8IHO
 
 | | Crystal structures of SARS-CoV-2 papain-like protease in complex with covalent inhibitors | | Descriptor: | Papain-like protease nsp3, ZINC ION, covalent inhibitor | | Authors: | Wang, Q, Hu, H, Li, M, Xu, Y. | | Deposit date: | 2023-02-23 | | Release date: | 2024-01-03 | | Method: | X-RAY DIFFRACTION (2.55 Å) | | Cite: | Structure-Based Design of Potent Peptidomimetic Inhibitors Covalently Targeting SARS-CoV-2 Papain-like Protease.
Int J Mol Sci, 24, 2023
|
|
6K87
 
 | |
6K82
 
 | | RGLG1 mutant-D338A E378A | | Descriptor: | E3 ubiquitin-protein ligase RGLG1, MAGNESIUM ION, SODIUM ION | | Authors: | Wang, Q, Wu, Y. | | Deposit date: | 2019-06-11 | | Release date: | 2020-06-24 | | Last modified: | 2024-03-27 | | Method: | X-RAY DIFFRACTION (1.402 Å) | | Cite: | RGLG1 mutant-D338A E378A
To Be Published
|
|
6K85
 
 | | The closed state of RGLG1 mutant-D338A | | Descriptor: | E3 ubiquitin-protein ligase RGLG1, MAGNESIUM ION | | Authors: | Wang, Q, Wu, Y. | | Deposit date: | 2019-06-11 | | Release date: | 2020-06-24 | | Last modified: | 2024-03-27 | | Method: | X-RAY DIFFRACTION (1.606 Å) | | Cite: | The closed state of RGLG1 mutant-D338A
To Be Published
|
|
6K8A
 
 | |
6K88
 
 | | RGLG1 MIDAS binds calcium ion | | Descriptor: | CALCIUM ION, E3 ubiquitin-protein ligase RGLG1 | | Authors: | Wang, Q, Wu, Y. | | Deposit date: | 2019-06-11 | | Release date: | 2020-06-24 | | Last modified: | 2024-03-27 | | Method: | X-RAY DIFFRACTION (1.788 Å) | | Cite: | RGLG1 MIDAS binds calcium ion
To Be Published
|
|
7VXN
 
 | |
5ZY5
 
 | | spCOMT apo structure | | Descriptor: | Probable catechol O-methyltransferase 1 | | Authors: | Wang, Q, Xu, L. | | Deposit date: | 2018-05-22 | | Release date: | 2019-05-22 | | Last modified: | 2023-11-22 | | Method: | X-RAY DIFFRACTION (2.295 Å) | | Cite: | structural and functional investigations of spCOMT
To Be Published
|
|
7VXL
 
 | |
6K86
 
 | | The closed state of RGLG1 mutant-E378A | | Descriptor: | E3 ubiquitin-protein ligase RGLG1, MAGNESIUM ION, SODIUM ION | | Authors: | Wang, Q, Wu, Y. | | Deposit date: | 2019-06-11 | | Release date: | 2020-06-24 | | Last modified: | 2024-03-27 | | Method: | X-RAY DIFFRACTION (1.593 Å) | | Cite: | The closed state of RGLG1 mutant-E378A
To Be Published
|
|
6K8B
 
 | | The open state of RGLG1 VWA domain | | Descriptor: | CALCIUM ION, E3 ubiquitin-protein ligase RGLG1, MAGNESIUM ION | | Authors: | Wang, Q, Wu, Y. | | Deposit date: | 2019-06-11 | | Release date: | 2020-06-24 | | Last modified: | 2024-03-27 | | Method: | X-RAY DIFFRACTION (2.21 Å) | | Cite: | The open state of RGLG1 VWA domain
To Be Published
|
|
5YWJ
 
 | | Global regulatory element SarX | | Descriptor: | HTH-type transcriptional regulator SarX | | Authors: | Wang, Q. | | Deposit date: | 2017-11-29 | | Release date: | 2018-12-05 | | Last modified: | 2023-11-22 | | Method: | X-RAY DIFFRACTION (2.102 Å) | | Cite: | Crystal structure of SarX from Staphylococcus aureus
To Be Published
|
|
5ZY6
 
 | | catechol methyltransferase spCOMT | | Descriptor: | Probable catechol O-methyltransferase 1, S-ADENOSYLMETHIONINE | | Authors: | Wang, Q, Xu, L. | | Deposit date: | 2018-05-22 | | Release date: | 2019-05-22 | | Last modified: | 2023-11-22 | | Method: | X-RAY DIFFRACTION (2.099 Å) | | Cite: | structural insight into spCOMT
To Be Published
|
|
6K89
 
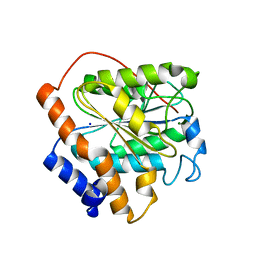 | | The closed state of RGLG1 VWA domain | | Descriptor: | E3 ubiquitin-protein ligase RGLG1, MAGNESIUM ION, SODIUM ION | | Authors: | Wang, Q, Wu, Y. | | Deposit date: | 2019-06-11 | | Release date: | 2020-06-24 | | Last modified: | 2024-03-27 | | Method: | X-RAY DIFFRACTION (1.689 Å) | | Cite: | The closed state of RGLG1 VWA domain
To Be Published
|
|
6K8E
 
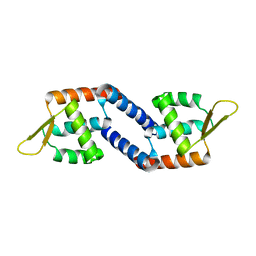 | | Global regulatory element SarX | | Descriptor: | HTH-type transcriptional regulator SarX | | Authors: | Wang, Q. | | Deposit date: | 2019-06-11 | | Release date: | 2020-06-17 | | Last modified: | 2023-11-22 | | Method: | X-RAY DIFFRACTION (2.002 Å) | | Cite: | Crystal structure of SarX from Staphylococcus aureus
To Be Published
|
|
7C2K
 
 | | COVID-19 RNA-dependent RNA polymerase pre-translocated catalytic complex | | Descriptor: | Non-structural protein 7, Non-structural protein 8, RNA (29-MER), ... | | Authors: | Wang, Q, Gao, Y, Ji, W, Mu, A, Rao, Z. | | Deposit date: | 2020-05-07 | | Release date: | 2020-06-03 | | Last modified: | 2024-03-27 | | Method: | ELECTRON MICROSCOPY (2.93 Å) | | Cite: | Structural Basis for RNA Replication by the SARS-CoV-2 Polymerase.
Cell, 182, 2020
|
|
7DTE
 
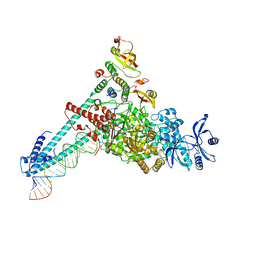 | | SARS-CoV-2 RdRP catalytic complex with T33-1 RNA | | Descriptor: | Non-structural protein 7, Non-structural protein 8, RNA (33-MER), ... | | Authors: | Wang, Q, Gong, P. | | Deposit date: | 2021-01-04 | | Release date: | 2021-10-20 | | Last modified: | 2024-06-05 | | Method: | ELECTRON MICROSCOPY (3 Å) | | Cite: | Remdesivir overcomes the S861 roadblock in SARS-CoV-2 polymerase elongation complex.
Cell Rep, 37, 2021
|
|
7DHX
 
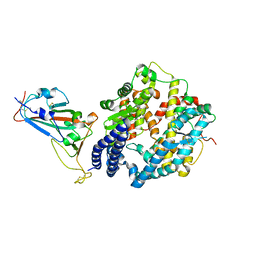 | | Crystal structure of SARS-CoV-2 RBD binding to pangolin ACE2 | | Descriptor: | 2-acetamido-2-deoxy-beta-D-glucopyranose, Spike glycoprotein, ZINC ION, ... | | Authors: | Wang, Q.H, Qi, J.X, Wu, L.L. | | Deposit date: | 2020-11-17 | | Release date: | 2021-09-29 | | Last modified: | 2023-11-29 | | Method: | X-RAY DIFFRACTION (2.3 Å) | | Cite: | Molecular basis of pangolin ACE2 engaged by COVID-19 virus
Chin.Sci.Bull., 66, 2021
|
|
7E4I
 
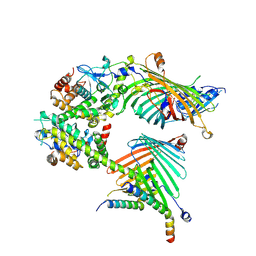 | | Cryo-EM structure of the yeast mitochondrial SAM-Tom40/Tom5/Tom6 complex at 3.0 angstrom | | Descriptor: | Mitochondrial import receptor subunit TOM40, Mitochondrial import receptor subunit TOM5, Mitochondrial import receptor subunit TOM6, ... | | Authors: | Wang, Q, Guan, Z.Y, Qi, L.B, Yan, C.Y, Yin, P. | | Deposit date: | 2021-02-13 | | Release date: | 2021-09-01 | | Last modified: | 2024-06-05 | | Method: | ELECTRON MICROSCOPY (3.05 Å) | | Cite: | Structural insight into the SAM-mediated assembly of the mitochondrial TOM core complex.
Science, 373, 2021
|
|
7E4H
 
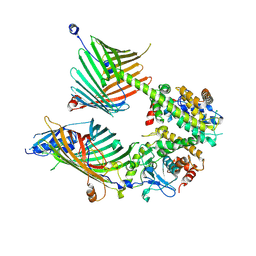 | | Cryo-EM structure of the yeast mitochondrial SAM-Tom40 complex at 3.0 angstrom | | Descriptor: | Mitochondrial import receptor subunit TOM40, Sorting assembly machinery 35 kDa subunit, Sorting assembly machinery 37 kDa subunit, ... | | Authors: | Wang, Q, Guan, Z.Y, Qi, L.B, Yan, C.Y, Yin, P. | | Deposit date: | 2021-02-13 | | Release date: | 2021-09-01 | | Last modified: | 2024-06-05 | | Method: | ELECTRON MICROSCOPY (3.01 Å) | | Cite: | Structural insight into the SAM-mediated assembly of the mitochondrial TOM core complex.
Science, 373, 2021
|
|
5DO2
 
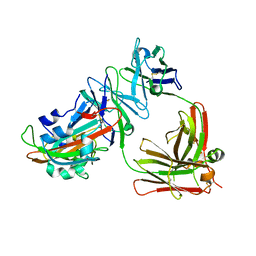 | | Complex structure of MERS-RBD bound with 4C2 antibody | | Descriptor: | 2-acetamido-2-deoxy-beta-D-glucopyranose, 4C2 heavy chain, 4C2 light chain, ... | | Authors: | Li, Y, Wan, Y, Liu, P, Zhao, J, Lu, G, Qi, J, Wang, Q, Lu, X, Wu, Y, Liu, W, Yuen, K.Y, Perlman, S, Gao, G.F, Yan, J. | | Deposit date: | 2015-09-10 | | Release date: | 2015-10-14 | | Last modified: | 2023-11-08 | | Method: | X-RAY DIFFRACTION (2.409 Å) | | Cite: | A humanized neutralizing antibody against MERS-CoV targeting the receptor-binding domain of the spike protein.
Cell Res., 25, 2015
|
|
6KWC
 
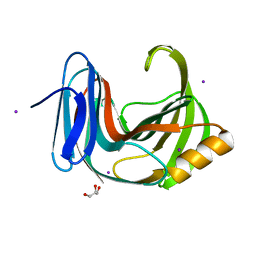 | | Crystal Structure Analysis of Endo-beta-1,4-xylanase II | | Descriptor: | Endo-1,4-beta-xylanase 2, GLYCEROL, IODIDE ION | | Authors: | Li, C, Wan, Q. | | Deposit date: | 2019-09-06 | | Release date: | 2021-01-27 | | Last modified: | 2023-11-22 | | Method: | X-RAY DIFFRACTION (1.3 Å) | | Cite: | Studying the Role of a Single Mutation of a Family 11 Glycoside Hydrolase Using High-Resolution X-ray Crystallography.
Protein J., 39, 2020
|
|
6KVV
 
 | |
6KWE
 
 | |
6KWH
 
 | | Crystal Structure Analysis of Endo-beta-1,4-xylanase II Complexed with Xylotriose | | Descriptor: | Endo-1,4-beta-xylanase 2, IODIDE ION, beta-D-xylopyranose-(1-4)-beta-D-xylopyranose-(1-4)-beta-D-xylopyranose | | Authors: | Li, C, Wan, Q. | | Deposit date: | 2019-09-06 | | Release date: | 2021-07-14 | | Last modified: | 2023-11-22 | | Method: | X-RAY DIFFRACTION (1.808 Å) | | Cite: | X-ray crystallographic studies of family 11 xylanase Michaelis and product complexes: implications for the catalytic mechanism
To Be Published
|
|
