5BOX
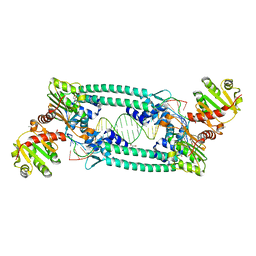 
 | | Structure of TrmBL2, an archaeal chromatin protein, shows a novel mode of DNA binding. | | Descriptor: | (4S)-2-METHYL-2,4-PENTANEDIOL, DNA (25-MER), DNA TGM (25-MER), ... | | Authors: | Ahmad, M.U, Diederichs, K, Welte, W. | | Deposit date: | 2015-05-27 | | Release date: | 2015-09-02 | | Last modified: | 2024-01-10 | | Method: | X-RAY DIFFRACTION (2.5 Å) | | Cite: | Structural Insights into Nonspecific Binding of DNA by TrmBL2, an Archaeal Chromatin Protein.
J.Mol.Biol., 427, 2015
|
|
4DF8
 
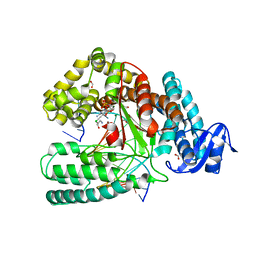 | | Crystal structure of the large fragment of DNA Polymerase I from Thermus aquaticus in a closed ternary complex with aminopentinyl-7-deaza-2-dATP | | Descriptor: | 1,2-ETHANEDIOL, 5-(5-aminopent-1-yn-1-yl)-7-{2-deoxy-5-O-[(S)-hydroxy{[(S)-hydroxy(phosphonooxy)phosphoryl]oxy}phosphoryl]-beta-D-erythro-pentofuranosyl}-7H-pyrrolo[2,3-d]pyrimidin-4-amine, CHLORIDE ION, ... | | Authors: | Bergen, K, Steck, A, Struett, S, Baccaro, A, Welte, W, Diederichs, K, Marx, A. | | Deposit date: | 2012-01-23 | | Release date: | 2012-05-16 | | Last modified: | 2023-09-13 | | Method: | X-RAY DIFFRACTION (2 Å) | | Cite: | Structures of KlenTaq DNA Polymerase Caught While Incorporating C5-Modified Pyrimidine and C7-Modified 7-Deazapurine Nucleoside Triphosphates.
J.Am.Chem.Soc., 134, 2012
|
|
4DF4
 
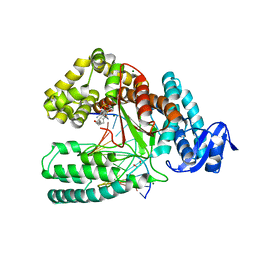 | | Crystal structure of the large fragment of DNA Polymerase I from Thermus aquaticus in a closed ternary complex with 7-(N-(10-hydroxydecanoyl)-aminopentinyl)-7-deaza-2 -dATP | | Descriptor: | 1,2-ETHANEDIOL, 7-{2-deoxy-5-O-[(S)-hydroxy{[(S)-hydroxy(phosphonooxy)phosphoryl]oxy}phosphoryl]-beta-D-erythro-pentofuranosyl}-5-{5-[(10-hydroxydecanoyl)amino]pent-1-yn-1-yl}-7H-pyrrolo[2,3-d]pyrimidin-4-amine, DNA (5'-D(*AP*AP*AP*TP*GP*GP*CP*GP*CP*CP*GP*TP*GP*GP*TP*C)-3'), ... | | Authors: | Bergen, K, Steck, A, Struett, S, Baccaro, A, Welte, W, Diederichs, K, Marx, A. | | Deposit date: | 2012-01-23 | | Release date: | 2012-05-16 | | Last modified: | 2023-09-13 | | Method: | X-RAY DIFFRACTION (2.2 Å) | | Cite: | Structures of KlenTaq DNA Polymerase Caught While Incorporating C5-Modified Pyrimidine and C7-Modified 7-Deazapurine Nucleoside Triphosphates.
J.Am.Chem.Soc., 134, 2012
|
|
1A0S
 
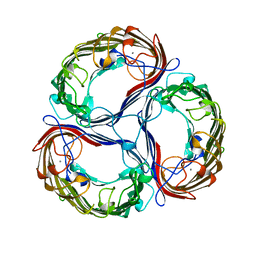 | | SUCROSE-SPECIFIC PORIN | | Descriptor: | CALCIUM ION, SUCROSE-SPECIFIC PORIN | | Authors: | Diederichs, K, Welte, W. | | Deposit date: | 1997-12-07 | | Release date: | 1998-06-10 | | Last modified: | 2024-02-07 | | Method: | X-RAY DIFFRACTION (2.4 Å) | | Cite: | Structure of the sucrose-specific porin ScrY from Salmonella typhimurium and its complex with sucrose.
Nat.Struct.Biol., 5, 1998
|
|
1A0T
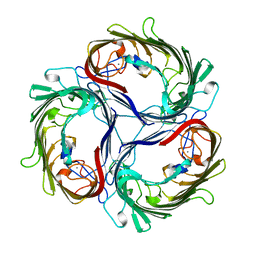 
 | | SUCROSE-SPECIFIC PORIN, WITH BOUND SUCROSE MOLECULES | | Descriptor: | CALCIUM ION, SUCROSE-SPECIFIC PORIN, beta-D-fructofuranose-(2-1)-alpha-D-glucopyranose | | Authors: | Diederichs, K, Welte, W. | | Deposit date: | 1997-12-08 | | Release date: | 1998-03-18 | | Last modified: | 2024-05-22 | | Method: | X-RAY DIFFRACTION (2.4 Å) | | Cite: | Structure of the sucrose-specific porin ScrY from Salmonella typhimurium and its complex with sucrose.
Nat.Struct.Biol., 5, 1998
|
|
5AG3
 
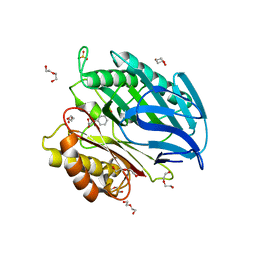 | | Chorismatase mechanisms reveal fundamentally different types of reaction in a single conserved protein fold | | Descriptor: | 3-(2-CARBOXYETHYL)BENZOIC ACID, DI(HYDROXYETHYL)ETHER, PUTATIVE PTERIDINE-DEPENDENT DIOXYGENASE, ... | | Authors: | Hubrich, F, Juneja, P, Mueller, M, Diederichs, K, Welte, W, Andexer, J.N. | | Deposit date: | 2015-01-28 | | Release date: | 2015-08-19 | | Last modified: | 2024-01-10 | | Method: | X-RAY DIFFRACTION (1.898 Å) | | Cite: | Chorismatase Mechanisms Reveal Fundamentally Different Types of Reaction in a Single Conserved Protein Fold.
J.Am.Chem.Soc., 137, 2015
|
|
1OH2
 
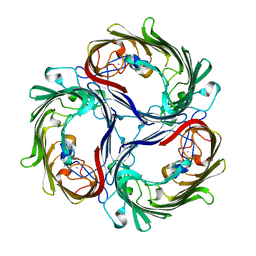 | |
1QXX
 
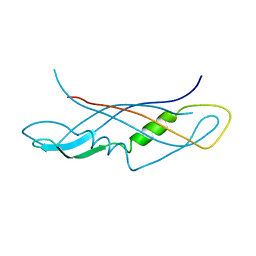 | | CRYSTAL STRUCTURE OF THE C-TERMINAL DOMAIN OF TONB | | Descriptor: | TonB protein | | Authors: | Koedding, J, Howard, P, Kaufmann, L, Polzer, P, Lustig, A, Welte, W. | | Deposit date: | 2003-09-09 | | Release date: | 2004-04-06 | | Last modified: | 2023-08-23 | | Method: | X-RAY DIFFRACTION (2.7 Å) | | Cite: | Dimerization of TonB is not essential for its binding to the outer membrane siderophore receptor FhuA of Escherichia coli.
J.Biol.Chem., 279, 2004
|
|
5A3K
 
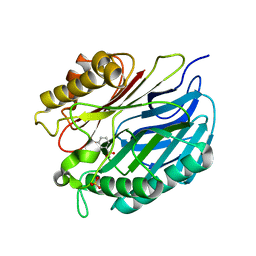 | | Chorismatase mechanisms reveal fundamentally different types of reaction in a single conserved protein fold | | Descriptor: | 3-HYDROXYBENZOIC ACID, PUTATIVE PTERIDINE-DEPENDENT DIOXYGENASE, SULFATE ION | | Authors: | Hubrich, F, Juneja, P, Mueller, M, Diederichs, K, Welte, W, Andexer, J.N. | | Deposit date: | 2015-06-01 | | Release date: | 2015-08-19 | | Last modified: | 2024-01-10 | | Method: | X-RAY DIFFRACTION (2.753 Å) | | Cite: | Chorismatase Mechanisms Reveal Fundamentally Different Types of Reaction in a Single Conserved Protein Fold.
J.Am.Chem.Soc., 137, 2015
|
|
1G29
 
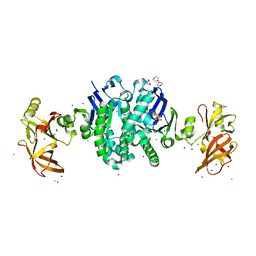 | | MALK | | Descriptor: | 1,4-DIETHYLENE DIOXIDE, AMMONIUM ION, CHLORIDE ION, ... | | Authors: | Diederichs, K, Diez, J, Greller, G, Mueller, C, Breed, J, Schnell, C, Vonrhein, C, Boos, W, Welte, W. | | Deposit date: | 2000-10-18 | | Release date: | 2000-12-06 | | Last modified: | 2024-02-07 | | Method: | X-RAY DIFFRACTION (1.9 Å) | | Cite: | Crystal structure of MalK, the ATPase subunit of the trehalose/maltose ABC transporter of the archaeon Thermococcus litoralis.
EMBO J., 19, 2000
|
|
1EU8
 
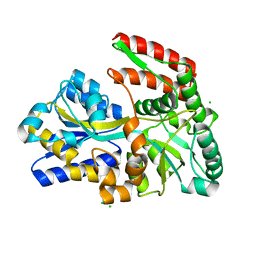 | | STRUCTURE OF TREHALOSE MALTOSE BINDING PROTEIN FROM THERMOCOCCUS LITORALIS | | Descriptor: | CHLORIDE ION, PLATINUM (II) ION, TREHALOSE/MALTOSE BINDING PROTEIN, ... | | Authors: | Diez, J, Diederichs, K, Greller, G, Horlacher, R, Boos, W, Welte, W. | | Deposit date: | 2000-04-14 | | Release date: | 2001-03-14 | | Last modified: | 2024-02-07 | | Method: | X-RAY DIFFRACTION (1.9 Å) | | Cite: | The crystal structure of a liganded trehalose/maltose-binding protein from the hyperthermophilic Archaeon Thermococcus litoralis at 1.85 A.
J.Mol.Biol., 305, 2001
|
|
1FCP
 
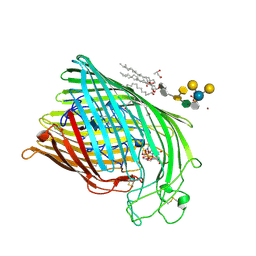 | | FERRIC HYDROXAMATE UPTAKE RECEPTOR (FHUA) FROM E.COLI IN COMPLEX WITH BOUND FERRICHROME-IRON | | Descriptor: | 2-TRIDECANOYLOXY-PENTADECANOIC ACID, 3-OXO-PENTADECANOIC ACID, ACETOACETIC ACID, ... | | Authors: | Hofmann, E, Ferguson, A.D, Diederichs, K, Welte, W. | | Deposit date: | 1998-10-14 | | Release date: | 1999-01-13 | | Last modified: | 2020-07-29 | | Method: | X-RAY DIFFRACTION (2.7 Å) | | Cite: | Siderophore-mediated iron transport: crystal structure of FhuA with bound lipopolysaccharide.
Science, 282, 1998
|
|
1FUE
 
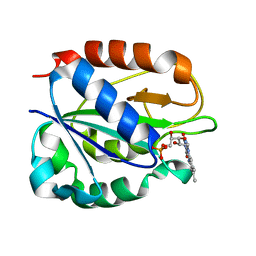 | | FLAVODOXIN FROM HELICOBACTER PYLORI | | Descriptor: | FLAVIN MONONUCLEOTIDE, FLAVODOXIN | | Authors: | Freigang, J, Diederichs, K, Schaefer, K.P, Welte, W, Paul, R. | | Deposit date: | 2000-09-15 | | Release date: | 2002-02-06 | | Last modified: | 2024-02-07 | | Method: | X-RAY DIFFRACTION (2.4 Å) | | Cite: | Crystal structure of oxidized flavodoxin, an essential protein in Helicobacter pylori.
Protein Sci., 11, 2002
|
|
1DUW
 
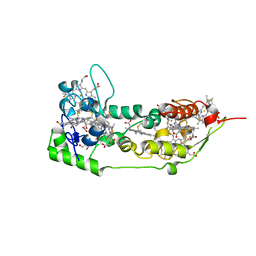 | | STRUCTURE OF NONAHEME CYTOCHROME C | | Descriptor: | GLYCEROL, HEME C, NONAHEME CYTOCHROME C | | Authors: | Umhau, S, Fritz, G, Diederichs, K, Breed, J, Kroneck, P.M, Welte, W. | | Deposit date: | 2000-01-19 | | Release date: | 2001-03-21 | | Last modified: | 2024-10-30 | | Method: | X-RAY DIFFRACTION (1.89 Å) | | Cite: | Three-dimensional structure of the nonaheme cytochrome c from Desulfovibrio desulfuricans Essex in the Fe(III) state at 1.89 A resolution.
Biochemistry, 40, 2001
|
|
3QPH
 
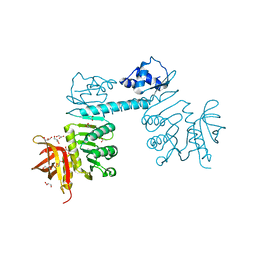 | | The three-dimensional structure of TrmB, a global transcriptional regulator of the hyperthermophilic archaeon Pyrococcus furiosus in complex with sucrose | | Descriptor: | ACETATE ION, GLYCEROL, TrmB, ... | | Authors: | Krug, M, Lee, S.-J, Boos, W, Welte, W, Diederichs, K. | | Deposit date: | 2011-02-13 | | Release date: | 2012-05-30 | | Last modified: | 2024-02-21 | | Method: | X-RAY DIFFRACTION (2.992 Å) | | Cite: | The three-dimensional structure of TrmB, a transcriptional regulator of dual function in the hyperthermophilic archaeon Pyrococcus furiosus in complex with sucrose.
Protein Sci., 22, 2013
|
|
1E54
 
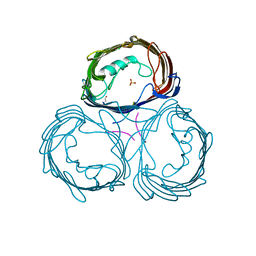 | | Anion-selective porin from Comamonas acidovorans | | Descriptor: | CALCIUM ION, OMP32, OUTER MEMBRANE PORIN PROTEIN 32, ... | | Authors: | Zeth, K, Diederichs, K, Welte, W, Engelhardt, H. | | Deposit date: | 2000-07-17 | | Release date: | 2001-07-12 | | Last modified: | 2024-10-16 | | Method: | X-RAY DIFFRACTION (2.1 Å) | | Cite: | Crystal Structure of Omp32, the Anion-Selective Porin from Comamonas Acidovorans, in Complex with a Periplasmic Peptideat 2.1 A Resolution
Structure, 8, 2000
|
|
1CS6
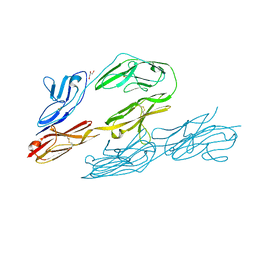 
 | | N-TERMINAL FRAGMENT OF AXONIN-1 FROM CHICKEN | | Descriptor: | AXONIN-1, GLYCEROL | | Authors: | Freigang, J, Proba, K, Diederichs, K, Sonderegger, P, Welte, W. | | Deposit date: | 1999-08-17 | | Release date: | 2000-05-19 | | Last modified: | 2024-10-30 | | Method: | X-RAY DIFFRACTION (1.8 Å) | | Cite: | The crystal structure of the ligand binding module of axonin-1/TAG-1 suggests a zipper mechanism for neural cell adhesion.
Cell(Cambridge,Mass.), 101, 2000
|
|
1U07
 
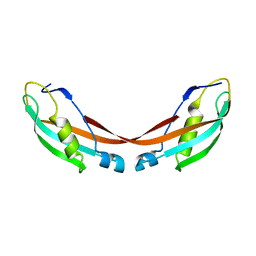 | | Crystal Structure of the 92-residue C-term. part of TonB with significant structural changes compared to shorter fragments | | Descriptor: | TonB protein | | Authors: | Koedding, J, Killig, F, Polzer, P, Howard, S.P, Diederichs, K, Welte, W. | | Deposit date: | 2004-07-13 | | Release date: | 2004-11-30 | | Last modified: | 2024-03-13 | | Method: | X-RAY DIFFRACTION (1.13 Å) | | Cite: | Crystal structure of a 92-residue c-terminal fragment of TonB from Escherichia coli reveals significant conformational changes compared to structures of smaller TonB fragments
J.Biol.Chem., 280, 2005
|
|
1B8D
 
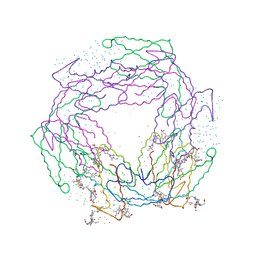 | | CRYSTAL STRUCTURE OF A PHYCOUROBILIN-CONTAINING PHYCOERYTHRIN | | Descriptor: | PHYCOERYTHROBILIN, PHYCOUROBILIN, PROTEIN (RHODOPHYTAN PHYCOERYTHRIN (ALPHA CHAIN)), ... | | Authors: | Ritter, S, Hiller, R.G, Wrench, P.M, Welte, W, Diederichs, K. | | Deposit date: | 1999-01-29 | | Release date: | 1999-02-18 | | Last modified: | 2023-08-09 | | Method: | X-RAY DIFFRACTION (1.9 Å) | | Cite: | Crystal structure of a phycourobilin-containing phycoerythrin at 1.90-A resolution.
J.Struct.Biol., 126, 1999
|
|
1C0L
 
 | | D-AMINO ACID OXIDASE: STRUCTURE OF SUBSTRATE COMPLEXES AT VERY HIGH RESOLUTION REVEAL THE CHEMICAL REACTTION MECHANISM OF FLAVIN DEHYDROGENATION | | Descriptor: | D-AMINO ACID OXIDASE, FLAVIN-ADENINE DINUCLEOTIDE, TRIFLUOROALANINE | | Authors: | Umhau, S, Molla, G, Diederichs, K, Pilone, M.S, Ghisla, S, Welte, W. | | Deposit date: | 1999-07-16 | | Release date: | 2000-11-22 | | Last modified: | 2024-02-07 | | Method: | X-RAY DIFFRACTION (1.73 Å) | | Cite: | The x-ray structure of D-amino acid oxidase at very high resolution identifies the chemical mechanism of flavin-dependent substrate dehydrogenation.
Proc.Natl.Acad.Sci.USA, 97, 2000
|
|
1C0I
 
 | | CRYSTAL STRUCTURE OF D-AMINO ACID OXIDASE IN COMPLEX WITH TWO ANTHRANYLATE MOLECULES | | Descriptor: | 2-AMINOBENZOIC ACID, D-AMINO ACID OXIDASE, FLAVIN-ADENINE DINUCLEOTIDE | | Authors: | Pollegioni, L, Diederichs, K, Molla, G, Umhau, S, Welte, W, Ghisla, S, Pilone, M.S. | | Deposit date: | 1999-07-16 | | Release date: | 2002-02-27 | | Last modified: | 2023-11-15 | | Method: | X-RAY DIFFRACTION (1.9 Å) | | Cite: | Yeast d-amino Acid oxidase: structural basis of its catalytic properties
J.Mol.Biol., 324, 2002
|
|
1C0K
 
 | | CRYSTAL STRUCTURE ANALYSIS OF D-AMINO ACID OXIDASE IN COMPLEX WITH L-LACTATE | | Descriptor: | FLAVIN-ADENINE DINUCLEOTIDE, LACTIC ACID, PROTEIN (D-AMINO ACID OXIDASE) | | Authors: | Umhau, S, Molla, G, Diederichs, K, Pilone, M.S, Ghisla, S, Welte, W, Pollegioni, L. | | Deposit date: | 1999-07-16 | | Release date: | 2000-11-22 | | Last modified: | 2023-11-15 | | Method: | X-RAY DIFFRACTION (1.46 Å) | | Cite: | The x-ray structure of D-amino acid oxidase at very high resolution identifies the chemical mechanism of flavin-dependent substrate dehydrogenation.
Proc.Natl.Acad.Sci.USA, 97, 2000
|
|
1C0P
 
 | | D-AMINO ACIC OXIDASE IN COMPLEX WITH D-ALANINE AND A PARTIALLY OCCUPIED BIATOMIC SPECIES | | Descriptor: | D-ALANINE, D-AMINO ACID OXIDASE, FLAVIN-ADENINE DINUCLEOTIDE, ... | | Authors: | Umhau, S, Pollegioni, L, Molla, G, Diederichs, K, Welte, W, Pilone, S.M, Ghisla, S. | | Deposit date: | 1999-07-19 | | Release date: | 2000-11-22 | | Last modified: | 2024-02-07 | | Method: | X-RAY DIFFRACTION (1.2 Å) | | Cite: | The x-ray structure of D-amino acid oxidase at very high resolution identifies the chemical mechanism of flavin-dependent substrate dehydrogenation.
Proc.Natl.Acad.Sci.USA, 97, 2000
|
|
3PY8
 
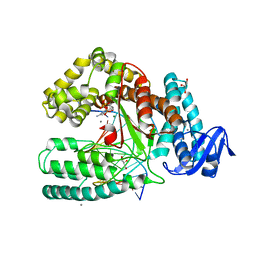 | | Crystal structure of a mutant of the large fragment of DNA polymerase I from thermus aquaticus in a closed ternary complex with DNA and ddCTP | | Descriptor: | 2',3'-DIDEOXYCYTIDINE 5'-TRIPHOSPHATE, DNA (5'-D(*AP*A*AP*GP*GP*GP*CP*GP*CP*CP*GP*TP*GP*GP*TP*C)-3'), DNA (5'-D(*GP*AP*CP*CP*AP*CP*GP*GP*CP*GP*CP*(DOC))-3'), ... | | Authors: | Schnur, A, Marx, A, Welte, W, Diederichs, K. | | Deposit date: | 2010-12-12 | | Release date: | 2011-06-15 | | Last modified: | 2024-02-21 | | Method: | X-RAY DIFFRACTION (1.74 Å) | | Cite: | Learning from Directed Evolution: Thermus aquaticus DNA Polymerase Mutants with Translesion Synthesis Activity.
Chembiochem, 12, 2011
|
|
1RYI
 
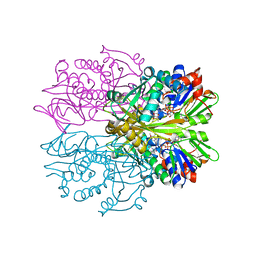 | | STRUCTURE OF GLYCINE OXIDASE WITH BOUND INHIBITOR GLYCOLATE | | Descriptor: | FLAVIN-ADENINE DINUCLEOTIDE, GLYCINE OXIDASE, GLYCOLIC ACID | | Authors: | Moertl, M, Diederichs, K, Welte, W, Pollegioni, L, Molla, G, Motteran, L, Andriolo, G, Pilone, M.S. | | Deposit date: | 2003-12-22 | | Release date: | 2005-02-22 | | Last modified: | 2023-11-15 | | Method: | X-RAY DIFFRACTION (1.8 Å) | | Cite: | Structure-function correlation in glycine oxidase from Bacillus subtilis
J.Biol.Chem., 279, 2004
|
|
