3SHI
 
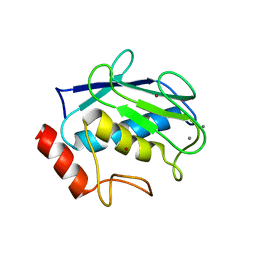 | | Crystal structure of human MMP1 catalytic domain at 2.2 A resolution | | Descriptor: | CALCIUM ION, Interstitial collagenase, ZINC ION | | Authors: | Bertini, I, Calderone, V, Cerofolini, L, Fragai, M, Geraldes, C.F.G.C, Hermann, P, Luchinat, C, Parigi, G, Teixeira, J. | | Deposit date: | 2011-06-16 | | Release date: | 2011-09-21 | | Last modified: | 2023-09-13 | | Method: | X-RAY DIFFRACTION (2.2 Å) | | Cite: | The catalytic domain of MMP-1 studied through tagged lanthanides.
Febs Lett., 586, 2012
|
|
3SIX
 
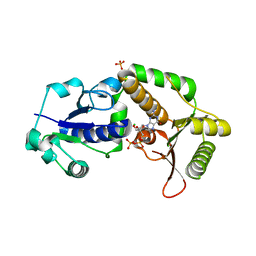 | | Crystal structure of NodZ alpha-1,6-fucosyltransferase soaked with GDP-fucose | | Descriptor: | CHLORIDE ION, GUANOSINE-5'-DIPHOSPHATE, Nodulation fucosyltransferase NodZ, ... | | Authors: | Brzezinski, K, Dauter, Z, Jaskolski, M. | | Deposit date: | 2011-06-20 | | Release date: | 2012-02-08 | | Last modified: | 2023-09-13 | | Method: | X-RAY DIFFRACTION (2.35 Å) | | Cite: | Structures of NodZ alpha-1,6-fucosyltransferase in complex with GDP and GDP-fucose
Acta Crystallogr.,Sect.D, 68, 2012
|
|
3SMH
 
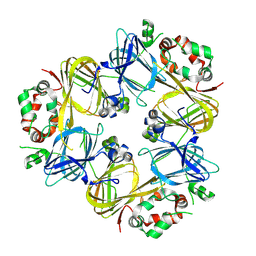 | |
2A33
 
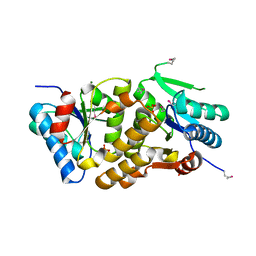 | | X-Ray Structure of a Lysine Decarboxylase-Like Protein from Arabidopsis Thaliana Gene AT2G37210 | | Descriptor: | MAGNESIUM ION, SULFATE ION, hypothetical protein | | Authors: | Wesenberg, G.E, Phillips Jr, G.N, Mccoy, J.G, Bitto, E, Bingman, C.A, Allard, S.T.M, Center for Eukaryotic Structural Genomics (CESG) | | Deposit date: | 2005-06-23 | | Release date: | 2005-07-19 | | Last modified: | 2023-11-15 | | Method: | X-RAY DIFFRACTION (1.95 Å) | | Cite: | X-ray crystal structures of the conserved hypothetical proteins from Arabidopsis thaliana gene loci At5g11950 and AT2g37210.
Proteins, 65, 2006
|
|
1F8X
 
 | |
2A9W
 
 | | E. coli TS complexed with dUMP and inhibitor GA9 | | Descriptor: | 2'-DEOXYURIDINE 5'-MONOPHOSPHATE, 2-BROMOPHENOL, 3,3-BIS(3-BROMO-4-HYDROXYPHENYL)-7-CHLORO-1H,3H-BENZO[DE]ISOCHROMEN-1-ONE, ... | | Authors: | Finer-Moore, J.S, Anderson, A.C, O'Neil, R.H, Costi, M.P, Ferrari, S, Krucinski, J, Stroud, R.M. | | Deposit date: | 2005-07-12 | | Release date: | 2005-10-11 | | Last modified: | 2023-08-23 | | Method: | X-RAY DIFFRACTION (1.65 Å) | | Cite: | The structure of Cryptococcus neoformans thymidylate synthase suggests strategies for using target dynamics for species-specific inhibition.
Acta Crystallogr.,Sect.D, 61, 2005
|
|
2AEI
 
 | | Crystal structure of a ternary complex of factor VIIa/tissue factor and 2-[[6-[3-(aminoiminomethyl)phenoxy]-3,5-difluro-4-[(1-methyl-3-phenylpropyl)amino]-2-pyridinyl]oxy]-benzoic acid | | Descriptor: | 2-({6-{3-[AMINO(IMINO)METHYL]PHENOXY}-3,5-DIFLUORO-4-[(1-METHYL-3-PHENYLPROPYL)AMINO]-2-PYRIDINYL}OXY)BENZOIC ACID, CACODYLATE ION, CALCIUM ION, ... | | Authors: | Adler, M, Whitlow, M. | | Deposit date: | 2005-07-22 | | Release date: | 2006-08-01 | | Last modified: | 2023-11-15 | | Method: | X-RAY DIFFRACTION (2.52 Å) | | Cite: | The discovery of fluoropyridine-based inhibitors of the Factor VIIa/TF complex.
Bioorg.Med.Chem.Lett., 15, 2005
|
|
2NWD
 
 | |
1FBL
 
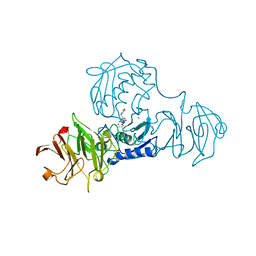 | | STRUCTURE OF FULL-LENGTH PORCINE SYNOVIAL COLLAGENASE (MMP1) REVEALS A C-TERMINAL DOMAIN CONTAINING A CALCIUM-LINKED, FOUR-BLADED BETA-PROPELLER | | Descriptor: | CALCIUM ION, FIBROBLAST (INTERSTITIAL) COLLAGENASE (MMP-1), N-[3-(N'-HYDROXYCARBOXAMIDO)-2-(2-METHYLPROPYL)-PROPANOYL]-O-TYROSINE-N-METHYLAMIDE, ... | | Authors: | Li, J, Brick, P, Blow, D.M. | | Deposit date: | 1995-04-24 | | Release date: | 1996-01-29 | | Last modified: | 2012-02-22 | | Method: | X-RAY DIFFRACTION (2.5 Å) | | Cite: | Structure of full-length porcine synovial collagenase reveals a C-terminal domain containing a calcium-linked, four-bladed beta-propeller.
Structure, 3, 1995
|
|
2O0R
 
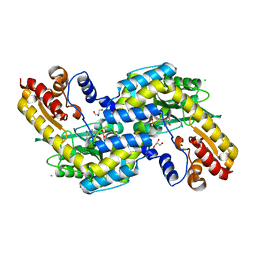 | | The three-dimensional structure of N-Succinyldiaminopimelate aminotransferase from Mycobacterium tuberculosis | | Descriptor: | CHLORIDE ION, GLYCEROL, Rv0858c (N-Succinyldiaminopimelate aminotransferase), ... | | Authors: | Weyand, S, Kefala, G, Weiss, M.S, TB Structural Genomics Consortium (TBSGC) | | Deposit date: | 2006-11-28 | | Release date: | 2007-02-27 | | Last modified: | 2023-11-15 | | Method: | X-RAY DIFFRACTION (2 Å) | | Cite: | The Three-dimensional Structure of N-Succinyldiaminopimelate Aminotransferase from Mycobacterium tuberculosis
J.Mol.Biol., 367, 2007
|
|
3SIW
 
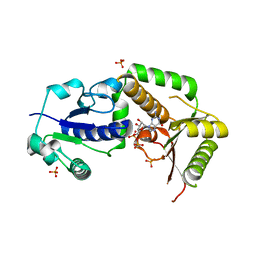 | | Crystal structure of NodZ alpha-1,6-fucosyltransferase co-crystallized with GDP | | Descriptor: | GUANOSINE-5'-DIPHOSPHATE, Nodulation fucosyltransferase NodZ, PHOSPHATE ION | | Authors: | Brzezinski, K, Dauter, Z, Jaskolski, M. | | Deposit date: | 2011-06-20 | | Release date: | 2012-02-08 | | Last modified: | 2023-09-13 | | Method: | X-RAY DIFFRACTION (1.98 Å) | | Cite: | Structures of NodZ alpha-1,6-fucosyltransferase in complex with GDP and GDP-fucose
Acta Crystallogr.,Sect.D, 68, 2012
|
|
1FGO
 
 | |
2O6Y
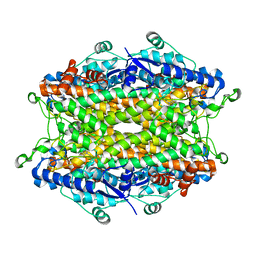 
 | | Tyrosine ammonia-lyase from Rhodobacter sphaeroides | | Descriptor: | Putative histidine ammonia-lyase | | Authors: | Louie, G.V, Bowman, M.E, Moffitt, M.C, Baiga, T.J, Moore, B.S, Noel, J.P. | | Deposit date: | 2006-12-09 | | Release date: | 2007-01-16 | | Last modified: | 2023-11-15 | | Method: | X-RAY DIFFRACTION (1.5 Å) | | Cite: | Structural determinants and modulation of substrate specificity in phenylalanine-tyrosine ammonia-lyases.
Chem.Biol., 13, 2006
|
|
3SNP
 
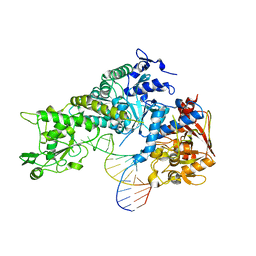 | |
2OCX
 
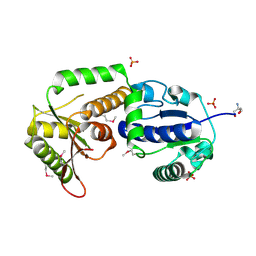 | | Crystal structure of Se-Met fucosyltransferase NodZ from Bradyrhizobium | | Descriptor: | 2-AMINO-2-HYDROXYMETHYL-PROPANE-1,3-DIOL, Nodulation fucosyltransferase NodZ, PHOSPHATE ION | | Authors: | Brzezinski, K, Stepkowski, T, Panjikar, S, Bujacz, G, Jaskolski, M. | | Deposit date: | 2006-12-21 | | Release date: | 2007-11-06 | | Last modified: | 2023-12-27 | | Method: | X-RAY DIFFRACTION (2.2 Å) | | Cite: | High-resolution structure of NodZ fucosyltransferase involved in the biosynthesis of the nodulation factor.
Acta Biochim.Pol., 54, 2007
|
|
3TRJ
 
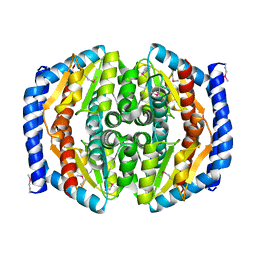 | | Structure of a phosphoheptose isomerase from Francisella tularensis | | Descriptor: | Phosphoheptose isomerase | | Authors: | Cheung, J, Franklin, M.C, Rudolph, M, Cassidy, M, Gary, E, Burshteyn, F, Love, J. | | Deposit date: | 2011-09-09 | | Release date: | 2011-09-21 | | Last modified: | 2023-12-06 | | Method: | X-RAY DIFFRACTION (2.8 Å) | | Cite: | Rapid countermeasure discovery against Francisella tularensis based on a metabolic network reconstruction.
Plos One, 8, 2013
|
|
1Y2X
 
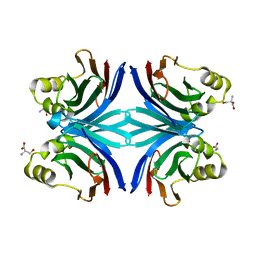 | | Crystal structure of the tetragonal form of the common edible mushroom (Agaricus bisporus) lectin in complex with T-antigen and N-acetylglucosamine | | Descriptor: | 2-acetamido-2-deoxy-beta-D-glucopyranose, SERINE, beta-D-galactopyranose-(1-3)-2-acetamido-2-deoxy-beta-D-galactopyranose, ... | | Authors: | Carrizo, M.E, Capaldi, S, Perduca, M, Irazoqui, F.J, Nores, G.A, Monaco, H.L. | | Deposit date: | 2004-11-23 | | Release date: | 2004-12-21 | | Last modified: | 2024-04-03 | | Method: | X-RAY DIFFRACTION (2.36 Å) | | Cite: | The Antineoplastic Lectin of the Common Edible Mushroom (Agaricus bisporus) Has Two Binding Sites, Each Specific for a Different Configuration at a Single Epimeric Hydroxyl
J.Biol.Chem., 280, 2005
|
|
1CV2
 
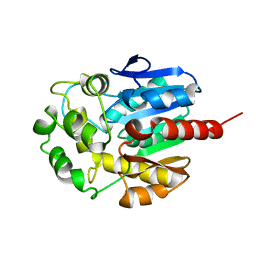 | | Hydrolytic haloalkane dehalogenase linb from sphingomonas paucimobilis UT26 AT 1.6 A resolution | | Descriptor: | HALOALKANE DEHALOGENASE | | Authors: | Marek, J, Vevodova, J, Damborsky, J, Smatanova, I, Svensson, L.A, Newman, J, Nagata, Y, Takagi, M. | | Deposit date: | 1999-08-22 | | Release date: | 2000-09-11 | | Last modified: | 2023-08-09 | | Method: | X-RAY DIFFRACTION (1.58 Å) | | Cite: | Crystal structure of the haloalkane dehalogenase from Sphingomonas paucimobilis UT26.
Biochemistry, 39, 2000
|
|
1CI9
 
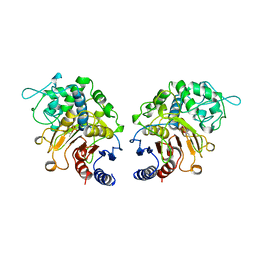 | | DFP-INHIBITED ESTERASE ESTB FROM BURKHOLDERIA GLADIOLI | | Descriptor: | DIISOPROPYL PHOSPHONATE, PROTEIN (CARBOXYLESTERASE) | | Authors: | Wagner, U.G, Petersen, E.I, Schwab, H, Kratky, C. | | Deposit date: | 1999-04-08 | | Release date: | 2001-12-12 | | Last modified: | 2024-04-03 | | Method: | X-RAY DIFFRACTION (1.8 Å) | | Cite: | EstB from Burkholderia gladioli: a novel esterase with a beta-lactamase fold reveals steric factors to discriminate between esterolytic and beta-lactam cleaving activity
Protein Sci., 11, 2002
|
|
1Y2T
 
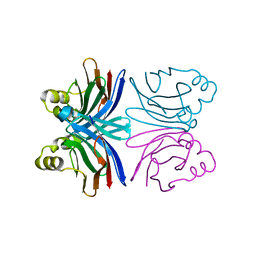 | | Crystal structure of the common edible mushroom (Agaricus bisporus) lectin | | Descriptor: | lectin | | Authors: | Carrizo, M.E, Capaldi, S, Perduca, M, Irazoqui, F.J, Nores, G.A, Monaco, H.L. | | Deposit date: | 2004-11-23 | | Release date: | 2004-12-21 | | Last modified: | 2024-03-13 | | Method: | X-RAY DIFFRACTION (1.5 Å) | | Cite: | The Antineoplastic Lectin of the Common Edible Mushroom (Agaricus bisporus) Has Two Binding Sites, Each Specific for a Different Configuration at a Single Epimeric Hydroxyl
J.Biol.Chem., 280, 2005
|
|
2PZM
 
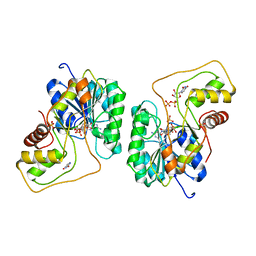 | | Crystal structure of the Bordetella bronchiseptica enzyme WbmG in complex with NAD and UDP | | Descriptor: | NICOTINAMIDE-ADENINE-DINUCLEOTIDE, Putative nucleotide sugar epimerase/ dehydratase, SULFATE ION, ... | | Authors: | Harmer, N.J, King, J.D, Palmer, C.M, Maskell, D, Blundell, T.L. | | Deposit date: | 2007-05-18 | | Release date: | 2007-10-02 | | Last modified: | 2023-08-30 | | Method: | X-RAY DIFFRACTION (2 Å) | | Cite: | Predicting protein function from structure--the roles of short-chain dehydrogenase/reductase enzymes in Bordetella O-antigen biosynthesis.
J.Mol.Biol., 374, 2007
|
|
1I19
 
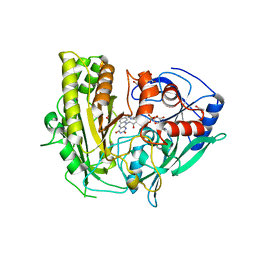 | | CRYSTAL STRUCTURE OF CHOLESTEROL OXIDASE FROM B.STEROLICUM | | Descriptor: | 1,2-ETHANEDIOL, CACODYLATE ION, CHOLESTEROL OXIDASE, ... | | Authors: | Coulombe, R, Yue, K.Q, Ghisla, S, Vrielink, A. | | Deposit date: | 2001-01-31 | | Release date: | 2001-08-08 | | Last modified: | 2011-07-13 | | Method: | X-RAY DIFFRACTION (1.7 Å) | | Cite: | Oxygen access to the active site of cholesterol oxidase through a narrow channel is gated by an Arg-Glu pair.
J.Biol.Chem., 276, 2001
|
|
1CPO
 
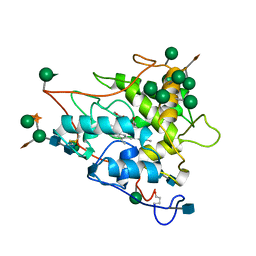 | | CHLOROPEROXIDASE | | Descriptor: | 2-acetamido-2-deoxy-beta-D-glucopyranose, 2-acetamido-2-deoxy-beta-D-glucopyranose-(1-4)-2-acetamido-2-deoxy-beta-D-glucopyranose, CHLOROPEROXIDASE, ... | | Authors: | Sundaramoorthy, M, Poulos, T.L. | | Deposit date: | 1996-02-10 | | Release date: | 1997-02-12 | | Last modified: | 2020-07-29 | | Method: | X-RAY DIFFRACTION (1.9 Å) | | Cite: | The crystal structure of chloroperoxidase: a heme peroxidase--cytochrome P450 functional hybrid.
Structure, 3, 1995
|
|
2PZK
 
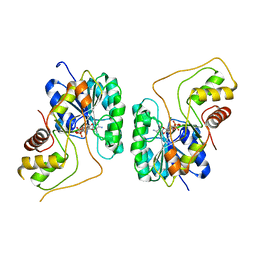 | | Crystal structure of the Bordetella bronchiseptica enzyme WbmG in complex with NAD | | Descriptor: | MAGNESIUM ION, NICOTINAMIDE-ADENINE-DINUCLEOTIDE, Putative nucleotide sugar epimerase/ dehydratase | | Authors: | King, J.D, Harmer, N.J, Maskell, D.J, Blundell, T.L. | | Deposit date: | 2007-05-18 | | Release date: | 2007-10-02 | | Last modified: | 2024-02-21 | | Method: | X-RAY DIFFRACTION (2.1 Å) | | Cite: | Predicting protein function from structure--the roles of short-chain dehydrogenase/reductase enzymes in Bordetella O-antigen biosynthesis.
J.Mol.Biol., 374, 2007
|
|
2Q1U
 
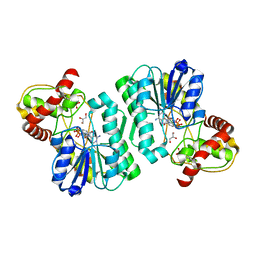 | | Crystal structure of the Bordetella bronchiseptica enzyme WbmF in complex with NAD+ and UDP | | Descriptor: | GLYCEROL, NICOTINAMIDE-ADENINE-DINUCLEOTIDE, Putative nucleotide sugar epimerase/ dehydratase, ... | | Authors: | Harmer, N.J, King, J.D, Palmer, C.M, Maskell, D, Blundell, T.L. | | Deposit date: | 2007-05-25 | | Release date: | 2007-10-02 | | Last modified: | 2023-08-30 | | Method: | X-RAY DIFFRACTION (1.7 Å) | | Cite: | Predicting protein function from structure--the roles of short-chain dehydrogenase/reductase enzymes in Bordetella O-antigen biosynthesis.
J.Mol.Biol., 374, 2007
|
|
