3F5D
 
 | |
3FD8
 
 | |
3FII
 
 | |
3CK5
 
 | |
3CO8
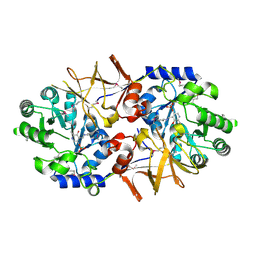 
 | | Crystal structure of alanine racemase from Oenococcus oeni | | Descriptor: | Alanine racemase, PYRIDOXAL-5'-PHOSPHATE | | Authors: | Palani, K, Burley, S.K, Swaminathan, S, New York SGX Research Center for Structural Genomics (NYSGXRC) | | Deposit date: | 2008-03-27 | | Release date: | 2008-04-08 | | Last modified: | 2021-02-03 | | Method: | X-RAY DIFFRACTION (1.7 Å) | | Cite: | Structure of alanine racemase from Oenococcus oeni with bound pyridoxal 5'-phosphate.
Acta Crystallogr.,Sect.F, 69, 2013
|
|
3CYG
 
 | |
3D8K
 
 | |
3DEB
 
 | |
3DLI
 
 | |
3EWN
 
 | |
3FJY
 
 | |
3FIJ
 | |
3CMN
 
 | |
3D3X
 
 | |
3CVG
 
 | |
3D3A
 
 | |
3D5L
 
 | |
3DMY
 
 | |
3DH0
 
 | |
3F6A
 
 | |
3FIE
 
 | |
3FND
 
 | |
3E3V
 
 | |
3G13
 
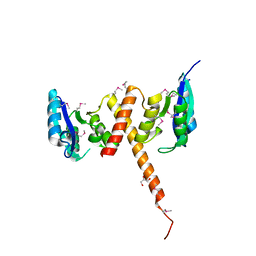 | |
3DEC
 
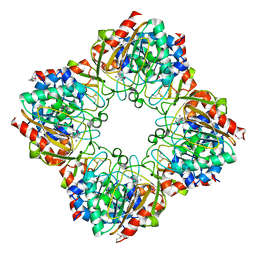
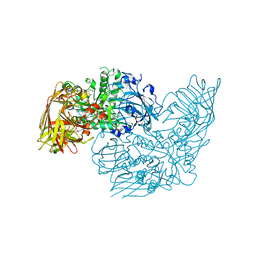 | | Crystal structure of a glycosyl hydrolases family 2 protein from Bacteroides thetaiotaomicron | | Descriptor: | Beta-galactosidase, POTASSIUM ION | | Authors: | Kumaran, D, Bonanno, J, Romero, R, Burley, S.K, Swaminathan, S, New York SGX Research Center for Structural Genomics (NYSGXRC) | | Deposit date: | 2008-06-09 | | Release date: | 2008-06-17 | | Last modified: | 2023-11-15 | | Method: | X-RAY DIFFRACTION (2.8 Å) | | Cite: | Crystal Structure of a Glycosyl Hydrolases Family 2 protein from Bacteroides thetaiotaomicron.
To be Published
|
|
