7Z01
 
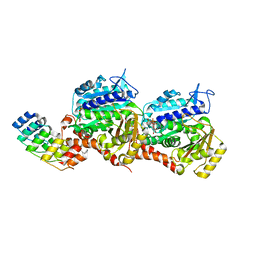 | | Z-SBTubA4 photoswitch bound to tubulin-DARPin D1 complex | | Descriptor: | 2-methoxy-5-[2-(5,6,7-trimethoxy-1,3-benzothiazol-2-yl)ethyl]phenol, CALCIUM ION, Designed Ankyrin Repeat Protein (DARPIN) D1, ... | | Authors: | Wranik, M, Weinert, T, Standfuss, J, Steinmetz, M. | | Deposit date: | 2022-02-21 | | Release date: | 2022-03-30 | | Last modified: | 2024-01-31 | | Method: | X-RAY DIFFRACTION (1.82 Å) | | Cite: | In Vivo Photocontrol of Microtubule Dynamics and Integrity, Migration and Mitosis, by the Potent GFP-Imaging-Compatible Photoswitchable Reagents SBTubA4P and SBTub2M.
J.Am.Chem.Soc., 144, 2022
|
|
7Z02
 
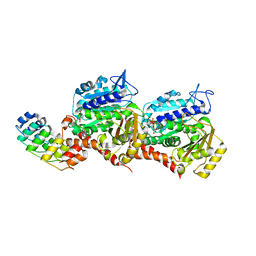 | | Z-SBTub2M photoswitch bound to tubulin-DARPin D1 complex | | Descriptor: | 6-methyl-2-[2-(3,4,5-trimethoxyphenyl)ethyl]-1,3-benzothiazole, Designed Ankyrin Repeat Protein (DARPIN) D1, GUANOSINE-5'-DIPHOSPHATE, ... | | Authors: | Wranik, M, Weinert, T, Standfuss, J, Steinmetz, M. | | Deposit date: | 2022-02-21 | | Release date: | 2022-03-30 | | Last modified: | 2024-01-31 | | Method: | X-RAY DIFFRACTION (2.36 Å) | | Cite: | In Vivo Photocontrol of Microtubule Dynamics and Integrity, Migration and Mitosis, by the Potent GFP-Imaging-Compatible Photoswitchable Reagents SBTubA4P and SBTub2M.
J.Am.Chem.Soc., 144, 2022
|
|
6RPH
 
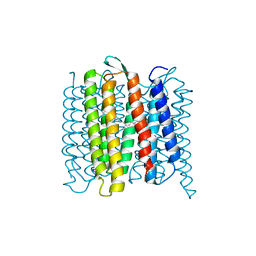 | | TR-SMX open state structure (10-15ms) of bacteriorhodopsin | | Descriptor: | Bacteriorhodopsin, RETINAL | | Authors: | Weinert, T, Skopintsev, P, James, D, Kekilli, D, Furrer, A, Bruenle, S, Mous, S, Nogly, P, Standfuss, J. | | Deposit date: | 2019-05-14 | | Release date: | 2019-07-17 | | Last modified: | 2019-07-24 | | Method: | X-RAY DIFFRACTION (2.6 Å) | | Cite: | Proton uptake mechanism in bacteriorhodopsin captured by serial synchrotron crystallography.
Science, 365, 2019
|
|
6EKA
 
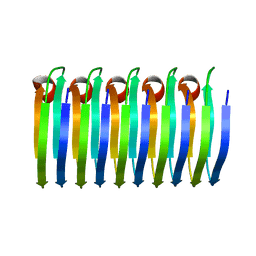 | | Solid-state MAS NMR structure of the HELLF prion amyloid fibrils | | Descriptor: | Podospora anserina S mat+ genomic DNA chromosome 3, supercontig 2 | | Authors: | Martinez, D, Daskalov, A, Andreas, L, Bardiaux, B, Coustou, V, Stanek, J, Berbon, M, Noubhani, M, Kauffmann, B, Wall, J.S, Pintacuda, G, Saupe, S.J, Habenstein, B, Loquet, A. | | Deposit date: | 2017-09-25 | | Release date: | 2018-10-10 | | Last modified: | 2024-06-19 | | Method: | SOLID-STATE NMR | | Cite: | Structural and molecular basis of cross-seeding barriers in amyloids
Proc.Natl.Acad.Sci.USA, 118, 2021
|
|
7YYW
 
 | | Molecular snapshots of drug release from tubulin: 10 nanoseconds after photoactivation. | | Descriptor: | Azo-Combretastatin A4 (trans), CALCIUM ION, Designed Ankyrin Repeat Protein (DARPIN) D1, ... | | Authors: | Wranik, M, Weinert, T, Standfuss, J. | | Deposit date: | 2022-02-18 | | Release date: | 2023-02-22 | | Last modified: | 2023-09-27 | | Method: | X-RAY DIFFRACTION (2.2 Å) | | Cite: | Watching the release of a photopharmacological drug from tubulin using time-resolved serial crystallography.
Nat Commun, 14, 2023
|
|
7YZ5
 
 | | Molecular snapshots of drug release from tubulin: 100 milliseconds (steady state) | | Descriptor: | CALCIUM ION, Designed Ankyrin Repeat Protein (DARPIN) D1, GUANOSINE-5'-DIPHOSPHATE, ... | | Authors: | Wranik, M, Weinert, T, Standfuss, J. | | Deposit date: | 2022-02-18 | | Release date: | 2023-02-22 | | Last modified: | 2024-02-07 | | Method: | X-RAY DIFFRACTION (2.11 Å) | | Cite: | Watching the release of a photopharmacological drug from tubulin using time-resolved serial crystallography.
Nat Commun, 14, 2023
|
|
7YZ2
 | | Molecular snapshots of drug release from tubulin: 10 milliseconds after photoactivation. | | Descriptor: | Designed Ankyrin Repeat Protein (DARPIN) D1, GUANOSINE-5'-DIPHOSPHATE, GUANOSINE-5'-TRIPHOSPHATE, ... | | Authors: | Wranik, M, Weinert, T, Standfuss, J. | | Deposit date: | 2022-02-18 | | Release date: | 2023-02-22 | | Last modified: | 2024-02-07 | | Method: | X-RAY DIFFRACTION (2.2 Å) | | Cite: | Watching the release of a photopharmacological drug from tubulin using time-resolved serial crystallography.
Nat Commun, 14, 2023
|
|
7YYV
 
 | | Molecular snapshots of drug release from tubulin: 1 nanosecond after photoactivation. | | Descriptor: | Azo-Combretastatin A4 (trans), CALCIUM ION, Designed Ankyrin Repeat Protein (DARPIN) D1, ... | | Authors: | Wranik, M, Weinert, T, Standfuss, J. | | Deposit date: | 2022-02-18 | | Release date: | 2023-02-22 | | Last modified: | 2024-02-07 | | Method: | X-RAY DIFFRACTION (2.2 Å) | | Cite: | Watching the release of a photopharmacological drug from tubulin using time-resolved serial crystallography.
Nat Commun, 14, 2023
|
|
7YYZ
 | | Molecular snapshots of drug release from tubulin: 10 microseconds after photoactivation. | | Descriptor: | Azo-Combretastatin A4 (trans), CALCIUM ION, Designed Ankyrin Repeat Protein (DARPIN) D1, ... | | Authors: | Wranik, M, Weinert, T, Standfuss, J. | | Deposit date: | 2022-02-18 | | Release date: | 2023-02-22 | | Last modified: | 2024-02-07 | | Method: | X-RAY DIFFRACTION (2.2 Å) | | Cite: | Watching the release of a photopharmacological drug from tubulin using time-resolved serial crystallography.
Nat Commun, 14, 2023
|
|
7YZ1
 
 | | Molecular snapshots of drug release from tubulin: 1 millisecond after photoactivation. | | Descriptor: | Azo-Combretastatin A4 (trans), CALCIUM ION, Designed Ankyrin Repeat Protein (DARPIN) D1, ... | | Authors: | Wranik, M, Weinert, T, Standfuss, J. | | Deposit date: | 2022-02-18 | | Release date: | 2023-02-22 | | Last modified: | 2023-09-27 | | Method: | X-RAY DIFFRACTION (2.2 Å) | | Cite: | Watching the release of a photopharmacological drug from tubulin using time-resolved serial crystallography.
Nat Commun, 14, 2023
|
|
7YZ6
 
 | | Molecular snapshots of drug release from tubulin: Dark (steady state) | | Descriptor: | Azo-Combretastatin A4 (cis), CALCIUM ION, Designed Ankyrin Repeat Protein (DARPIN) D1, ... | | Authors: | Wranik, M, Weinert, T, Standfuss, J. | | Deposit date: | 2022-02-18 | | Release date: | 2023-02-22 | | Last modified: | 2023-09-27 | | Method: | X-RAY DIFFRACTION (2.1 Å) | | Cite: | Watching the release of a photopharmacological drug from tubulin using time-resolved serial crystallography.
Nat Commun, 14, 2023
|
|
7YYX
 
 | | Molecular snapshots of drug release from tubulin: 100 nanoseconds after photoactivation. | | Descriptor: | Azo-Combretastatin A4 (trans), CALCIUM ION, Designed Ankyrin Repeat Protein (DARPIN) D1, ... | | Authors: | Wranik, M, Weinert, T, Standfuss, J. | | Deposit date: | 2022-02-18 | | Release date: | 2023-02-22 | | Last modified: | 2023-09-27 | | Method: | X-RAY DIFFRACTION (2.2 Å) | | Cite: | Watching the release of a photopharmacological drug from tubulin using time-resolved serial crystallography.
Nat Commun, 14, 2023
|
|
7YZ0
 
 | | Molecular snapshots of drug release from tubulin: 100 microseconds after photoactivation. | | Descriptor: | Azo-Combretastatin A4 (trans), CALCIUM ION, Designed Ankyrin Repeat Protein (DARPIN) D1, ... | | Authors: | Wranik, M, Weinert, T, Standfuss, J. | | Deposit date: | 2022-02-18 | | Release date: | 2023-02-22 | | Last modified: | 2024-02-07 | | Method: | X-RAY DIFFRACTION (2.2 Å) | | Cite: | Watching the release of a photopharmacological drug from tubulin using time-resolved serial crystallography.
Nat Commun, 14, 2023
|
|
7YZ3
 
 | | Molecular snapshots of drug release from tubulin: Apo state | | Descriptor: | CALCIUM ION, Designed Ankyrin Repeat Protein (DARPIN) D1, GUANOSINE-5'-DIPHOSPHATE, ... | | Authors: | Wranik, M, Weinert, T, Standfuss, J. | | Deposit date: | 2022-02-18 | | Release date: | 2023-02-22 | | Last modified: | 2024-02-07 | | Method: | X-RAY DIFFRACTION (1.8 Å) | | Cite: | Watching the release of a photopharmacological drug from tubulin using time-resolved serial crystallography.
Nat Commun, 14, 2023
|
|
7YYY
 
 | | Molecular snapshots of drug release from tubulin: 1 microsecond after photoactivation | | Descriptor: | Azo-Combretastatin A4 (trans), CALCIUM ION, Designed Ankyrin Repeat Protein (DARPIN) D1, ... | | Authors: | Wranik, M, Weinert, T, Standfuss, J. | | Deposit date: | 2022-02-18 | | Release date: | 2023-02-22 | | Last modified: | 2023-09-27 | | Method: | X-RAY DIFFRACTION (2.2 Å) | | Cite: | Watching the release of a photopharmacological drug from tubulin using time-resolved serial crystallography.
Nat Commun, 14, 2023
|
|
7YYQ
 
 | | Molecular snapshots of drug release from tubulin: Dark state | | Descriptor: | Azo-Combretastatin A4 (cis), CALCIUM ION, Designed Ankyrin Repeat Protein (DARPIN) D1, ... | | Authors: | Wranik, M, Weinert, T, Standfuss, J. | | Deposit date: | 2022-02-18 | | Release date: | 2023-02-22 | | Last modified: | 2023-09-27 | | Method: | X-RAY DIFFRACTION (1.7 Å) | | Cite: | Watching the release of a photopharmacological drug from tubulin using time-resolved serial crystallography.
Nat Commun, 14, 2023
|
|
6F2S
 
 | | CryoEM structure of Ageratum Yellow Vein virus (AYVV) | | Descriptor: | Capsid protein, coat protein subunit H, coat protein subunit I, ... | | Authors: | Hesketh, E.L, Saunders, K, Fisher, C, Potze, J, Stanley, J, Lomonossoff, G.P, Ranson, N.A. | | Deposit date: | 2017-11-27 | | Release date: | 2018-06-27 | | Last modified: | 2024-05-15 | | Method: | ELECTRON MICROSCOPY (3.3 Å) | | Cite: | The 3.3 angstrom structure of a plant geminivirus using cryo-EM.
Nat Commun, 9, 2018
|
|
5J78
 
 | | Crystal structure of an Acetylating Aldehyde Dehydrogenase from Geobacillus thermoglucosidasius | | Descriptor: | ACETATE ION, Acetaldehyde dehydrogenase (Acetylating), GLYCEROL, ... | | Authors: | Crennell, S.J, Extance, J.P, Danson, M.J. | | Deposit date: | 2016-04-06 | | Release date: | 2016-09-07 | | Last modified: | 2024-01-10 | | Method: | X-RAY DIFFRACTION (2.1 Å) | | Cite: | Structure of an acetylating aldehyde dehydrogenase from the thermophilic ethanologen Geobacillus thermoglucosidasius.
Protein Sci., 25, 2016
|
|
5EDS
 
 | | Crystal structure of human PI3K-gamma in complex with benzimidazole inhibitor 5 | | Descriptor: | 4-azanyl-6-[[(1~{S})-1-[6-fluoranyl-1-(3-methylsulfonylphenyl)benzimidazol-2-yl]ethyl]amino]pyrimidine-5-carbonitrile, Phosphatidylinositol 4,5-bisphosphate 3-kinase catalytic subunit gamma isoform, SULFATE ION | | Authors: | Whittington, D.A, Tang, J, Yakowec, P. | | Deposit date: | 2015-10-21 | | Release date: | 2015-12-30 | | Last modified: | 2023-09-27 | | Method: | X-RAY DIFFRACTION (2.8 Å) | | Cite: | Discovery, Optimization, and in Vivo Evaluation of Benzimidazole Derivatives AM-8508 and AM-9635 as Potent and Selective PI3K delta Inhibitors.
J.Med.Chem., 59, 2016
|
|
6T4D
 
 | | Crystal structure of Plasmodium falciparum Morn1 | | Descriptor: | Morn1, ZINC ION | | Authors: | Grishkovskaya, I, Kostan, J, Sajko, S, Morriswood, B, Djinovic-Carugo, K. | | Deposit date: | 2019-10-13 | | Release date: | 2020-11-18 | | Last modified: | 2024-05-15 | | Method: | X-RAY DIFFRACTION (2.14 Å) | | Cite: | Structures of three MORN repeat proteins and a re-evaluation of the proposed lipid-binding properties of MORN repeats.
Plos One, 15, 2020
|
|
6FKA
 
 | | Crystal structure of N2C/D282C stabilized opsin bound to RS11 | | Descriptor: | (2~{S})-2-(3,4-dichlorophenyl)-3-methyl-1-spiro[1,3-benzodioxole-2,4'-piperidine]-1'-yl-butan-1-one, PALMITIC ACID, Rhodopsin, ... | | Authors: | Mattle, D, Standfuss, J, Dawson, R. | | Deposit date: | 2018-01-23 | | Release date: | 2018-04-04 | | Last modified: | 2024-01-17 | | Method: | X-RAY DIFFRACTION (2.7 Å) | | Cite: | Ligand channel in pharmacologically stabilized rhodopsin.
Proc. Natl. Acad. Sci. U.S.A., 115, 2018
|
|
6FKC
 
 | | Crystal structure of N2C/D282C stabilized opsin bound to RS15 | | Descriptor: | 3-[1'-[(2~{S})-2-(4-chlorophenyl)-3-methyl-butanoyl]spiro[1,3-benzodioxole-2,4'-piperidine]-5-yl]propanoic acid, PALMITIC ACID, Rhodopsin, ... | | Authors: | Mattle, D, Standfuss, J, Dawson, R. | | Deposit date: | 2018-01-23 | | Release date: | 2018-04-04 | | Last modified: | 2024-01-17 | | Method: | X-RAY DIFFRACTION (2.46 Å) | | Cite: | Ligand channel in pharmacologically stabilized rhodopsin.
Proc. Natl. Acad. Sci. U.S.A., 115, 2018
|
|
6FK7
 
 | | Crystal structure of N2C/D282C stabilized opsin bound to RS06 | | Descriptor: | (2~{R},3~{R})-2-(4-chlorophenyl)-3-oxidanyl-1-spiro[1,3-benzodioxole-2,4'-piperidine]-1'-yl-butan-1-one, PALMITIC ACID, Rhodopsin, ... | | Authors: | Mattle, D, Standfuss, J, Dawson, R. | | Deposit date: | 2018-01-23 | | Release date: | 2018-04-04 | | Last modified: | 2024-01-17 | | Method: | X-RAY DIFFRACTION (2.62 Å) | | Cite: | Ligand channel in pharmacologically stabilized rhodopsin.
Proc. Natl. Acad. Sci. U.S.A., 115, 2018
|
|
6FKD
 
 | | Crystal structure of N2C/D282C stabilized opsin bound to RS16 | | Descriptor: | 5-chloranyl-2-(2-oxidanylidene-2-spiro[1,3-benzodioxole-2,4'-piperidine]-1'-yl-ethyl)-3~{H}-pyridin-6-one, PALMITIC ACID, Rhodopsin, ... | | Authors: | Mattle, D, Standfuss, J, Dawson, R. | | Deposit date: | 2018-01-23 | | Release date: | 2018-04-04 | | Last modified: | 2024-01-17 | | Method: | X-RAY DIFFRACTION (2.49 Å) | | Cite: | Ligand channel in pharmacologically stabilized rhodopsin.
Proc. Natl. Acad. Sci. U.S.A., 115, 2018
|
|
6FUF
 
 | | Crystal structure of the rhodopsin-mini-Go complex | | Descriptor: | 2-acetamido-2-deoxy-beta-D-glucopyranose, Guanine nucleotide-binding protein G(o) subunit alpha, RETINAL, ... | | Authors: | Tsai, C.-J, Weinert, T, Muehle, J, Pamula, F, Nehme, R, Flock, T, Nogly, P, Edwards, P.C, Carpenter, B, Gruhl, T, Ma, P, Deupi, X, Standfuss, J, Tate, C.G, Schertler, G.F.X. | | Deposit date: | 2018-02-27 | | Release date: | 2018-10-03 | | Last modified: | 2024-01-17 | | Method: | X-RAY DIFFRACTION (3.117 Å) | | Cite: | Crystal structure of rhodopsin in complex with a mini-Gosheds light on the principles of G protein selectivity.
Sci Adv, 4, 2018
|
|
