: | 1,2-ETHANEDIOL, 4-(2-HYDROXYETHYL)-1-PIPERAZINE ETHANESULFONIC ACID, Shiga toxin 2K subunit A, ... | | Authors: | Zhang, Y.Z, He, X.H. | | Deposit date: | 2019-08-22 | | Release date: | 2020-07-01 | | Last modified: | 2024-10-16 | | Method: | X-RAY DIFFRACTION (2.287 Å) | | Cite: | Structural and Functional Characterization of Stx2k, a New Subtype of Shiga Toxin 2.
Microorganisms, 8, 2019
|
|
4P2C
 
 | | Complex of Shiga toxin 2e with a neutralizing single-domain antibody | | Descriptor: | Nanobody 1, Anti-F4+ETEC bacteria VHH variable region, Shiga toxin 2e, ... | | Authors: | Lo, A.W.H, Moonens, K, De Kerpel, M, Brys, L, Pardon, E, Remaut, H, De Greve, H. | | Deposit date: | 2014-03-03 | | Release date: | 2014-07-30 | | Last modified: | 2024-10-30 | | Method: | X-RAY DIFFRACTION (2.818 Å) | | Cite: | The Molecular Mechanism of Shiga Toxin Stx2e Neutralization by a Single-domain Antibody Targeting the Cell Receptor-binding Domain.
J.Biol.Chem., 289, 2014
|
|
5ZW2
 
 | | FAD complex of PigA | | Descriptor: | 1,2-ETHANEDIOL, 1,4,7,10,13,16-HEXAOXACYCLOOCTADECANE, ACETATE ION, ... | | Authors: | Lee, C.-C, Ko, T.-P, Wang, A.H.J. | | Deposit date: | 2018-05-14 | | Release date: | 2018-09-05 | | Last modified: | 2023-11-22 | | Method: | X-RAY DIFFRACTION (1.803 Å) | | Cite: | Crystal Structure of PigA: A Prolyl Thioester-Oxidizing Enzyme in Prodigiosin Biosynthesis.
Chembiochem, 20, 2019
|
|
2VQZ
 
 | | Structure of the cap-binding domain of influenza virus polymerase subunit PB2 with bound m7GTP | | Descriptor: | 7N-METHYL-8-HYDROGUANOSINE-5'-TRIPHOSPHATE, POLYMERASE BASIC PROTEIN 2 | | Authors: | Guilligay, D, Tarendeau, F, Resa-Infante, P, Coloma, R, Crepin, T, Sehr, P, Lewis, J, Ruigrok, R.W.H, Ortin, J, Hart, D.J, Cusack, S. | | Deposit date: | 2008-03-21 | | Release date: | 2008-05-13 | | Last modified: | 2024-10-23 | | Method: | X-RAY DIFFRACTION (2.3 Å) | | Cite: | The Structural Basis for CAP Binding by Influenza Virus Polymerase Subunit Pb2.
Nat.Struct.Mol.Biol., 15, 2008
|
|
5ZW0
 
 | | Apo-form PigA | | Descriptor: | L-prolyl-[peptidyl-carrier protein] dehydrogenase | | Authors: | Lee, C.-C, Ko, T.-P, Wang, A.H.J. | | Deposit date: | 2018-05-14 | | Release date: | 2018-09-05 | | Last modified: | 2023-11-22 | | Method: | X-RAY DIFFRACTION (2.54 Å) | | Cite: | Crystal Structure of PigA: A Prolyl Thioester-Oxidizing Enzyme in Prodigiosin Biosynthesis.
Chembiochem, 20, 2019
|
|
4AJZ
 
 | | Ligand controlled assembly of hexamers, dihexamers, and linear multihexamer structures by an engineered acylated insulin | | Descriptor: | CHLORIDE ION, INSULIN A CHAIN, INSULIN B CHAIN, ... | | Authors: | Steensgaard, D.B, Schluckebier, G, Strauss, H.M, Norrman, M, Thomsen, J.K, Friderichsen, A.V, Havelund, S, Jonassen, I. | | Deposit date: | 2012-02-21 | | Release date: | 2013-01-09 | | Last modified: | 2024-11-20 | | Method: | X-RAY DIFFRACTION (1.8 Å) | | Cite: | Ligand Controlled Assembly of Hexamers, Dihexamers, and Linear Multihexamer Structures by the Engineered Acylated Insulin Degludec.
Biochemistry, 52, 2013
|
|
4AJX
 | | Ligand controlled assembly of hexamers, dihexamers, and linear multihexamer structures by an engineered acylated insulin | | Descriptor: | IMIDAZOLE, INSULIN, N-(16-Carboxyhexadecanoyl)-L-glutamic acid, ... | | Authors: | Steensgaard, D.B, Schluckebier, G, Strauss, H.M, Norrman, M, Thomsen, J.K, Friderichsen, A.V, Havelund, S, Jonassen, I. | | Deposit date: | 2012-02-20 | | Release date: | 2013-01-09 | | Last modified: | 2024-11-06 | | Method: | X-RAY DIFFRACTION (1.2 Å) | | Cite: | Ligand Controlled Assembly of Hexamers, Dihexamers, and Linear Multihexamer Structures by the Engineered Acylated Insulin Degludec.
Biochemistry, 52, 2013
|
|
2BOS
 
 | | A MUTANT SHIGA-LIKE TOXIN IIE BOUND TO ITS RECEPTOR | | Descriptor: | N-BUTANE, PROTEIN (SHIGA-LIKE TOXIN IIE B SUBUNIT), alpha-D-galactopyranose-(1-4)-beta-D-galactopyranose, ... | | Authors: | Ling, H, Boodhoo, A, Armstrong, G.D, Clark, C.G, Brunton, J.L, Read, R.J. | | Deposit date: | 1998-10-20 | | Release date: | 1999-10-20 | | Last modified: | 2024-10-16 | | Method: | X-RAY DIFFRACTION (2 Å) | | Cite: | A mutant Shiga-like toxin IIe bound to its receptor Gb(3): structure of a group II Shiga-like toxin with altered binding specificity.
Structure Fold.Des., 8, 2000
|
|
