6CT6
 
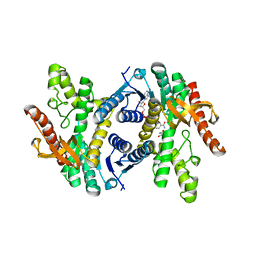 | | Crystal structure of lactate dehydrogenase from Eimeria maxima with NADH and oxamate | | Descriptor: | 1,4-DIHYDRONICOTINAMIDE ADENINE DINUCLEOTIDE, Lactate dehydrogenase, OXAMIC ACID, ... | | Authors: | Wirth, J.D, Xu, C, Theobald, D.L. | | Deposit date: | 2018-03-22 | | Release date: | 2019-03-27 | | Last modified: | 2023-10-04 | | Method: | X-RAY DIFFRACTION (1.705 Å) | | Cite: | The Mechanistic, Structural, and Evolutionary Origin of Lactate Dehydrogenase Substrate Specificity in Apicomplexa
To Be Published
|
|
7SJ6
 
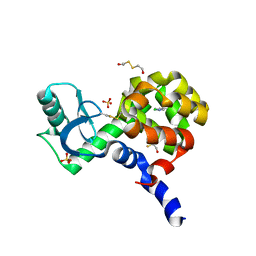 | | T4 Lysozyme L99A/M102H with 1,2-Azaborine bound | | Descriptor: | 1,2-dihydro-1,2-azaborinine, 2-HYDROXYETHYL DISULFIDE, ACETATE ION, ... | | Authors: | Yao, L, Wirth, J. | | Deposit date: | 2021-10-16 | | Release date: | 2022-10-26 | | Last modified: | 2023-11-29 | | Method: | X-RAY DIFFRACTION (1.72 Å) | | Cite: | T4 Lysozyme L99A/M102H with 1,2-Azaborine bound
to be published
|
|
6VDH
 
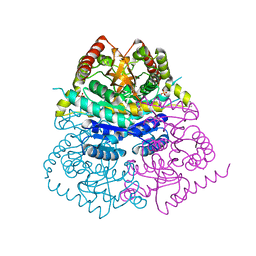 | |
6VDI
 
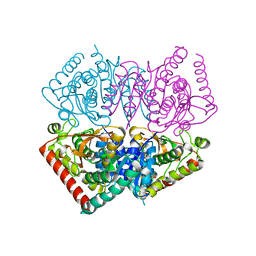 | |
6VDJ
 
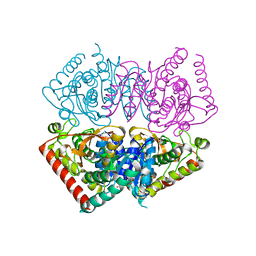 | | Crystal structure of ancestral apicomplexan lactate dehydrogenase with sulfate and NADH4. | | Descriptor: | 1,4,5,6-TETRAHYDRONICOTINAMIDE ADENINE DINUCLEOTIDE, SULFATE ION, lactate dehydrogenase | | Authors: | Theobald, D.L, Wirth, J.D, Classen, S, Perlmutter, N. | | Deposit date: | 2019-12-27 | | Release date: | 2020-01-29 | | Last modified: | 2023-10-11 | | Method: | X-RAY DIFFRACTION (2 Å) | | Cite: | Mechanism of Bifunctionality in an Ancestral Apicomplexan Malate/Lactate Dehydrogenase
To Be Published
|
|
5OMF
 
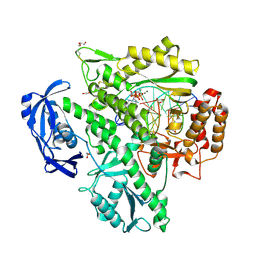 | | Closed, ternary structure of KOD DNA polymerase | | Descriptor: | 1,2-ETHANEDIOL, 2'-DEOXYADENOSINE 5'-TRIPHOSPHATE, DNA (5'-D(*GP*AP*CP*CP*AP*CP*GP*GP*CP*CP*AP*C)-3'), ... | | Authors: | Kropp, H.M, Betz, K, Wirth, J, Diederichs, K, Marx, A. | | Deposit date: | 2017-07-31 | | Release date: | 2017-12-20 | | Last modified: | 2024-01-17 | | Method: | X-RAY DIFFRACTION (2.092 Å) | | Cite: | Crystal structures of ternary complexes of archaeal B-family DNA polymerases.
PLoS ONE, 12, 2017
|
|
8TFN
 
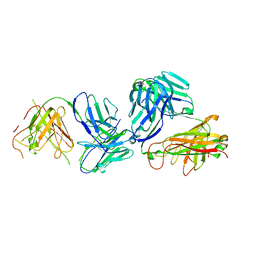 | | Structure of anti-TCRvbeta6-5 antibody in complex with the cognate TCR | | Descriptor: | Anti-TCRVb6-5 Fab heavy chain, Anti-TCRVb6-5 Fab light chain, TRAV12-3, ... | | Authors: | Katragadda, M, Servatalab, R, Wirth, J. | | Deposit date: | 2023-07-11 | | Release date: | 2023-11-08 | | Last modified: | 2024-05-22 | | Method: | X-RAY DIFFRACTION (2.54 Å) | | Cite: | Innate TCR beta-chain engagement drives human T cells toward distinct memory-like effector phenotypes with immunotherapeutic potentials.
Sci Adv, 9, 2023
|
|
7NBN
 
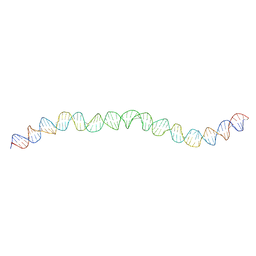 | | Allostery through DNA drives phenotype switching | | Descriptor: | AddAB promoter | | Authors: | Rosenblum, G, Elad, N, Rozenberg, H, Wiggers, F, Jungwirth, J, Hofmann, H. | | Deposit date: | 2021-01-27 | | Release date: | 2021-04-07 | | Last modified: | 2024-05-01 | | Method: | ELECTRON MICROSCOPY (7 Å) | | Cite: | Allostery through DNA drives phenotype switching.
Nat Commun, 12, 2021
|
|
6Z0S
 
 | | Allostery through DNA drives phenotype switching | | Descriptor: | comG promoter DNA - strand A, comG promoter DNA - strand B | | Authors: | Rosenblum, G, Elad, N, Rozenberg, H, Wiggers, F, Jungwirth, J, Hofmann, H. | | Deposit date: | 2020-05-11 | | Release date: | 2021-04-07 | | Last modified: | 2024-05-22 | | Method: | ELECTRON MICROSCOPY (5.7 Å) | | Cite: | Allostery through DNA drives phenotype switching.
Nat Commun, 12, 2021
|
|
