8VTN
 
 | |
8VTK
 
 | |
8VTL
 
 | |
8VTO
 
 | |
8VTM
 
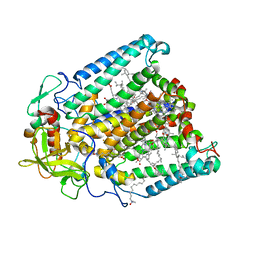 | |
1H0O
 
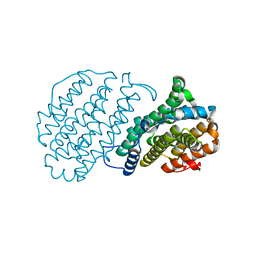 | |
1H0N
 
 | |
1FOV
 
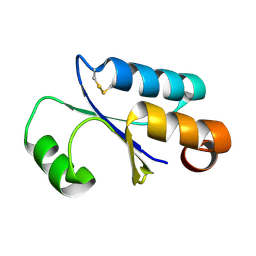 | | GLUTAREDOXIN 3 FROM ESCHERICHIA COLI IN THE FULLY OXIDIZED FORM | | Descriptor: | GLUTAREDOXIN 3 | | Authors: | Nordstrand, K, Sandstrom, A, Aslund, F, Holmgren, A, Otting, G, Berndt, K.D. | | Deposit date: | 2000-08-29 | | Release date: | 2000-10-26 | | Last modified: | 2022-02-23 | | Method: | SOLUTION NMR | | Cite: | NMR structure of oxidized glutaredoxin 3 from Escherichia coli.
J.Mol.Biol., 303, 2000
|
|
1X9F
 
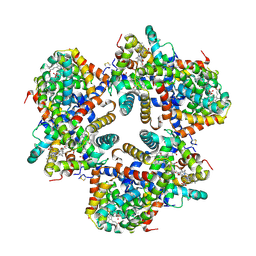 | | Hemoglobin Dodecamer from Lumbricus Erythrocruorin | | Descriptor: | CARBON MONOXIDE, Globin II, extracellular, ... | | Authors: | Strand, K, Knapp, J.E, Bhyravbhatla, B, Royer Jr, W.E. | | Deposit date: | 2004-08-20 | | Release date: | 2004-11-30 | | Last modified: | 2024-04-03 | | Method: | X-RAY DIFFRACTION (2.6 Å) | | Cite: | Crystal structure of the hemoglobin dodecamer from lumbricus erythrocruorin: allosteric core of giant annelid respiratory complexes
J.Mol.Biol., 344, 2004
|
|
3GRX
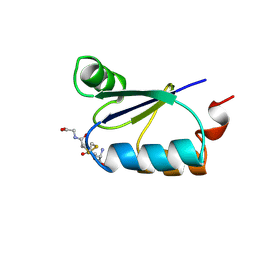 
 | | NMR STRUCTURE OF ESCHERICHIA COLI GLUTAREDOXIN 3-GLUTATHIONE MIXED DISULFIDE COMPLEX, 20 STRUCTURES | | Descriptor: | GLUTAREDOXIN 3, GLUTATHIONE | | Authors: | Nordstrand, K, Aslund, F, Holmgren, A, Otting, G, Berndt, K.D. | | Deposit date: | 1998-08-17 | | Release date: | 1999-03-30 | | Last modified: | 2018-03-14 | | Method: | SOLUTION NMR | | Cite: | NMR structure of Escherichia coli glutaredoxin 3-glutathione mixed disulfide complex: implications for the enzymatic mechanism.
J.Mol.Biol., 286, 1999
|
|
1MZ4
 
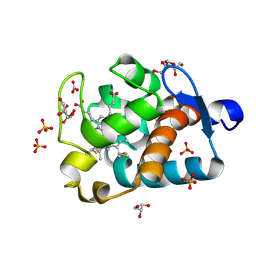 | | Crystal Structure of Cytochrome c550 from Thermosynechococcus elongatus | | Descriptor: | BICARBONATE ION, GLYCEROL, HEME C, ... | | Authors: | Kerfeld, C.A, Sawaya, M.R, Bottin, H, Tran, K.T, Sugiura, M, Kirilovsky, D, Krogmann, D, Yeates, T.O, Boussac, A. | | Deposit date: | 2002-10-05 | | Release date: | 2003-09-23 | | Last modified: | 2021-03-03 | | Method: | X-RAY DIFFRACTION (1.8 Å) | | Cite: | Structural and EPR characterization of the soluble form of cytochrome c-550 and of the psbV2 gene product from the cyanobacterium Thermosynechococcus elongatus.
Plant Cell.Physiol., 44, 2003
|
|
8VTJ
 
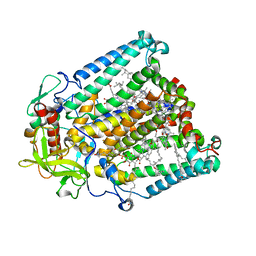 | |
1W69
 
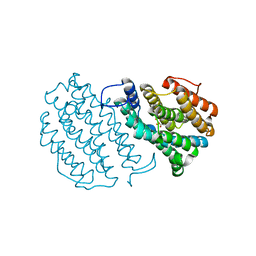 | | Crystal Structure of Mouse Ribonucleotide Reductase Subunit R2 under Reducing Conditions. A Fully Occupied Dinuclear Iron Cluster and Bound Acetate. | | Descriptor: | ACETIC ACID, FE (II) ION, RIBONUCLEOSIDE-DIPHOSPHATE REDUCTASE M2 CHAIN | | Authors: | Karlsen, S, Strand, K.R, Kolberg, M, Rohr, A.K, Gorbitz, C.H, Andersson, K.K. | | Deposit date: | 2004-08-16 | | Release date: | 2004-08-26 | | Last modified: | 2023-12-13 | | Method: | X-RAY DIFFRACTION (2.2 Å) | | Cite: | Crystal Structural Studies of Changes in the Native Dinuclear Iron Center of Ribonucleotide Reductase Protein R2 from Mouse
J.Biol.Chem., 279, 2004
|
|
1W68
 
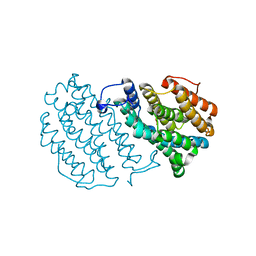 | | Crystal Structure of Mouse Ribonucleotide Reductase Subunit R2 under Oxidizing Conditions. A Fully Occupied Dinuclear Iron Cluster. | | Descriptor: | MU-OXO-DIIRON, RIBONUCLEOSIDE-DIPHOSPHATE REDUCTASE M2 CHAIN | | Authors: | Karlsen, S, Strand, K.R, Kolberg, M, Rohr, A.K, Gorbitz, C.H, Andersson, K.K. | | Deposit date: | 2004-08-16 | | Release date: | 2004-08-26 | | Last modified: | 2023-12-13 | | Method: | X-RAY DIFFRACTION (2.2 Å) | | Cite: | Crystal Structural Studies of Changes in the Native Dinuclear Iron Center of Ribonucleotide Reductase Protein R2 from Mouse
J.Biol.Chem., 279, 2004
|
|
2GTL
 
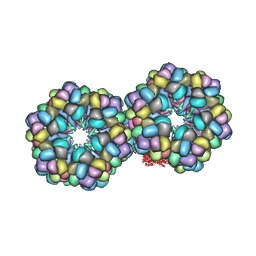 | | Lumbricus Erythrocruorin at 3.5A resolution | | Descriptor: | CALCIUM ION, CARBON MONOXIDE, Extracellular globin 2, ... | | Authors: | Royer Jr, W.E, Sharma, H, Strand, K, Knapp, J.E, Bhyravbhatla, B. | | Deposit date: | 2006-04-28 | | Release date: | 2006-07-18 | | Last modified: | 2011-07-13 | | Method: | X-RAY DIFFRACTION (3.5 Å) | | Cite: | Lumbricus erythrocruorin at 3.5 a resolution: architecture of a megadalton respiratory complex.
Structure, 14, 2006
|
|
2AMB
 
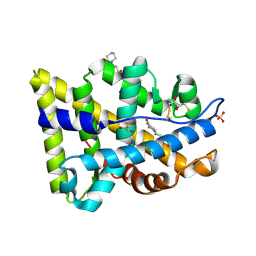 | | Crystal structure of human androgen receptor ligand binding domain in complex with tetrahydrogestrinone | | Descriptor: | 17-HYDROXY-18A-HOMO-19-NOR-17ALPHA-PREGNA-4,9,11-TRIEN-3-ONE, 2,3-DIHYDROXY-1,4-DITHIOBUTANE, 4-(2-HYDROXYETHYL)-1-PIPERAZINE ETHANESULFONIC ACID, ... | | Authors: | Pereira de Jesus-Tran, K, Cote, P.-L, Cantin, L, Blanchet, J, Labrie, F, Breton, R. | | Deposit date: | 2005-08-09 | | Release date: | 2006-05-16 | | Last modified: | 2023-08-23 | | Method: | X-RAY DIFFRACTION (1.75 Å) | | Cite: | Comparison of crystal structures of human androgen receptor ligand-binding domain complexed with various agonists reveals molecular determinants responsible for binding affinity.
Protein Sci., 15, 2006
|
|
4Q2Z
 
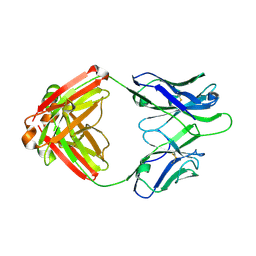 | | Fab fragment of HIV vaccine-elicited CD4bs-directed antibody, GE356, from a non-human primate | | Descriptor: | Heavy chain of Fab fragment of HIV vaccine-elicited CD4bs-directed antibody, Light chain of Fab fragment of HIV vaccine-elicited CD4bs-directed antibody | | Authors: | Navis, M, Tran, K, Bale, S, Phad, G, Guenaga, J, Wilson, R, Soldemo, M, McKee, K, Sundling, C, Mascola, J, Li, Y, Wyatt, R.T, Hedestam, G.B.K. | | Deposit date: | 2014-04-10 | | Release date: | 2014-09-17 | | Last modified: | 2023-09-20 | | Method: | X-RAY DIFFRACTION (1.93 Å) | | Cite: | HIV-1 Receptor Binding Site-Directed Antibodies Using a VH1-2 Gene Segment Orthologue Are Activated by Env Trimer Immunization.
Plos Pathog., 10, 2014
|
|
2IPJ
 
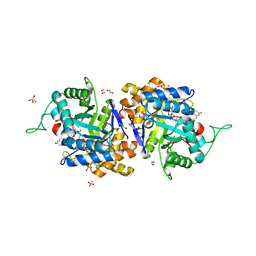 | | Crystal structure of h3alpha-hydroxysteroid dehydrogenase type 3 mutant Y24A in complex with NADP+ and epi-testosterone | | Descriptor: | (10ALPHA,13ALPHA,14BETA,17ALPHA)-17-HYDROXYANDROST-4-EN-3-ONE, 1,2-ETHANEDIOL, Aldo-keto reductase family 1 member C2, ... | | Authors: | Faucher, F, Cantin, L, Pereira de Jesus-Tran, K, Luu-the, V, Labrie, F, Breton, R. | | Deposit date: | 2006-10-12 | | Release date: | 2007-06-19 | | Last modified: | 2023-08-30 | | Method: | X-RAY DIFFRACTION (1.8 Å) | | Cite: | Mouse 17alpha-Hydroxysteroid Dehydrogenase (AKR1C21) Binds Steroids Differently from other Aldo-keto Reductases: Identification and Characterization of Amino Acid Residues Critical for Substrate Binding.
J.Mol.Biol., 369, 2007
|
|
2PNU
 
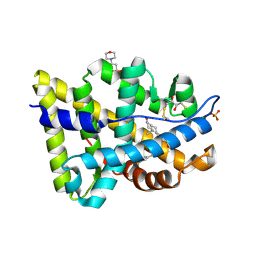 | | Crystal structure of human androgen receptor ligand-binding domain in complex with EM-5744 | | Descriptor: | (5S,8R,9S,10S,13R,14S,17S)-13-{2-[(3,5-DIFLUOROBENZYL)OXY]ETHYL}-17-HYDROXY-10-METHYLHEXADECAHYDRO-3H-CYCLOPENTA[A]PHENANTHREN-3-ONE, 2,3-DIHYDROXY-1,4-DITHIOBUTANE, 2-(N-MORPHOLINO)-ETHANESULFONIC ACID, ... | | Authors: | Cantin, L, Faucher, F, Couture, J.F, Pereira de Jesus-Tran, K, Legrand, P, Ciobanu, C.L, Singh, S.M, Labrie, F, Breton, R. | | Deposit date: | 2007-04-25 | | Release date: | 2007-09-11 | | Last modified: | 2023-08-30 | | Method: | X-RAY DIFFRACTION (1.65 Å) | | Cite: | Structural Characterization of the Human Androgen Receptor Ligand-binding Domain Complexed with EM5744, a Rationally Designed Steroidal Ligand Bearing a Bulky Chain Directed toward Helix 12.
J.Biol.Chem., 282, 2007
|
|
2IPG
 
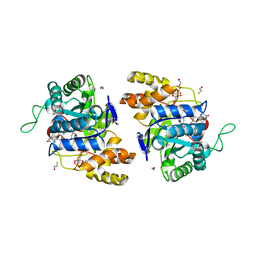 | | Crystal structure of 17alpha-hydroxysteroid dehydrogenase mutant K31A in complex with NADP+ and epi-testosterone | | Descriptor: | (10ALPHA,13ALPHA,14BETA,17ALPHA)-17-HYDROXYANDROST-4-EN-3-ONE, 1,2-ETHANEDIOL, 3(17)alpha-hydroxysteroid dehydrogenase, ... | | Authors: | Faucher, F, Cantin, L, Pereira de Jesus-Tran, K, Lemieux, M, Luu-the, V, Labrie, F, Breton, R. | | Deposit date: | 2006-10-12 | | Release date: | 2007-06-19 | | Last modified: | 2023-08-30 | | Method: | X-RAY DIFFRACTION (1.9 Å) | | Cite: | Mouse 17alpha-Hydroxysteroid Dehydrogenase (AKR1C21) Binds Steroids Differently from other Aldo-keto Reductases: Identification and Characterization of Amino Acid Residues Critical for Substrate Binding.
J.Mol.Biol., 369, 2007
|
|
2IPF
 
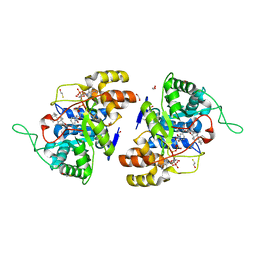 | | Crystal structure of 17alpha-hydroxysteroid dehydrogenase in complex with NADP+ and epi-testosterone | | Descriptor: | (10ALPHA,13ALPHA,14BETA,17ALPHA)-17-HYDROXYANDROST-4-EN-3-ONE, (3(17)alpha-hydroxysteroid dehydrogenase), 1,2-ETHANEDIOL, ... | | Authors: | Faucher, F, Cantin, L, Pereira de Jesus-Tran, K, Luu-the, V, Labrie, F, Breton, R. | | Deposit date: | 2006-10-12 | | Release date: | 2007-06-19 | | Last modified: | 2023-08-30 | | Method: | X-RAY DIFFRACTION (1.85 Å) | | Cite: | Mouse 17alpha-Hydroxysteroid Dehydrogenase (AKR1C21) Binds Steroids Differently from other Aldo-keto Reductases: Identification and Characterization of Amino Acid Residues Critical for Substrate Binding.
J.Mol.Biol., 369, 2007
|
|
2HE8
 
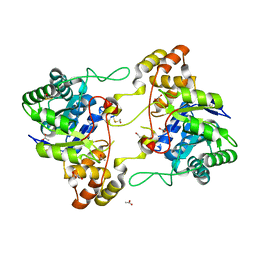 | | Crystal structure of 17alpha-hydroxysteroid dehydrogenase in its apo-form | | Descriptor: | ACETATE ION, Aldo-keto reductase family 1, member C21, ... | | Authors: | Faucher, F, Pereira de Jesus-Tran, K, Cantin, L, Luu-the, V, Labrie, F, Breton, R. | | Deposit date: | 2006-06-21 | | Release date: | 2006-12-05 | | Last modified: | 2023-12-06 | | Method: | X-RAY DIFFRACTION (1.9 Å) | | Cite: | Crystal Structures of Mouse 17alpha-Hydroxysteroid Dehydrogenase (Apoenzyme and Enzyme-NADP(H) Binary Complex): Identification of Molecular Determinants Responsible for the Unique 17alpha-reductive Activity of this Enzyme.
J.Mol.Biol., 364, 2006
|
|
2HDJ
 
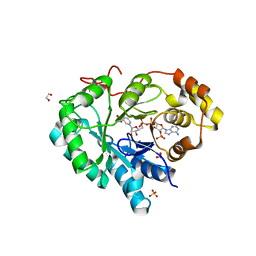 | | Crystal structure of human type 3 3alpha-hydroxysteroid dehydrogenase in complex with NADP(H) | | Descriptor: | 1,2-ETHANEDIOL, Aldo-keto reductase family 1 member C2, NADPH DIHYDRO-NICOTINAMIDE-ADENINE-DINUCLEOTIDE PHOSPHATE, ... | | Authors: | Faucher, F, Pereira de Jesus-Tran, K, Cantin, L, Luu-the, V, Labrie, F, Breton, R. | | Deposit date: | 2006-06-20 | | Release date: | 2006-12-05 | | Last modified: | 2023-08-30 | | Method: | X-RAY DIFFRACTION (2 Å) | | Cite: | Crystal Structures of Mouse 17alpha-Hydroxysteroid Dehydrogenase (Apoenzyme and Enzyme-NADP(H) Binary Complex): Identification of Molecular Determinants Responsible for the Unique 17alpha-reductive Activity of this Enzyme.
J.Mol.Biol., 364, 2006
|
|
2HE5
 
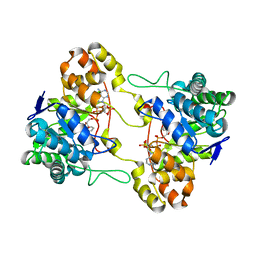 | | Crystal structure of 17alpha-hydroxysteroid dehydrogenase in binary complex with NADP(H) in an open conformation | | Descriptor: | ACETATE ION, Aldo-keto reductase family 1, member C21, ... | | Authors: | Faucher, F, Pereira de Jesus-Tran, K, Cantin, L, Luu-the, V, Labrie, F, Breton, R. | | Deposit date: | 2006-06-21 | | Release date: | 2006-12-05 | | Last modified: | 2024-02-14 | | Method: | X-RAY DIFFRACTION (2.9 Å) | | Cite: | Crystal Structures of Mouse 17alpha-Hydroxysteroid Dehydrogenase (Apoenzyme and Enzyme-NADP(H) Binary Complex): Identification of Molecular Determinants Responsible for the Unique 17alpha-reductive Activity of this Enzyme.
J.Mol.Biol., 364, 2006
|
|
2HEJ
 
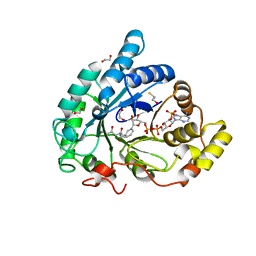 | | Crystal structure of 17alpha-hydroxysteroid dehydrogenase in complex with NADP(H) in a closed conformation | | Descriptor: | (4S)-2-METHYL-2,4-PENTANEDIOL, 1,2-ETHANEDIOL, Aldo-keto reductase family 1, ... | | Authors: | Faucher, F, Pereira de Jesus-Tran, K, Cantin, L, Luu-the, V, Labrie, F, Breton, R. | | Deposit date: | 2006-06-21 | | Release date: | 2006-12-05 | | Last modified: | 2023-08-30 | | Method: | X-RAY DIFFRACTION (1.35 Å) | | Cite: | Crystal Structures of Mouse 17alpha-Hydroxysteroid Dehydrogenase (Apoenzyme and Enzyme-NADP(H) Binary Complex): Identification of Molecular Determinants Responsible for the Unique 17alpha-reductive Activity of this Enzyme.
J.Mol.Biol., 364, 2006
|
|
