5XWY
 
 | | Electron cryo-microscopy structure of LbuCas13a-crRNA binary complex | | Descriptor: | A type VI-A CRISPR-Cas RNA-guided RNA ribonuclease, Cas13a, RNA (59-MER) | | Authors: | Zhang, X, Wang, Y, Ma, J, Liu, L, Li, X, Li, Z, You, L, Wang, J, Wang, M. | | Deposit date: | 2017-06-30 | | Release date: | 2017-09-13 | | Last modified: | 2024-03-27 | | Method: | ELECTRON MICROSCOPY (3.2 Å) | | Cite: | The Molecular Architecture for RNA-Guided RNA Cleavage by Cas13a.
Cell, 170, 2017
|
|
6P8D
 
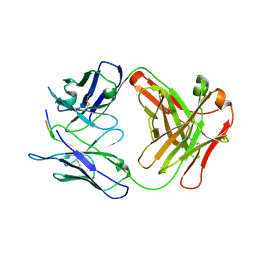 | |
6PEC
 
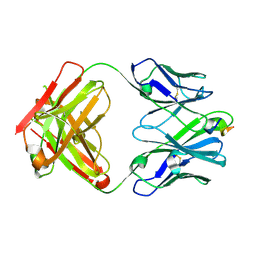 | |
6PEF
 
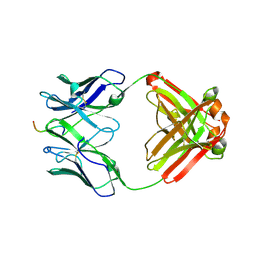 | |
6DXE
 
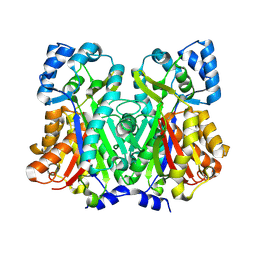 | |
6DX8
 
 | |
6PDS
 
 | | Vaccine-elicited NHP FP-targeting antibody 0PV-a.04 in complex with HIV fusion peptide (residue 512-519) | | Descriptor: | HIV-1 fusion peptide residue 512-519, SULFATE ION, antibody 0PV-a.04 heavy chain, ... | | Authors: | Xu, K, Liu, K, Wang, Y, Kwong, P.D. | | Deposit date: | 2019-06-19 | | Release date: | 2020-06-24 | | Method: | X-RAY DIFFRACTION (1.893 Å) | | Cite: | Modular recognition of antigens provides a mechanism that improves vaccine-elicited antibody-class frequencies
To Be Published
|
|
6DXA
 
 | |
8TOO
 
 | | Crystal structure of Epstein-Barr virus gp42 in complex with antibody 4C12 | | Descriptor: | 4C12 heavy chain, 4C12 light chain, Glycoprotein 42 | | Authors: | Bu, W, Kumar, A, Board, N, Kim, J, Dowdell, K, Zhang, S, Lei, Y, Hostal, A, Krogmann, T, Wang, Y, Pittaluga, S, Marcotrigiano, J, Cohen, J.I. | | Deposit date: | 2023-08-03 | | Release date: | 2024-03-27 | | Method: | X-RAY DIFFRACTION (2.6 Å) | | Cite: | Epstein-Barr virus gp42 antibodies reveal sites of vulnerability for receptor binding and fusion to B cells.
Immunity, 57, 2024
|
|
8TNN
 
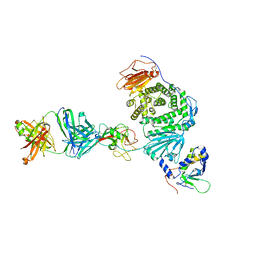 | | Crystal structure of Epstein-Barr virus gH/gL/gp42 in complex with gp42 antibody A10 | | Descriptor: | 2-acetamido-2-deoxy-beta-D-glucopyranose, A10 heavy chain, A10 light chain, ... | | Authors: | Bu, W, Kumar, A, Board, N, Kim, J, Dowdell, K, Zhang, S, Lei, Y, Hostal, A, Krogmann, T, Wang, Y, Pittaluga, S, Marcotrigiano, J, Cohen, J.I. | | Deposit date: | 2023-08-02 | | Release date: | 2024-03-27 | | Last modified: | 2024-05-22 | | Method: | X-RAY DIFFRACTION (3.36 Å) | | Cite: | Epstein-Barr virus gp42 antibodies reveal sites of vulnerability for receptor binding and fusion to B cells.
Immunity, 57, 2024
|
|
8TNT
 
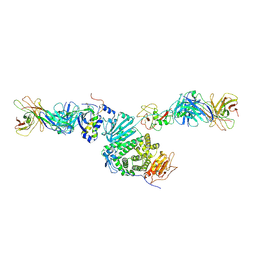 | | Crystal structure of Epstein-Barr virus gH/gL/gp42 in complex with antibodies F-2-1 and 769C2 | | Descriptor: | 2-acetamido-2-deoxy-beta-D-glucopyranose, 769C2 heavy chain, 769C2 light chain, ... | | Authors: | Bu, W, Kumar, A, Board, N, Kim, J, Dowdell, K, Zhang, S, Lei, Y, Hostal, A, Krogmann, T, Wang, Y, Pittaluga, S, Marcotrigiano, J, Cohen, J.I. | | Deposit date: | 2023-08-02 | | Release date: | 2024-03-27 | | Method: | X-RAY DIFFRACTION (3.15 Å) | | Cite: | Epstein-Barr virus gp42 antibodies reveal sites of vulnerability for receptor binding and fusion to B cells.
Immunity, 57, 2024
|
|
5YEL
 
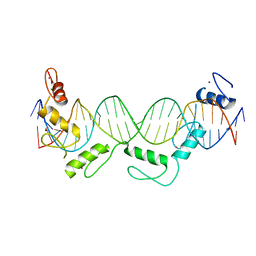 | | Crystal structure of CTCF ZFs6-11-gb7CSE | | Descriptor: | DNA (26-MER), Transcriptional repressor CTCF, ZINC ION | | Authors: | Yin, M, Wang, J, Wang, M, Li, X, Wang, Y. | | Deposit date: | 2017-09-18 | | Release date: | 2017-11-29 | | Last modified: | 2024-03-27 | | Method: | X-RAY DIFFRACTION (2.96 Å) | | Cite: | Molecular mechanism of directional CTCF recognition of a diverse range of genomic sites
Cell Res., 27, 2017
|
|
1ZOI
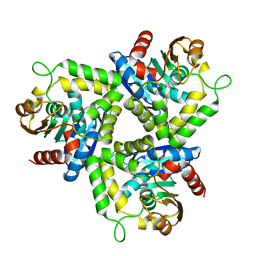 
 | | Crystal Structure of a Stereoselective Esterase from Pseudomonas putida IFO12996 | | Descriptor: | esterase | | Authors: | Elmi, F, Lee, H.T, Huang, J.Y, Hsieh, Y.C, Wang, Y.L, Chen, Y.J, Shaw, S.Y, Chen, C.J. | | Deposit date: | 2005-05-13 | | Release date: | 2006-05-02 | | Last modified: | 2024-03-13 | | Method: | X-RAY DIFFRACTION (1.6 Å) | | Cite: | Stereoselective esterase from Pseudomonas putida IFO12996 reveals alpha/beta hydrolase folds for D-beta-acetylthioisobutyric acid synthesis
J.Bacteriol., 187, 2005
|
|
8YM2
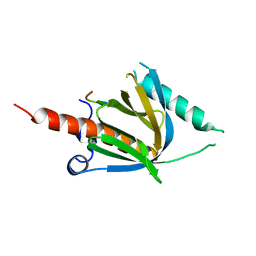 
 | | Crystal structure of AIDA-1 PTB domain in complex with SynGAP NPxF motif | | Descriptor: | Ankyrin repeat and sterile alpha motif domain-containing protein 1B, Ras/Rap GTPase-activating protein SynGAP | | Authors: | Wang, X, Wang, Y, Cai, Q, Zhang, M. | | Deposit date: | 2024-03-08 | | Release date: | 2024-05-29 | | Last modified: | 2024-06-05 | | Method: | X-RAY DIFFRACTION (2 Å) | | Cite: | AIDA-1/ANKS1B Binds to the SynGAP Family RasGAPs with High Affinity and Specificity.
J.Mol.Biol., 436, 2024
|
|
4W9P
 
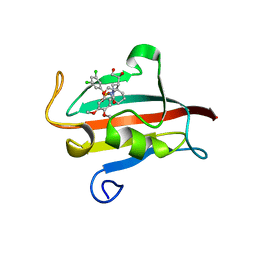 | | The Fk1 domain of FKBP51 in complex with (1S,5S,6R)-10-[(3,5-dichlorophenyl)sulfonyl]-5-[(1S)-1,2-dihydroxyethyl]-3-[2-(3,4-dimethoxyphenoxy)ethyl]-3,10-diazabicyclo[4.3.1]decan-2-one | | Descriptor: | (1S,5S,6R)-10-[(3,5-dichlorophenyl)sulfonyl]-5-[(1S)-1,2-dihydroxyethyl]-3-[2-(3,4-dimethoxyphenoxy)ethyl]-3,10-diazabicyclo[4.3.1]decan-2-one, ACETATE ION, Peptidyl-prolyl cis-trans isomerase FKBP5 | | Authors: | Pomplun, S, Wang, Y, Kirschner, K, Kozany, C, Bracher, A, Hausch, F. | | Deposit date: | 2014-08-27 | | Release date: | 2014-12-03 | | Last modified: | 2024-01-10 | | Method: | X-RAY DIFFRACTION (1.5 Å) | | Cite: | Rational Design and Asymmetric Synthesis of Potent and Neurotrophic Ligands for FK506-Binding Proteins (FKBPs).
Angew.Chem.Int.Ed.Engl., 54, 2015
|
|
6CJ7
 
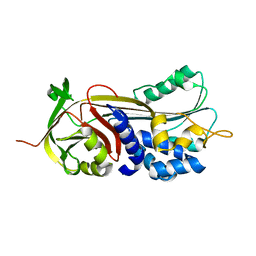 | | Crystal structure of Manduca sexta Serine protease inhibitor (Serpin)-12 | | Descriptor: | Serpin-12 | | Authors: | Gulati, M, Hu, Y, Peng, S, Pathak, P.K, Wang, Y, Deng, J, Jiang, H. | | Deposit date: | 2018-02-26 | | Release date: | 2018-07-04 | | Last modified: | 2023-10-04 | | Method: | X-RAY DIFFRACTION (1.6 Å) | | Cite: | Manduca sexta serpin-12 controls the prophenoloxidase activation system in larval hemolymph.
Insect Biochem. Mol. Biol., 99, 2018
|
|
4W9Q
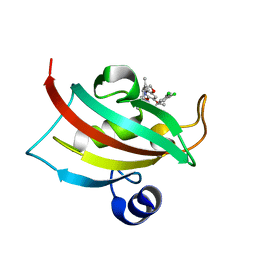 
 | | The Fk1 domain of FKBP51 in complex with (1S,5S,6R)-10-[(3,5-dichlorophenyl)sulfonyl]-3-[2-(3,4-dimethoxyphenoxy)ethyl]-5-ethyl-3,10-diazabicyclo[4.3.1]decan-2-one | | Descriptor: | (1S,5S,6R)-10-[(3,5-dichlorophenyl)sulfonyl]-3-[2-(3,4-dimethoxyphenoxy)ethyl]-5-ethyl-3,10-diazabicyclo[4.3.1]decan-2-one, Peptidyl-prolyl cis-trans isomerase FKBP5 | | Authors: | Pomplun, S, Wang, Y, Kirschner, K, Kozany, C, Bracher, A, Hausch, F. | | Deposit date: | 2014-08-27 | | Release date: | 2014-12-03 | | Last modified: | 2024-01-10 | | Method: | X-RAY DIFFRACTION (1.08 Å) | | Cite: | Rational Design and Asymmetric Synthesis of Potent and Neurotrophic Ligands for FK506-Binding Proteins (FKBPs).
Angew.Chem.Int.Ed.Engl., 54, 2015
|
|
8SNL
 
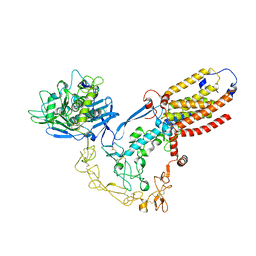 | | Structure of human ADAM17/iRhom2 sheddase complex | | Descriptor: | CALCIUM ION, Disintegrin and metalloproteinase domain-containing protein 17, Inactive rhomboid protein 2, ... | | Authors: | Zhao, H, Dai, Y, Wang, Y, Lee, C.H. | | Deposit date: | 2023-04-27 | | Release date: | 2024-05-29 | | Method: | ELECTRON MICROSCOPY (2.78 Å) | | Cite: | Structure of human ADAM17/iRhom2 sheddase complex
To Be Published
|
|
8SNO
 
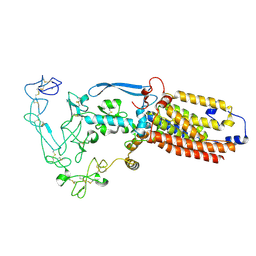 | | Structure of mature human ADAM17/iRhom2 sheddase complex, conformation 2 | | Descriptor: | CALCIUM ION, Disintegrin and metalloproteinase domain-containing protein 17, Inactive rhomboid protein 2 | | Authors: | Zhao, H, Dai, Y, Wang, Y, Lee, C.H. | | Deposit date: | 2023-04-27 | | Release date: | 2024-05-29 | | Method: | ELECTRON MICROSCOPY (2.78 Å) | | Cite: | Structure of mature human ADAM17/iRhom2 sheddase complex, conformation 2
To Be Published
|
|
8SNN
 
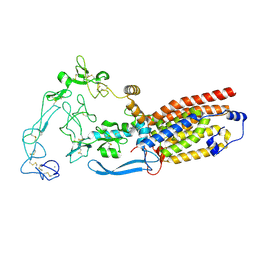 | | Structure of mature human ADAM17/iRhom2 sheddase complex, conformation 1 | | Descriptor: | CALCIUM ION, Disintegrin and metalloproteinase domain-containing protein 17, Inactive rhomboid protein 2 | | Authors: | Zhao, H, Dai, Y, Wang, Y, Lee, C.H. | | Deposit date: | 2023-04-27 | | Release date: | 2024-05-29 | | Method: | ELECTRON MICROSCOPY (2.32 Å) | | Cite: | Structure of mature human ADAM17/iRhom2 sheddase complex, conformation 1
To Be Published
|
|
8SNM
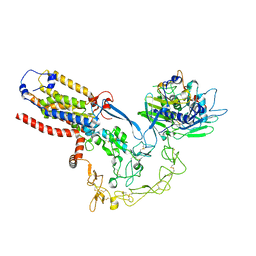 
 | | Structure of mature human ADAM17/iRhom2 sheddase complex in complex with ADAM17 prodomain | | Descriptor: | CALCIUM ION, Disintegrin and metalloproteinase domain-containing protein 17, Disintegrin and metalloproteinase domain-containing protein 17 propeptide, ... | | Authors: | Zhao, H, Dai, Y, Wang, Y, Lee, C.H. | | Deposit date: | 2023-04-27 | | Release date: | 2024-05-29 | | Method: | ELECTRON MICROSCOPY (3.84 Å) | | Cite: | Structure of mature human ADAM17/iRhom2 sheddase complex in complex with ADAM17 prodomain
To Be Published
|
|
5XT4
 
 | | Crystal Structure of Transketolase in complex with TPP intermediate VIII' from Pichia Stipitis | | Descriptor: | 2-C-{3-[(4-amino-2-methylpyrimidin-5-yl)methyl]-5-(2-{[(R)-hydroxy(phosphonooxy)phosphoryl]oxy}ethyl)-4-methyl-1,3-thiazol-3-ium-2-yl}-6-O-phosphono-D-glucitol, CALCIUM ION, DI(HYDROXYETHYL)ETHER, ... | | Authors: | Li, T.L, Hsu, N.S, Wang, Y.L. | | Deposit date: | 2017-06-16 | | Release date: | 2018-06-20 | | Last modified: | 2023-11-22 | | Method: | X-RAY DIFFRACTION (1.06 Å) | | Cite: | Evidence of Diradicals Involved in the Yeast Transketolase Catalyzed Keto-Transferring Reactions.
Chembiochem, 19, 2018
|
|
5XU2
 
 | | Crystal Structure of Transketolase in complex with TPP_III and fructose-6-phosphate from Pichia Stipitis | | Descriptor: | 2-[3-[(4-azanyl-2-methyl-pyrimidin-5-yl)methyl]-4-methyl-2H-1,3-thiazol-5-yl]ethyl phosphono hydrogen phosphate, CALCIUM ION, DI(HYDROXYETHYL)ETHER, ... | | Authors: | Li, T.L, Hsu, N.S, Wang, Y.L. | | Deposit date: | 2017-06-22 | | Release date: | 2018-06-27 | | Last modified: | 2023-11-22 | | Method: | X-RAY DIFFRACTION (0.97 Å) | | Cite: | Evidence of Diradicals Involved in the Yeast Transketolase Catalyzed Keto-Transferring Reactions.
Chembiochem, 19, 2018
|
|
5XT0
 
 | | Crystal Structure of Transketolase in complex with TPP intermediate VIII from Pichia Stipitis | | Descriptor: | 2-C-{3-[(4-amino-2-methylpyrimidin-5-yl)methyl]-5-(2-{[(R)-hydroxy(phosphonooxy)phosphoryl]oxy}ethyl)-4-methyl-1,3-thiazol-3-ium-2-yl}-6-O-phosphono-D-glucitol, CALCIUM ION, DI(HYDROXYETHYL)ETHER, ... | | Authors: | Li, T.L, Hsu, N.S, Wang, Y.L. | | Deposit date: | 2017-06-16 | | Release date: | 2018-06-20 | | Last modified: | 2023-11-22 | | Method: | X-RAY DIFFRACTION (1.15 Å) | | Cite: | Evidence of Diradicals Involved in the Yeast Transketolase Catalyzed Keto-Transferring Reactions.
Chembiochem, 19, 2018
|
|
5XPS
 
 | | Crystal Structure of Transketolase in complex with erythrose-4-phosphate from Pichia Stipitis | | Descriptor: | CALCIUM ION, DI(HYDROXYETHYL)ETHER, ERYTHOSE-4-PHOSPHATE, ... | | Authors: | Li, T.L, Hsu, N.S, Wang, Y.L. | | Deposit date: | 2017-06-04 | | Release date: | 2018-06-06 | | Last modified: | 2023-11-22 | | Method: | X-RAY DIFFRACTION (1.07 Å) | | Cite: | Evidence of Diradicals Involved in the Yeast Transketolase Catalyzed Keto-Transferring Reactions.
Chembiochem, 19, 2018
|
|
