7R8F
 
 | |
7R8K
 
 | |
7R8G
 
 | |
7R8H
 
 | |
7R8I
 
 | |
7SZJ
 
 | |
7SZK
 
 | | Cryo-EM structure of 27a bound to E. coli RNAP and rrnBP1 promoter complex | | Descriptor: | (2S,7R,7aR,13aP,16Z,18E,20S,21S,22R,23R,24R,25S,26R,27S,28E)-5,21,23-trihydroxy-27-methoxy-2,4,16,20,22,24,26-heptamethyl-10-[4-(2-methylpropyl)piperazin-1-yl]-12-({4-[(morpholin-4-yl)methyl]phenyl}methoxy)-1,6,15-trioxo-1,2,7,7a-tetrahydro-6H-2,7-(epoxypentadeca[1,11,13]trienoimino)[1]benzofuro[4,5-a]phenoxazin-25-yl acetate, DNA (5'-D(P*CP*TP*CP*GP*TP*AP*GP*AP*GP*TP*CP*CP*GP*TP*GP*TP*CP*A)-3'), DNA-directed RNA polymerase subunit alpha, ... | | Authors: | Shin, Y, Murakami, K.S. | | Deposit date: | 2021-11-28 | | Release date: | 2022-07-13 | | Last modified: | 2024-02-28 | | Method: | ELECTRON MICROSCOPY (2.94 Å) | | Cite: | Optimization of Benzoxazinorifamycins to Improve Mycobacterium tuberculosis RNA Polymerase Inhibition and Treatment of Tuberculosis.
Acs Infect Dis., 8, 2022
|
|
7R8J
 
 | |
7KHE
 
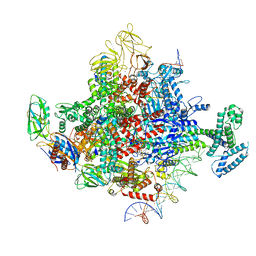 | | Escherichia coli RNA polymerase and rrnBP1 promoter pre-open complex with DksA/ppGpp | | Descriptor: | CHAPSO, DNA (46-MER), DNA (54-MER), ... | | Authors: | Shin, Y, Qayyum, M.Z, Murakami, K.S. | | Deposit date: | 2020-10-21 | | Release date: | 2020-12-09 | | Last modified: | 2024-03-06 | | Method: | ELECTRON MICROSCOPY (3.58 Å) | | Cite: | Structural basis of ribosomal RNA transcription regulation.
Nat Commun, 12, 2021
|
|
7KHB
 
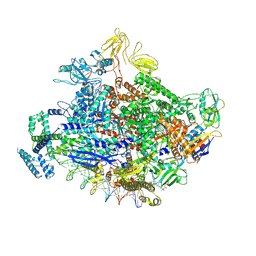 | | Escherichia coli RNA polymerase and rrnBP1 promoter open complex | | Descriptor: | CHAPSO, DNA (60-MER), DNA (64-MER), ... | | Authors: | Shin, Y, Qayyum, M.Z, Murakami, K.S. | | Deposit date: | 2020-10-20 | | Release date: | 2020-10-28 | | Last modified: | 2024-03-06 | | Method: | ELECTRON MICROSCOPY (3.53 Å) | | Cite: | Structural basis of ribosomal RNA transcription regulation.
Nat Commun, 12, 2021
|
|
7KHC
 
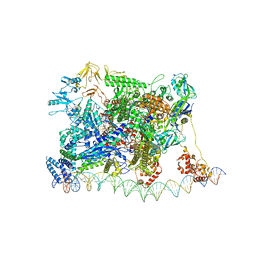 | | Escherichia coli RNA polymerase and rrnBP1 promoter closed complex | | Descriptor: | CHAPSO, DNA (18 MER), DNA (63-MER), ... | | Authors: | Shin, Y, Qayyum, M.Z, Murakami, K.S. | | Deposit date: | 2020-10-20 | | Release date: | 2020-10-28 | | Last modified: | 2024-03-06 | | Method: | ELECTRON MICROSCOPY (4.14 Å) | | Cite: | Structural basis of ribosomal RNA transcription regulation.
Nat Commun, 12, 2021
|
|
7KHI
 
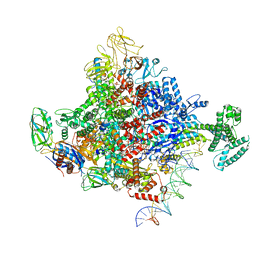 | | Escherichia coli RNA polymerase and rrnBP1 promoter complex with DksA/ppGpp | | Descriptor: | CHAPSO, DNA (28-MER), DNA (36-MER), ... | | Authors: | Shin, Y, Qayyum, M.Z, Murakami, K.S. | | Deposit date: | 2020-10-21 | | Release date: | 2020-10-28 | | Last modified: | 2024-03-06 | | Method: | ELECTRON MICROSCOPY (3.62 Å) | | Cite: | Structural basis of ribosomal RNA transcription regulation.
Nat Commun, 12, 2021
|
|
6WOX
 
 | |
6WOY
 
 | |
6OY7
 
 | |
6OVY
 
 | |
6OVR
 
 | |
6OW3
 
 | |
6OY6
 
 | |
6OY5
 
 | |
6P70
 
 | | X-ray crystal structure of bacterial RNA polymerase and pyrBI promoter complex | | Descriptor: | DNA (5'-D(*TP*AP*TP*AP*AP*TP*CP*GP*AP*TP*CP*TP*TP*TP*GP*CP*CP*GP*GP*G)-3'), DNA (5'-D(P*TP*CP*CP*CP*GP*GP*CP*AP*AP*AP*TP*TP*GP*TP*CP*CP*G)-3'), DNA-directed RNA polymerase subunit alpha, ... | | Authors: | Shin, Y, Murakami, K.S. | | Deposit date: | 2019-06-04 | | Release date: | 2019-06-19 | | Last modified: | 2023-10-11 | | Method: | X-RAY DIFFRACTION (3.052 Å) | | Cite: | Structural basis of reiterative transcription from the pyrG and pyrBI promoters by bacterial RNA polymerase.
Nucleic Acids Res., 48, 2020
|
|
6P71
 
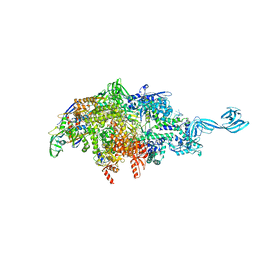 | |
4HJD
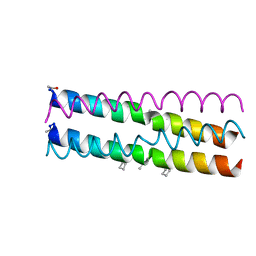 
 | | GCN4pLI derivative with alpha/beta/acyclic-gamma amino acid substitution pattern | | Descriptor: | GCN4pLI(alpha/beta/acyclic gamma) | | Authors: | Shin, Y.H, Mortenson, D.E, Satyshur, K.A, Forest, K.T, Gellman, S.H. | | Deposit date: | 2012-10-12 | | Release date: | 2013-06-12 | | Last modified: | 2023-11-15 | | Method: | X-RAY DIFFRACTION (1.7 Å) | | Cite: | Differential Impact of beta and gamma Residue Preorganization on alpha / beta / gamma-Peptide Helix Stability in Water.
J.Am.Chem.Soc., 135, 2013
|
|
4HJB
 
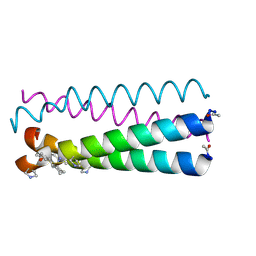 | | GCN4pLI derivative with alpha/beta/cyclic-gamma amino acid substitution pattern | | Descriptor: | GCN4pLI(alpha/beta/cyclic-gamma) | | Authors: | Shin, Y.H, Mortenson, D.E, Satyshur, K.A, Forest, K.T, Gellman, S.H. | | Deposit date: | 2012-10-12 | | Release date: | 2013-06-12 | | Last modified: | 2023-11-15 | | Method: | X-RAY DIFFRACTION (1.25 Å) | | Cite: | Differential Impact of beta and gamma Residue Preorganization on alpha / beta / gamma-Peptide Helix Stability in Water.
J.Am.Chem.Soc., 135, 2013
|
|
5EZ5
 
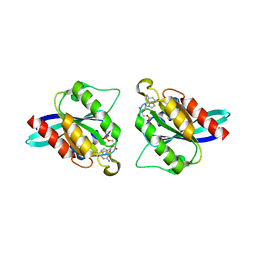 | |
