2RIM
 
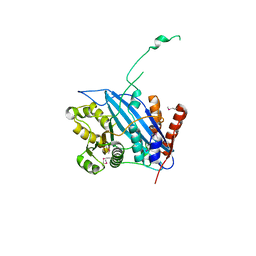 | | Crystal structure of Rtt109 | | Descriptor: | Regulator of Ty1 transposition protein 109 | | Authors: | Yuan, Y.A. | | Deposit date: | 2007-10-12 | | Release date: | 2008-09-02 | | Last modified: | 2011-07-13 | | Method: | X-RAY DIFFRACTION (2.2 Å) | | Cite: | Structural insights into histone h3 lysine 56 acetylation by rtt109
Structure, 16, 2008
|
|
3VZ1
 
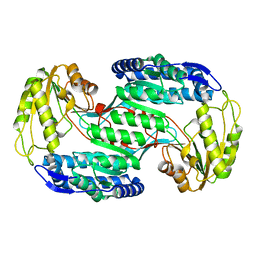 | |
3VZ3
 
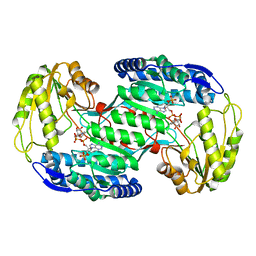 | | Structural insights into substrate and cofactor selection by sp2771 | | Descriptor: | 4-oxobutanoic acid, NADP NICOTINAMIDE-ADENINE-DINUCLEOTIDE PHOSPHATE, Succinate-semialdehyde dehydrogenase | | Authors: | Yuan, Y.A, Yuan, Z, Yin, B, Wei, D. | | Deposit date: | 2012-10-09 | | Release date: | 2013-07-10 | | Last modified: | 2023-11-08 | | Method: | X-RAY DIFFRACTION (1.69 Å) | | Cite: | Structural basis for cofactor and substrate selection by cyanobacterium succinic semialdehyde dehydrogenase
J.Struct.Biol., 182, 2013
|
|
3VZ2
 
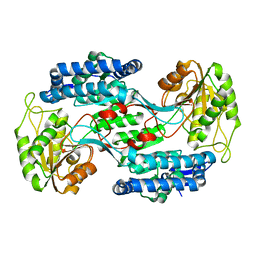 | |
3VNB
 
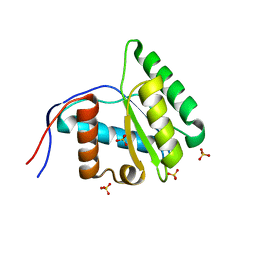 | |
3VZ0
 
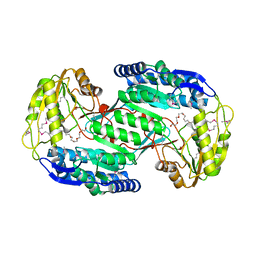 | | Structural insights into cofactor and substrate selection by Gox0499 | | Descriptor: | NONAETHYLENE GLYCOL, Putative NAD-dependent aldehyde dehydrogenase | | Authors: | Yuan, Y.A, Yuan, Z, Yin, B, Wei, D. | | Deposit date: | 2012-10-09 | | Release date: | 2013-07-10 | | Method: | X-RAY DIFFRACTION (2.3 Å) | | Cite: | Structural basis for cofactor and substrate selection by cyanobacterium succinic semialdehyde dehydrogenase
J.Struct.Biol., 182, 2013
|
|
3WA8
 
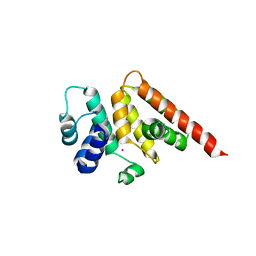 | | Crystal structure of M. ruber CasB | | Descriptor: | CRISPR-associated protein, Cse2 family, MERCURY (II) ION | | Authors: | Yuan, Y.A, Yuan, Z. | | Deposit date: | 2013-04-28 | | Release date: | 2014-04-30 | | Last modified: | 2024-03-20 | | Method: | X-RAY DIFFRACTION (2.8 Å) | | Cite: | Structural insights into crRNA G-rich sequence binding and R-loop formation facilitated by Meiothermus ruber CasB
To be Published
|
|
3VPY
 
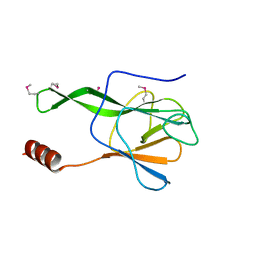 | | Crystal structure of Arabidopsis DDL FHA domain | | Descriptor: | FHA domain-containing protein DDL | | Authors: | Yuan, Y.A, Machida, S. | | Deposit date: | 2012-03-15 | | Release date: | 2013-02-06 | | Method: | X-RAY DIFFRACTION (1.7 Å) | | Cite: | Crystal Structure of Arabidopsis thaliana Dawdle Forkhead-Associated Domain reveals a conserved phospho-threonine recognition cleft for Dicer-like1 binding.
Mol Plant, 2013
|
|
5B1Z
 
 | |
5B4M
 
 | |
3UST
 
 | | Structure of BmNPV ORF075 (p33) | | Descriptor: | AcMNPV orf92, FLAVIN-ADENINE DINUCLEOTIDE | | Authors: | Yuan, Y.A, Hou, Y, Xia, Q. | | Deposit date: | 2011-11-23 | | Release date: | 2012-10-10 | | Method: | X-RAY DIFFRACTION (2.1 Å) | | Cite: | Crystal structure of Bombyx mori nucleopolyhedrovirus ORF75 reveals a pseudo-dimer of thiol oxidase domains with a putative substrate-binding pocket
J.Gen.Virol., 93, 2012
|
|
3ADG
 
 | |
3ADJ
 
 | |
3AIA
 
 | | Crystal structure of DUF358 reveals a putative SPOUT-class methltransferase | | Descriptor: | S-ADENOSYLMETHIONINE, UPF0217 protein MJ1640, pentane-2,2,4,4-tetrol | | Authors: | Yuan, Y.A, Chen, H.Y. | | Deposit date: | 2010-05-11 | | Release date: | 2011-03-30 | | Last modified: | 2023-11-01 | | Method: | X-RAY DIFFRACTION (1.4 Å) | | Cite: | Crystal structure of Mj1640/DUF358 protein reveals a putative SPOUT-class RNA methyltransferase
J Mol Cell Biol, 2, 2010
|
|
3ADI
 
 | |
3ADL
 
 | |
2ZI0
 
 | | Crystal structure of Tav2b/siRNA complex | | Descriptor: | Protein 2b, RNA (5'-D(P*AP*GP*AP*CP*AP*GP*CP*AP*UP*UP*AP*UP*GP*CP*UP*GP*UP*CP*UP*UP*U)-3') | | Authors: | Yuan, Y.A, Chen, H.-Y. | | Deposit date: | 2008-02-12 | | Release date: | 2008-07-22 | | Last modified: | 2011-07-13 | | Method: | X-RAY DIFFRACTION (2.82 Å) | | Cite: | Structural basis for RNA-silencing suppression by Tomato aspermy virus protein 2b
Embo Rep., 9, 2008
|
|
2ZFN
 
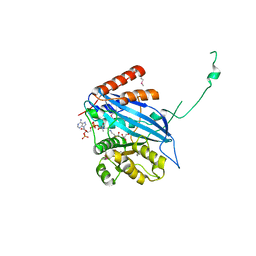 | |
3AI9
 
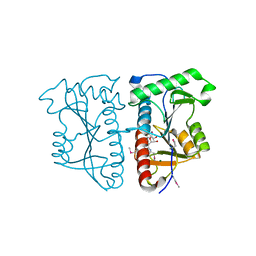 | |
3AJF
 
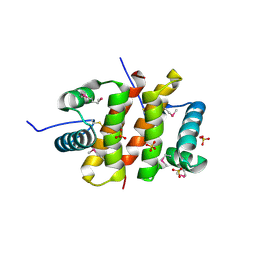 | |
2ZKO
 
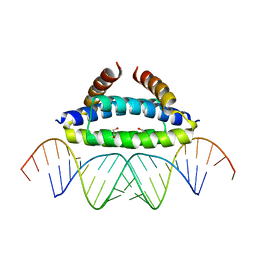 | |
3AX1
 
 | |
3AXJ
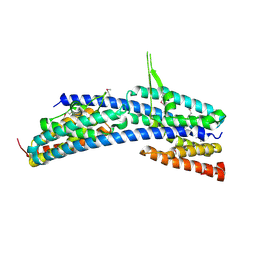 
 | | High resolution crystal structure of C3PO | | Descriptor: | GM27569p, Translin associated factor X, isoform B | | Authors: | Yuan, Y.A, Yang, X. | | Deposit date: | 2011-04-07 | | Release date: | 2011-04-20 | | Last modified: | 2011-07-13 | | Method: | X-RAY DIFFRACTION (2.1 Å) | | Cite: | High resolution crystal structure of C3PO
To be Published
|
|
3VYY
 
 | |
3VNA
 
 | |
