1E54
 
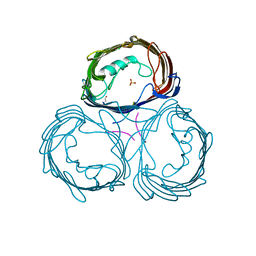 | | Anion-selective porin from Comamonas acidovorans | | Descriptor: | CALCIUM ION, OMP32, OUTER MEMBRANE PORIN PROTEIN 32, ... | | Authors: | Zeth, K, Diederichs, K, Welte, W, Engelhardt, H. | | Deposit date: | 2000-07-17 | | Release date: | 2001-07-12 | | Last modified: | 2020-03-11 | | Method: | X-RAY DIFFRACTION (2.1 Å) | | Cite: | Crystal Structure of Omp32, the Anion-Selective Porin from Comamonas Acidovorans, in Complex with a Periplasmic Peptideat 2.1 A Resolution
Structure, 8, 2000
|
|
7PDC
 
 | |
6SHK
 
 | |
6FHG
 
 | |
1LZW
 
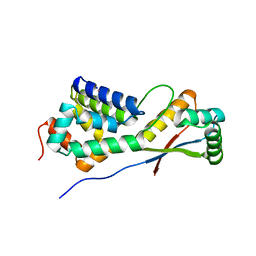 | | Structural basis of ClpS-mediated switch in ClpA substrate recognition | | Descriptor: | ATP-dependent clp protease ATP-binding subunit ClpA, PLATINUM (II) ION, Protein yljA | | Authors: | Zeth, K, Ravelli, R.B, Paal, K, Cusack, S, Bukau, B, Dougan, D.A. | | Deposit date: | 2002-06-11 | | Release date: | 2002-11-27 | | Last modified: | 2024-02-14 | | Method: | X-RAY DIFFRACTION (2.5 Å) | | Cite: | Structural analysis of the adaptor protein ClpS in complex with the N-terminal domain of ClpA
Nat.Struct.Biol., 9, 2002
|
|
1MG9
 
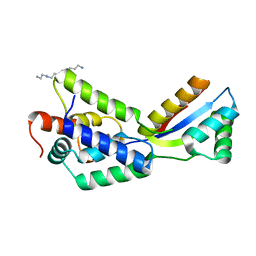 | | The structural basis of ClpS-mediated switch in ClpA substrate recognition | | Descriptor: | ATP dependent clp protease ATP-binding subunit clpA, SPERMINE (FULLY PROTONATED FORM), protein yljA | | Authors: | Zeth, K, Ravelli, R.B, Paal, K, Cusack, S, Bukau, B, Dougan, D.A. | | Deposit date: | 2002-08-15 | | Release date: | 2002-11-27 | | Last modified: | 2024-02-14 | | Method: | X-RAY DIFFRACTION (2.3 Å) | | Cite: | Structural analysis of the adaptor protein ClpS in complex with the N-terminal domain of ClpA
Nat.Struct.Biol., 9, 2002
|
|
6ZLX
 
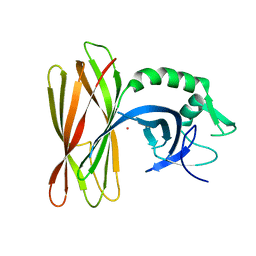 | |
5NNT
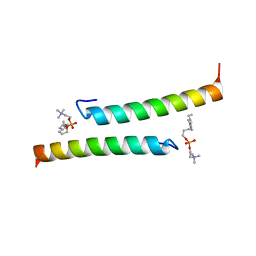 
 | | The dimeric structure of LL-37 crystallized in DPC | | Descriptor: | Cathelicidin antimicrobial peptide, dodecyl 2-(trimethylammonio)ethyl phosphate | | Authors: | Zeth, K, Sancho-Vaello, E. | | Deposit date: | 2017-04-10 | | Release date: | 2018-01-24 | | Last modified: | 2019-10-16 | | Method: | X-RAY DIFFRACTION (2.209 Å) | | Cite: | Structural remodeling and oligomerization of human cathelicidin on membranes suggest fibril-like structures as active species.
Sci Rep, 7, 2017
|
|
5NNK
 
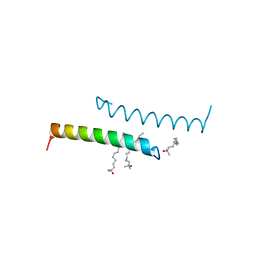 | | The structure of LL-37 crystallized in the presence LDAO | | Descriptor: | Cathelicidin antimicrobial peptide, LAURYL DIMETHYLAMINE-N-OXIDE | | Authors: | Zeth, K, Sancho-Vaello, E. | | Deposit date: | 2017-04-10 | | Release date: | 2018-01-24 | | Last modified: | 2019-10-16 | | Method: | X-RAY DIFFRACTION (1.798 Å) | | Cite: | Structural remodeling and oligomerization of human cathelicidin on membranes suggest fibril-like structures as active species.
Sci Rep, 7, 2017
|
|
3H7Z
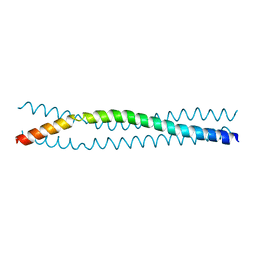 
 | |
3H7X
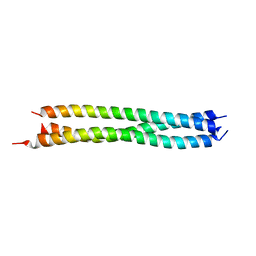 
 | |
6SEV
 
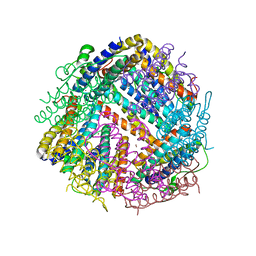 | |
4N59
 
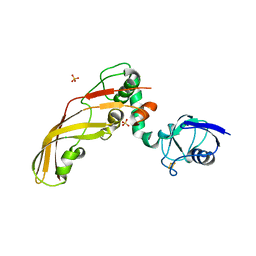 | | The Crystal Structure of Pectocin M2 at 2.3 Angstroms | | Descriptor: | CHLORIDE ION, FE2/S2 (INORGANIC) CLUSTER, Pectocin M2, ... | | Authors: | Zeth, K, Grinter, R, Roszak, A.W, Cogdell, R.J, Walker, D. | | Deposit date: | 2013-10-09 | | Release date: | 2014-06-04 | | Last modified: | 2023-09-20 | | Method: | X-RAY DIFFRACTION (2.3 Å) | | Cite: | Structure of the atypical bacteriocin pectocin M2 implies a novel mechanism of protein uptake.
Mol.Microbiol., 93, 2014
|
|
6Z4E
 
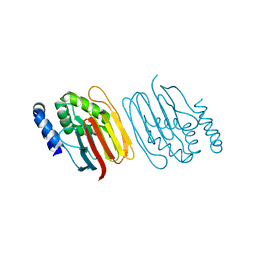 | | The structure of the C-terminal domain of RssB from E. coli | | Descriptor: | Regulator of RpoS | | Authors: | Zeth, K, Dimce, M, Terrence, D.M, Schuenemann, V, Dougan, D. | | Deposit date: | 2020-05-25 | | Release date: | 2020-07-29 | | Last modified: | 2020-09-30 | | Method: | X-RAY DIFFRACTION (2 Å) | | Cite: | Insight into the RssB-Mediated Recognition and Delivery of sigma s to the AAA+ Protease, ClpXP.
Biomolecules, 10, 2020
|
|
6Z4C
 
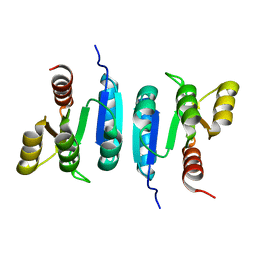 | | The structure of the N-terminal domain of RssB from E. coli | | Descriptor: | Regulator of RpoS | | Authors: | Zeth, K, Dimce, M, Terrence, D.M, Schuenemann, V, Dougan, D. | | Deposit date: | 2020-05-25 | | Release date: | 2020-07-29 | | Last modified: | 2024-01-24 | | Method: | X-RAY DIFFRACTION (2 Å) | | Cite: | Insight into the RssB-Mediated Recognition and Delivery of sigma s to the AAA+ Protease, ClpXP.
Biomolecules, 10, 2020
|
|
5NMN
 
 | |
5NNM
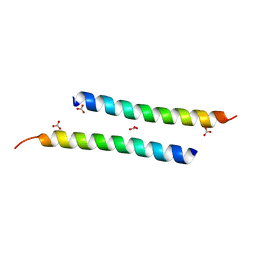 
 | | The crystal structure of dimeric LL-37 | | Descriptor: | CARBONATE ION, Cathelicidin antimicrobial peptide | | Authors: | Zeth, K, Sancho-Vaello, E. | | Deposit date: | 2017-04-10 | | Release date: | 2018-01-24 | | Last modified: | 2019-10-16 | | Method: | X-RAY DIFFRACTION (1.9 Å) | | Cite: | Structural remodeling and oligomerization of human cathelicidin on membranes suggest fibril-like structures as active species.
Sci Rep, 7, 2017
|
|
3D9X
 
 | |
1MOJ
 
 | | Crystal structure of an archaeal dps-homologue from Halobacterium salinarum | | Descriptor: | Dps-like ferritin, FE (III) ION, MAGNESIUM ION, ... | | Authors: | Zeth, K, Offermann, S, Essen, L.O, Oesterhelt, D. | | Deposit date: | 2002-09-09 | | Release date: | 2004-04-20 | | Last modified: | 2024-02-14 | | Method: | X-RAY DIFFRACTION (1.9 Å) | | Cite: | Iron-oxo clusters biomineralizing on protein surfaces: Structural analysis of Halobacterium salinarum DpsA in its low- and high-iron states.
Proc.Natl.Acad.Sci.USA, 101, 2004
|
|
3LT6
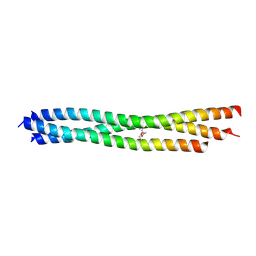 
 | |
3LT7
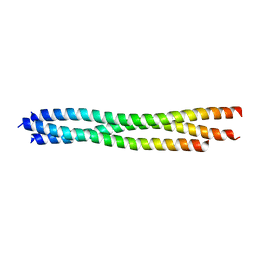 
 | |
6HX2
 
 | |
6HUI
 
 | |
6HV1
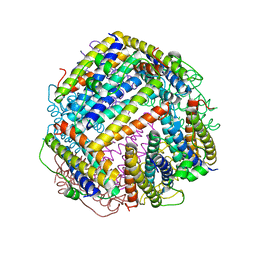 
 | |
6HVQ
 
 | |
