1E0V
 
 | | Xylanase 10A from Sreptomyces lividans. cellobiosyl-enzyme intermediate at 1.7 A | | Descriptor: | ENDO-1,4-BETA-XYLANASE A, beta-D-glucopyranose-(1-4)-2-deoxy-2-fluoro-alpha-D-glucopyranose | | Authors: | Ducros, V, Charnock, S.J, Derewenda, U, Derewenda, Z.S, Dauter, Z, Dupont, C, Shareck, F, Morosoli, R, Kluepfel, D, Davies, G.J. | | Deposit date: | 2000-04-10 | | Release date: | 2001-04-05 | | Last modified: | 2024-05-01 | | Method: | X-RAY DIFFRACTION (1.7 Å) | | Cite: | Substrate Specificity in Glycoside Hydrolase Family 10. Structural and Kinetic Analysis of the Streptomyces Lividans Xylanase 10A
J.Biol.Chem., 275, 2000
|
|
1E0X
 
 | | XYLANASE 10A FROM SREPTOMYCES LIVIDANS. XYLOBIOSYL-ENZYME INTERMEDIATE AT 1.65 A | | Descriptor: | ENDO-1,4-BETA-XYLANASE A, GLYCEROL, beta-D-xylopyranose-(1-4)-2-deoxy-2-fluoro-alpha-D-xylopyranose | | Authors: | Ducros, V, Charnock, S.J, Derewenda, U, Derewenda, Z.S, Dauter, Z, Dupont, C, Shareck, F, Morosoli, R, Kluepfel, D, Davies, G.J. | | Deposit date: | 2000-04-10 | | Release date: | 2001-04-05 | | Last modified: | 2024-05-01 | | Method: | X-RAY DIFFRACTION (1.65 Å) | | Cite: | Substrate Specificity in Glycoside Hydrolase Family 10. Structural and Kinetic Analysis of the Streptomyces Lividans Xylanase 10A
J.Biol.Chem., 275, 2000
|
|
5YPR
 
 | | Crystal Structure of PSD-95 SH3-GK domain in complex with a synthesized inhibitor | | Descriptor: | Disks large homolog 4, Synthesized GK inhibitor | | Authors: | Zhu, J, Zhou, Q, Shang, Y, Weng, Z, Zhu, R, Zhang, M. | | Deposit date: | 2017-11-02 | | Release date: | 2018-03-14 | | Last modified: | 2024-10-30 | | Method: | X-RAY DIFFRACTION (2.349 Å) | | Cite: | Synaptic Targeting and Function of SAPAPs Mediated by Phosphorylation-Dependent Binding to PSD-95 MAGUKs.
Cell Rep, 21, 2017
|
|
1E0W
 
 | | Xylanase 10A from Sreptomyces lividans. native structure at 1.2 angstrom resolution | | Descriptor: | ENDO-1,4-BETA-XYLANASE A | | Authors: | Ducros, V, Charnock, S.J, Derewenda, U, Derewenda, Z.S, Dauter, Z, Dupont, C, Shareck, F, Morosoli, R, Kluepfel, D, Davies, G.J. | | Deposit date: | 2000-04-10 | | Release date: | 2001-04-05 | | Last modified: | 2014-02-05 | | Method: | X-RAY DIFFRACTION (1.2 Å) | | Cite: | Substrate Specificity in Glycoside Hydrolase Family 10. Structural and Kinetic Analysis of the Streptomyces Lividans Xylanase 10A
J.Biol.Chem., 275, 2000
|
|
3N0U
 
 | |
2BXW
 | | CRYSTAL STRUCTURE OF RHOGDI Lys(135,138,141)Tyr MUTANT | | Descriptor: | 2-AMINO-2-HYDROXYMETHYL-PROPANE-1,3-DIOL, FORMIC ACID, RHO GDP-DISSOCIATION INHIBITOR 1 | | Authors: | Sikorska, M, Cooper, D.R, Otlewski, J, Derewenda, Z.S. | | Deposit date: | 2005-07-27 | | Release date: | 2005-08-19 | | Last modified: | 2023-12-13 | | Method: | X-RAY DIFFRACTION (2.4 Å) | | Cite: | Protein Crystallization by Surface Entropy Reduction: Optimization of the Ser Strategy
Acta Crystallogr.,Sect.D, 63, 2007
|
|
3E57
 
 | |
2BR6
 
 | | Crystal Structure of Quorum-Quenching N-Acyl Homoserine Lactone Lactonase | | Descriptor: | AIIA-LIKE PROTEIN, GLYCEROL, HOMOSERINE LACTONE, ... | | Authors: | Kim, M.H, Choi, W.C, Kang, H.O, Kang, B.S, Kim, K.J, Derewenda, Z.S, Lee, J.K, Oh, T.K, Lee, C.H. | | Deposit date: | 2005-05-03 | | Release date: | 2005-12-07 | | Last modified: | 2024-05-08 | | Method: | X-RAY DIFFRACTION (1.7 Å) | | Cite: | The Molecular Structure and Catalytic Mechanism of a Quorum-Quenching N-Acyl-L-Homoserine Lactone Hydrolase.
Proc.Natl.Acad.Sci.USA, 102, 2005
|
|
6YOO
 
 | | Structure of SAMM50 LIR bound to GABARAPL1 | | Descriptor: | 1,2-ETHANEDIOL, CHLORIDE ION, Gamma-aminobutyric acid receptor-associated protein-like 1, ... | | Authors: | Mouilleron, S, Wenxin, Z, Johansen, T, Tooze, S, Abudu, Y.P, Lamark, T. | | Deposit date: | 2020-04-14 | | Release date: | 2021-04-28 | | Last modified: | 2024-01-24 | | Method: | X-RAY DIFFRACTION (1.06 Å) | | Cite: | Structure of SAMM50 LIR bound to GABARAPL1
To Be Published
|
|
1KMQ
 
 | | Crystal Structure of a Constitutively Activated RhoA Mutant (Q63L) | | Descriptor: | 1,4-DIETHYLENE DIOXIDE, MAGNESIUM ION, PHOSPHOAMINOPHOSPHONIC ACID-GUANYLATE ESTER, ... | | Authors: | Longenecker, K, Read, P, Lin, S.-K, Somlyo, A.P, Nakamoto, R.K, Derewenda, Z.S. | | Deposit date: | 2001-12-17 | | Release date: | 2003-05-06 | | Last modified: | 2024-02-14 | | Method: | X-RAY DIFFRACTION (1.55 Å) | | Cite: | Structure of a constitutively activated RhoA mutant (Q63L) at 1.55 A resolution.
Acta Crystallogr.,Sect.D, 59, 2003
|
|
2BTN
 
 | | Crystal Structure and Catalytic Mechanism of the Quorum-Quenching N- Acyl Homoserine Lactone Hydrolase | | Descriptor: | AIIA-LIKE PROTEIN, GLYCEROL, ZINC ION | | Authors: | Kim, M.H, Choi, W.C, Kang, H.O, Lee, J.S, Kang, B.S, Kim, K.J, Derewenda, Z.S, Oh, T.K, Lee, C.H, Lee, J.K. | | Deposit date: | 2005-06-03 | | Release date: | 2005-12-07 | | Last modified: | 2024-05-08 | | Method: | X-RAY DIFFRACTION (2 Å) | | Cite: | The Molecular Structure and Catalytic Mechanism of a Quorum-Quenching N-Acyl-L-Homoserine Lactone Hydrolase.
Proc.Natl.Acad.Sci.USA, 102, 2005
|
|
1TGL
 
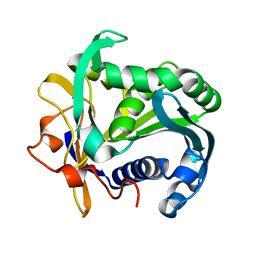 | | A SERINE PROTEASE TRIAD FORMS THE CATALYTIC CENTRE OF A TRIACYLGLYCEROL LIPASE | | Descriptor: | TRIACYL-GLYCEROL ACYLHYDROLASE | | Authors: | Brady, L, Brzozowski, A.M, Derewenda, Z.S, Dodson, E.J, Dodson, G.G, Tolley, S.P, Turkenburg, J.P, Christiansen, L, Huge-Jensen, B, Norskov, L, Thim, L. | | Deposit date: | 1990-02-05 | | Release date: | 1990-10-15 | | Last modified: | 2024-02-14 | | Method: | X-RAY DIFFRACTION (1.9 Å) | | Cite: | A serine protease triad forms the catalytic centre of a triacylglycerol lipase.
Nature, 343, 1990
|
|
3GOR
 
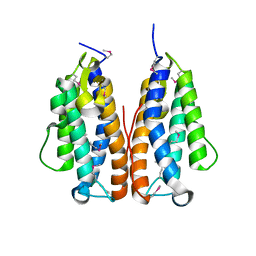 | | Crystal structure of putative metal-dependent hydrolase APC36150 | | Descriptor: | NICKEL (II) ION, Putative metal-dependent hydrolase | | Authors: | Cooper, D.R, Grelewska, K, Derewenda, Z.S, Integrated Center for Structure and Function Innovation (ISFI) | | Deposit date: | 2009-03-19 | | Release date: | 2009-05-19 | | Last modified: | 2024-10-30 | | Method: | X-RAY DIFFRACTION (2.511 Å) | | Cite: | The structure of DinB from Geobacillus stearothermophilus: a representative of a unique four-helix-bundle superfamily.
Acta Crystallogr.,Sect.F, 66, 2010
|
|
1JFR
 
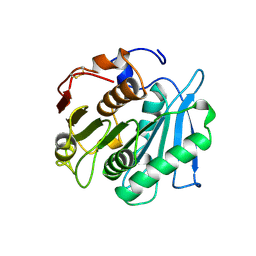 | |
1W9Q
 
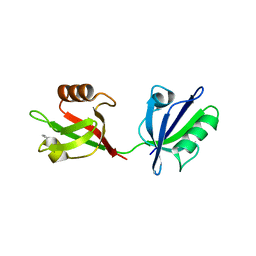 | | Crystal structure of the PDZ tandem of human syntenin in complex with TNEFAF peptide | | Descriptor: | BENZOIC ACID, SYNTENIN 1, TNEFAF PEPTIDE | | Authors: | Grembecka, J, Cierpicki, T, Devedjiev, Y, Cooper, D.R, Derewenda, Z.S. | | Deposit date: | 2004-10-15 | | Release date: | 2006-03-22 | | Last modified: | 2023-12-13 | | Method: | X-RAY DIFFRACTION (1.7 Å) | | Cite: | The Binding of the Pdz Tandem of Syntenin to Target Proteins.
Biochemistry, 45, 2006
|
|
1VYH
 
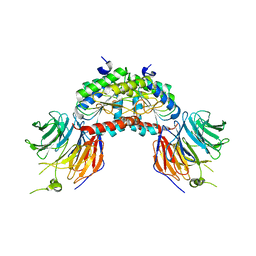 | | PAF-AH Holoenzyme: Lis1/Alfa2 | | Descriptor: | PLATELET-ACTIVATING FACTOR ACETYLHYDROLASE IB ALPHA SUBUNIT, PLATELET-ACTIVATING FACTOR ACETYLHYDROLASE IB BETA SUBUNIT | | Authors: | Tarricone, C, Perrina, F, Monzani, S, Massimiliano, L, Knapp, S, Tsai, L.-H, Derewenda, Z.S, Musacchio, A. | | Deposit date: | 2004-04-30 | | Release date: | 2005-05-26 | | Last modified: | 2023-12-13 | | Method: | X-RAY DIFFRACTION (3.4 Å) | | Cite: | Coupling Paf Signaling to Dynein Regulation: Structure of Lis1 in Complex with Paf-Acetylhydrolase.
Neuron, 44, 2004
|
|
7W7P
 | |
7YOX
 
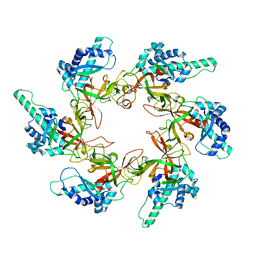 | |
2JHX
 | |
2JHY
 
 | |
2JHW
 
 | | CRYSTAL STRUCTURE OF RHOGDI E155A, E157A MUTANT | | Descriptor: | RHO GDP-DISSOCIATION INHIBITOR 1, SULFATE ION | | Authors: | Cooper, D.R, Grelewska, K, Derewenda, Z.S. | | Deposit date: | 2007-02-23 | | Release date: | 2007-05-08 | | Last modified: | 2023-12-13 | | Method: | X-RAY DIFFRACTION (2.5 Å) | | Cite: | Protein Crystallization by Surface Entropy Reduction: Optimization of the Ser Strategy
Acta Crystallogr.,Sect.D, 63, 2007
|
|
2JHU
 
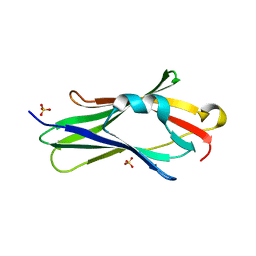 | | CRYSTAL STRUCTURE OF RHOGDI E154A,E155A MUTANT | | Descriptor: | RHO GDP-DISSOCIATION INHIBITOR 1, SULFATE ION | | Authors: | Cooper, D.R, Pinkowska, M, Derewenda, Z.S. | | Deposit date: | 2007-02-23 | | Release date: | 2007-05-08 | | Last modified: | 2023-12-13 | | Method: | X-RAY DIFFRACTION (1.65 Å) | | Cite: | Protein Crystallization by Surface Entropy Reduction: Optimization of the Ser Strategy
Acta Crystallogr.,Sect.D, 63, 2007
|
|
2JHV
 
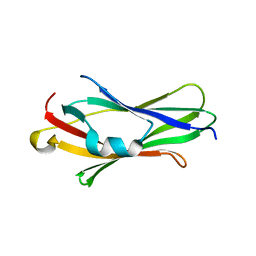 | |
2JI0
 
 | | CRYSTAL STRUCTURE OF RHOGDI K138Y, K141Y MUTANT | | Descriptor: | RHO GDP-DISSOCIATION INHIBITOR 1, SULFATE ION | | Authors: | Cooper, D.R, Boczek, T, Derewenda, Z.S. | | Deposit date: | 2007-02-23 | | Release date: | 2007-05-08 | | Last modified: | 2023-12-13 | | Method: | X-RAY DIFFRACTION (2.1 Å) | | Cite: | Protein Crystallization by Surface Entropy Reduction: Optimization of the Ser Strategy
Acta Crystallogr.,Sect.D, 63, 2007
|
|
2JHZ
 
 | |
