8VQB
 
 | |
8VQ9
 
 | |
8VQA
 
 | |
8WP9
 | | Small-heat shock protein from Methanocaldococcus jannaschii, Hsp16.5 | | Descriptor: | Small heat shock protein HSP16.5 | | Authors: | Lee, J, Ryu, B, Kim, T, Kim, K.K. | | Deposit date: | 2023-10-09 | | Release date: | 2023-12-27 | | Method: | ELECTRON MICROSCOPY (2.49 Å) | | Cite: | Cryo-EM structure of a 16.5-kDa small heat-shock protein from Methanocaldococcus jannaschii.
Int.J.Biol.Macromol., 258, 2024
|
|
9CL9
 
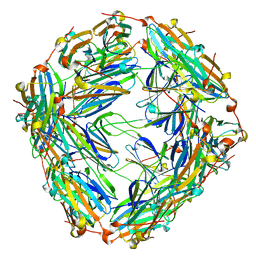
 | | WT 12C IM fraction, B-b3 with RluB bound | | Descriptor: | 23S rRNA, Large ribosomal subunit protein bL20, Large ribosomal subunit protein bL21, ... | | Authors: | Lee, J, Sheng, K, Williamson, J.R. | | Deposit date: | 2024-07-10 | | Release date: | 2024-07-24 | | Method: | ELECTRON MICROSCOPY (5.04 Å) | | Cite: | 50S ribosome assembly intermediates at low temperature reveal bound RluB
To Be Published
|
|
3BAG
 
 | |
3BAU
 
 | |
3BAQ
 
 | |
3BAH
 
 | |
3BAO
 
 | |
3BAV
 
 | |
3BB2
 | |
2R64
 
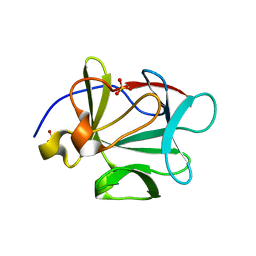
 | | Crystal structure of a 3-aminoindazole compound with CDK2 | | Descriptor: | Cell division protein kinase 2, N-[5-(1,1-DIOXIDOISOTHIAZOLIDIN-2-YL)-1H-INDAZOL-3-YL]-2-(4-PIPERIDIN-1-YLPHENYL)ACETAMIDE | | Authors: | Lee, J, Choi, H, Kim, K.H, Jeong, S, Park, J.W, Baek, C.S, Lee, S.H. | | Deposit date: | 2007-09-05 | | Release date: | 2008-09-09 | | Last modified: | 2024-03-13 | | Method: | X-RAY DIFFRACTION (2.3 Å) | | Cite: | Synthesis and biological evaluation of 3,5-diaminoindazoles as cyclin-dependent kinase inhibitors.
Bioorg.Med.Chem.Lett., 18, 2008
|
|
7Y7O
 | |
7WBK
 
 | |
7WBM
 
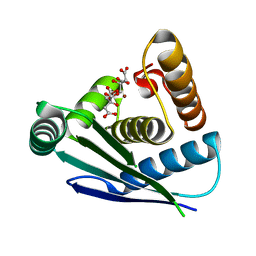
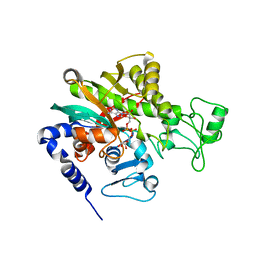 | | Crystal structure of Legionella pneumophila effector protein Lpg0081 | | Descriptor: | Lpg0081, [(2R,3S,4R,5R)-5-(6-AMINOPURIN-9-YL)-3,4-DIHYDROXY-OXOLAN-2-YL]METHYL [HYDROXY-[[(2R,3S,4R,5S)-3,4,5-TRIHYDROXYOXOLAN-2-YL]METHOXY]PHOSPHORYL] HYDROGEN PHOSPHATE | | Authors: | Lee, J, Kim, H, Oh, B.H. | | Deposit date: | 2021-12-17 | | Release date: | 2022-06-15 | | Last modified: | 2023-11-29 | | Method: | X-RAY DIFFRACTION (2.7 Å) | | Cite: | Reversible modification of mitochondrial ADP/ATP translocases by paired Legionella effector proteins.
Proc.Natl.Acad.Sci.USA, 119, 2022
|
|
5CRV
 
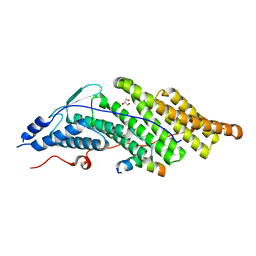 | | Crystal structure of the Bro domain of HD-PTP in a complex with the core region of STAM2 | | Descriptor: | GLYCEROL, Signal transducing adapter molecule 2, Tyrosine-protein phosphatase non-receptor type 23 | | Authors: | Lee, J, Ku, B, Kim, S.J. | | Deposit date: | 2015-07-23 | | Release date: | 2016-02-24 | | Last modified: | 2023-11-08 | | Method: | X-RAY DIFFRACTION (2.001 Å) | | Cite: | Structural Study of the HD-PTP Bro1 Domain in a Complex with the Core Region of STAM2, a Subunit of ESCRT-0
Plos One, 11, 2016
|
|
5CRU
 
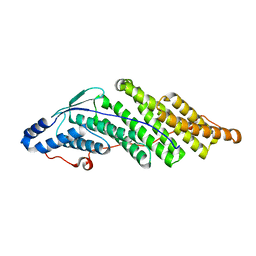 | | Crystal structure of the Bro domain of HD-PTP | | Descriptor: | Tyrosine-protein phosphatase non-receptor type 23 | | Authors: | Lee, J, Ku, B, Kim, S.J. | | Deposit date: | 2015-07-23 | | Release date: | 2016-02-24 | | Last modified: | 2023-11-08 | | Method: | X-RAY DIFFRACTION (2.4 Å) | | Cite: | Structural Study of the HD-PTP Bro1 Domain in a Complex with the Core Region of STAM2, a Subunit of ESCRT-0
Plos One, 11, 2016
|
|
4QXL
 
 | | Crystal Structure of FLHE | | Descriptor: | Flagellar protein flhE | | Authors: | Lee, J, Monzingo, A.F, Keatinge-Clay, A.T, Harshey, R.M. | | Deposit date: | 2014-07-21 | | Release date: | 2015-01-14 | | Last modified: | 2017-11-22 | | Method: | X-RAY DIFFRACTION (1.512 Å) | | Cite: | Structure of Salmonella FlhE, Conserved Member of a Flagellar Type III Secretion Operon.
J.Mol.Biol., 427, 2015
|
|
8T4H
 
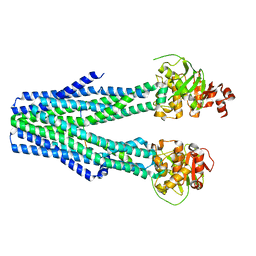 | |
8T46
 
 | |
8T4F
 
 | |
8T4G
 
 | |
8T4I
 
 | |
8T4J
 
 | |
