3VA0
 
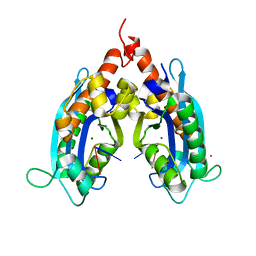 | |
3V9X
 
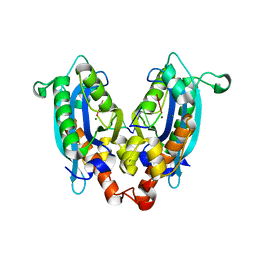 | |
3TAT
 | | TYROSINE AMINOTRANSFERASE FROM E. COLI | | Descriptor: | PYRIDOXAL-5'-PHOSPHATE, TYROSINE AMINOTRANSFERASE | | Authors: | Ko, T.P, Yang, W.Z, Wu, S.P, Tsai, H, Yuan, H.S. | | Deposit date: | 1998-08-12 | | Release date: | 1999-08-12 | | Last modified: | 2023-08-09 | | Method: | X-RAY DIFFRACTION (3.5 Å) | | Cite: | Crystallization and preliminary crystallographic analysis of the Escherichia coli tyrosine aminotransferase.
Acta Crystallogr.,Sect.D, 55, 1999
|
|
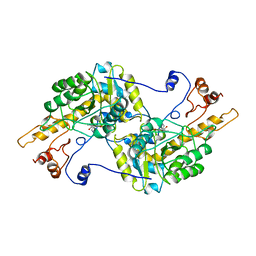
3U1K
 
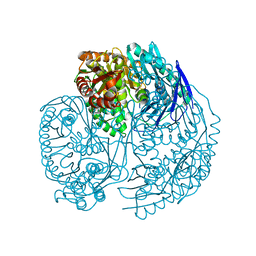 | | Crystal structure of human PNPase | | Descriptor: | CITRIC ACID, Polyribonucleotide nucleotidyltransferase 1, mitochondrial | | Authors: | Lin, C.L, Yuan, H.S. | | Deposit date: | 2011-09-30 | | Release date: | 2012-02-01 | | Last modified: | 2023-11-01 | | Method: | X-RAY DIFFRACTION (2.13 Å) | | Cite: | Crystal structure of human polynucleotide phosphorylase: insights into its domain function in RNA binding and degradation
Nucleic Acids Res., 40, 2012
|
|
3V9Z
 
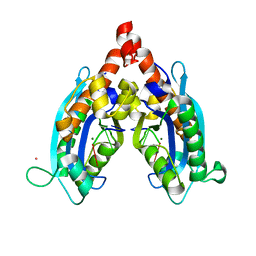 | |
3V9S
 
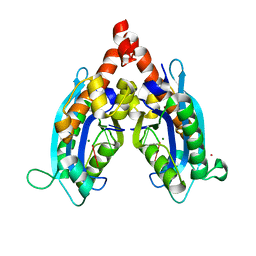 | |
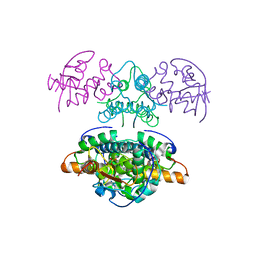
2JBG
 
 | |
2JB0
 
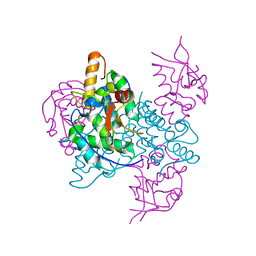 | |
1ETQ
 
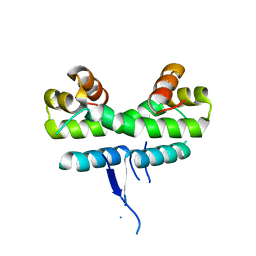 | |
1ETV
 
 | |
1ETW
 
 | |
1ETX
 
 | |
1ETK
 
 | |
1ETY
 
 | |
1CEI
 
 | | STRUCTURE DETERMINATION OF THE COLICIN E7 IMMUNITY PROTEIN (IMME7) THAT BINDS SPECIFICALLY TO THE DNASE-TYPE COLICIN E7 AND INHIBITS ITS BACTERIOCIDAL ACTIVITY | | Descriptor: | COLICIN E7 IMMUNITY PROTEIN | | Authors: | Chak, K.-F, Safo, M.K, Ku, W.-Y, Hsieh, S.-Y, Yuan, H.S. | | Deposit date: | 1996-03-19 | | Release date: | 1997-03-12 | | Last modified: | 2024-02-07 | | Method: | X-RAY DIFFRACTION (1.8 Å) | | Cite: | The crystal structure of the immunity protein of colicin E7 suggests a possible colicin-interacting surface.
Proc.Natl.Acad.Sci.USA, 93, 1996
|
|
1ETO
 
 | |
1F36
 
 | |
5ZKI
 
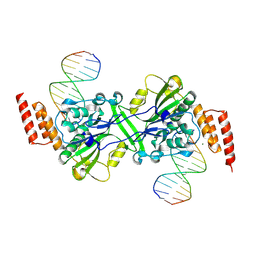 | | Human EXOG-H140A in complex with duplex DNA | | Descriptor: | CHLORIDE ION, DNA (5'-D(*CP*GP*GP*GP*AP*TP*AP*TP*CP*CP*CP*G)-3'), MAGNESIUM ION, ... | | Authors: | Wu, C.C, Lin, J.L.J, Yuan, H.S. | | Deposit date: | 2018-03-24 | | Release date: | 2019-04-03 | | Last modified: | 2023-11-22 | | Method: | X-RAY DIFFRACTION (2.321 Å) | | Cite: | A unique exonuclease ExoG cleaves between RNA and DNA in mitochondrial DNA replication.
Nucleic Acids Res., 47, 2019
|
|
5ZKJ
 
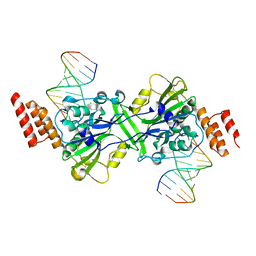 | | Human EXOG-H140A in complex with RNA/DNA hybrid duplex | | Descriptor: | CHLORIDE ION, DNA (5'-D(*CP*GP*TP*GP*AP*CP*AP*TP*CP*CP*CP*G)-3'), MAGNESIUM ION, ... | | Authors: | Wu, C.C, Lin, J.L.J, Yuan, H.S. | | Deposit date: | 2018-03-24 | | Release date: | 2019-04-03 | | Last modified: | 2023-11-22 | | Method: | X-RAY DIFFRACTION (2.798 Å) | | Cite: | A unique exonuclease ExoG cleaves between RNA and DNA in mitochondrial DNA replication.
Nucleic Acids Res., 47, 2019
|
|
7DA6
 
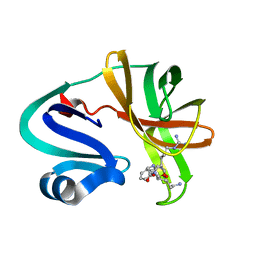 | |
7W1R
 
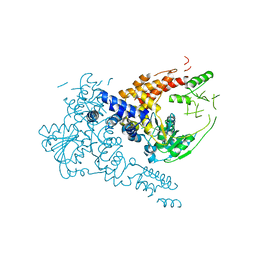 | | Crystal structure of human Suv3 monomer | | Descriptor: | ATP-dependent RNA helicase SUPV3L1, mitochondrial | | Authors: | Jain, M, Yuan, H.S. | | Deposit date: | 2021-11-20 | | Release date: | 2022-05-11 | | Last modified: | 2023-11-29 | | Method: | X-RAY DIFFRACTION (3.2 Å) | | Cite: | Dimeric assembly of human Suv3 helicase promotes its RNA unwinding function in mitochondrial RNA degradosome for RNA decay.
Protein Sci., 31, 2022
|
|
6IID
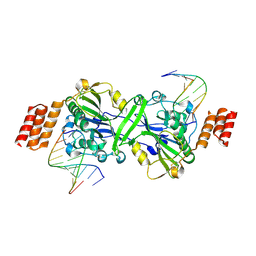 
 | | Human EXOG-H140A in complex with RNA-DNA chimeric duplex | | Descriptor: | DNA (5'-D(*CP*GP*TP*GP*AP*CP*AP*TP*CP*CP*CP*G)-3'), DNA/RNA (5'-R(P*CP*GP*GP*GP*A)-D(P*T)-R(P*G)-D(P*T)-R(P*CP*AP*CP*G)-3'), MAGNESIUM ION, ... | | Authors: | Wu, C.C, Lin, J.L.J, Yuan, H.S. | | Deposit date: | 2018-10-04 | | Release date: | 2019-04-03 | | Last modified: | 2023-11-22 | | Method: | X-RAY DIFFRACTION (2.986 Å) | | Cite: | A unique exonuclease ExoG cleaves between RNA and DNA in mitochondrial DNA replication.
Nucleic Acids Res., 47, 2019
|
|
6J7Y
 
 | | Human mitochondrial Oligoribonuclease in complex with DNA | | Descriptor: | DNA (5'-D(P*TP*T)-3'), MAGNESIUM ION, Oligoribonuclease, ... | | Authors: | Chu, L.Y, Agrawal, S, Yuan, H.S. | | Deposit date: | 2019-01-18 | | Release date: | 2019-08-28 | | Last modified: | 2023-11-22 | | Method: | X-RAY DIFFRACTION (2.203 Å) | | Cite: | Structural insights into nanoRNA degradation by human Rexo2.
Rna, 25, 2019
|
|
6J80
 
 | | Human mitochondrial Oligoribonuclease in complex with poly-dT DNA | | Descriptor: | CITRIC ACID, DNA (5'-D(P*TP*TP*TP*TP*TP*TP*T)-3'), MAGNESIUM ION, ... | | Authors: | Chu, L.Y, Agrawal, S, Yuan, H.S. | | Deposit date: | 2019-01-18 | | Release date: | 2019-08-28 | | Last modified: | 2023-11-22 | | Method: | X-RAY DIFFRACTION (1.812 Å) | | Cite: | Structural insights into nanoRNA degradation by human Rexo2.
Rna, 25, 2019
|
|
6J7Z
 
 | | Human mitochondrial Oligoribonuclease in complex with RNA | | Descriptor: | CITRIC ACID, MAGNESIUM ION, Oligoribonuclease, ... | | Authors: | Chu, L.Y, Agrawal, S, Yuan, H.S. | | Deposit date: | 2019-01-18 | | Release date: | 2019-08-28 | | Last modified: | 2023-11-22 | | Method: | X-RAY DIFFRACTION (2.005 Å) | | Cite: | Structural insights into nanoRNA degradation by human Rexo2.
Rna, 25, 2019
|
|
