5O7A
 
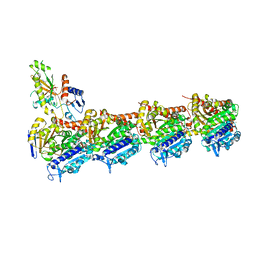 | | Quinolin-6-yloxyacetamides are microtubule destabilizing agents that bind to the colchicine site of tubulin | | Descriptor: | (2~{R})-2-(3-ethynylquinolin-6-yl)oxy-2-methoxy-~{N}-[(1~{E})-1-methoxyimino-2-methyl-propan-2-yl]ethanamide, GUANOSINE-5'-DIPHOSPHATE, GUANOSINE-5'-TRIPHOSPHATE, ... | | Authors: | Sharma, A, Olieric, N, Steinmetz, M.O. | | Deposit date: | 2017-06-08 | | Release date: | 2017-07-05 | | Last modified: | 2024-05-08 | | Method: | X-RAY DIFFRACTION (2.495 Å) | | Cite: | Quinolin-6-Yloxyacetamides Are Microtubule Destabilizing Agents That Bind to the Colchicine Site of Tubulin.
Int J Mol Sci, 18, 2017
|
|
3MIU
 
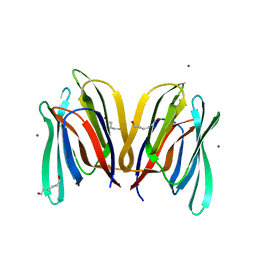 | | Structure of Banana Lectin-pentamannose complex | | Descriptor: | HEXANE-1,6-DIOL, Lectin, ZINC ION, ... | | Authors: | Sharma, A, Vijayan, M. | | Deposit date: | 2010-04-12 | | Release date: | 2010-09-08 | | Last modified: | 2023-11-01 | | Method: | X-RAY DIFFRACTION (2.63 Å) | | Cite: | Influence of glycosidic linkage on the nature of carbohydrate binding in beta-prism I fold lectins: an X-ray and molecular dynamics investigation on banana lectin-carbohydrate complexes
Glycobiology, 21, 2011
|
|
3MIV
 
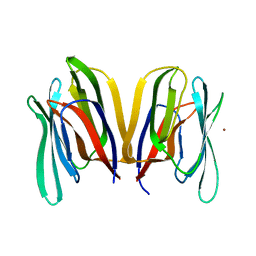 | | Structure of Banana lectin - Glc-alpha(1,2)-Glc complex | | Descriptor: | Lectin, ZINC ION | | Authors: | Sharma, A, Vijayan, M. | | Deposit date: | 2010-04-12 | | Release date: | 2010-09-08 | | Last modified: | 2023-11-01 | | Method: | X-RAY DIFFRACTION (3.5 Å) | | Cite: | Influence of glycosidic linkage on the nature of carbohydrate binding in beta-prism I fold lectins: an X-ray and molecular dynamics investigation on banana lectin-carbohydrate complexes
Glycobiology, 21, 2011
|
|
2OGW
 
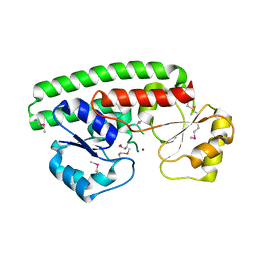 | | Structure of ABC type zinc transporter from E. coli | | Descriptor: | High-affinity zinc uptake system protein znuA precursor, ZINC ION | | Authors: | Sharma, A, Yogavel, M. | | Deposit date: | 2007-01-09 | | Release date: | 2007-03-06 | | Last modified: | 2023-12-27 | | Method: | X-RAY DIFFRACTION (1.83 Å) | | Cite: | Structural Analysis of ABC-family Periplasmic Zinc Binding Protein Provides New Insights Into Mechanism of Ligand Uptake and Release
J.Mol.Biol., 367, 2007
|
|
6S8L
 
 | | Structure, Thermodynamics, and Kinetics of Plinabulin Binding to two Tubulin Isotypes | | Descriptor: | (3Z,6Z)-3-benzylidene-6-[(5-tert-butyl-1H-imidazol-4-yl)methylidene]piperazine-2,5-dione, Designed ankyrin repeat protein (DARPIN) D1, GUANOSINE-5'-DIPHOSPHATE, ... | | Authors: | Sharma, A, Olieric, N, Steinmetz, M. | | Deposit date: | 2019-07-10 | | Release date: | 2019-11-27 | | Last modified: | 2024-05-15 | | Method: | X-RAY DIFFRACTION (1.801 Å) | | Cite: | Structure, Thermodynamics, and Kinetics of Plinabulin Binding to Two Tubulin Isotypes
Chem, 2019
|
|
5ZH2
 
 | | CRYSTAL STRUCTURE OF PfKRS WITH INHIBITOR CLADO-5 | | Descriptor: | (3R)-6,8-dihydroxy-3-{[(2R,6R)-6-methyloxan-2-yl]methyl}-3,4-dihydro-1H-2-benzopyran-1-one, 3[N-MORPHOLINO]PROPANE SULFONIC ACID, CHLORIDE ION, ... | | Authors: | Babbar, P, Malhotra, N, Sharma, M, Harlos, K, Reddy, D.S, Manickam, Y, Sharma, A. | | Deposit date: | 2018-03-11 | | Release date: | 2018-06-27 | | Last modified: | 2023-11-22 | | Method: | X-RAY DIFFRACTION (2.66 Å) | | Cite: | Specific Stereoisomeric Conformations Determine the Drug Potency of Cladosporin Scaffold against Malarial Parasite
J. Med. Chem., 61, 2018
|
|
5ZKE
 
 | | Crystal Structure of N-terminal Domain of Plasmodium vivax p43 in space group P212121 | | Descriptor: | Aminoacyl tRNA Synthetase Complex-Interacting Multifunctional Protein p43 | | Authors: | Gupta, S, Sharma, M, Harlos, K, Manickam, Y, Sharma, A. | | Deposit date: | 2018-03-23 | | Release date: | 2019-04-24 | | Last modified: | 2024-03-06 | | Method: | X-RAY DIFFRACTION (1.492 Å) | | Cite: | Crystal structures of the two domains that constitute the Plasmodium vivax p43 protein.
Acta Crystallogr D Struct Biol, 76, 2020
|
|
5ZKF
 
 | | Crystal Structure of N-terminal Domain of Plasmodium vivax p43 in space group P21 | | Descriptor: | Aminoacyl tRNA Synthetase Complex-Interacting Multifunctional Protein p43 | | Authors: | Gupta, S, Sharma, M, Harlos, K, Manickam, Y, Sharma, A. | | Deposit date: | 2018-03-23 | | Release date: | 2019-04-24 | | Last modified: | 2020-03-11 | | Method: | X-RAY DIFFRACTION (2.75 Å) | | Cite: | Crystal structures of the two domains that constitute the Plasmodium vivax p43 protein.
Acta Crystallogr D Struct Biol, 76, 2020
|
|
5Z79
 
 | | Crystal Structure Analysis of the HPPK-DHPS in complex with substrates | | Descriptor: | 4-AMINOBENZOIC ACID, 5'-DEOXYADENOSINE, 6-HYDROXYMETHYLPTERIN-DIPHOSPHATE, ... | | Authors: | Manickam, Y, Karl, H, Sharma, A. | | Deposit date: | 2018-01-27 | | Release date: | 2018-08-29 | | Last modified: | 2023-11-22 | | Method: | X-RAY DIFFRACTION (2.9 Å) | | Cite: | Structure of 6-hydroxymethyl-7,8-dihydropterin pyrophosphokinase-dihydropteroate synthase fromPlasmodium vivaxsheds light on drug resistance
J. Biol. Chem., 293, 2018
|
|
7WAK
 
 | | Glutamyl-tRNA synthetase from Plasmodium falciparum (PfERS) in complex with ADP | | Descriptor: | ADENOSINE-5'-DIPHOSPHATE, CHLORIDE ION, Glutamyl-tRNA synthetase, ... | | Authors: | Sharma, V, Manickam, Y, Babbar, P, Sharma, A. | | Deposit date: | 2021-12-14 | | Release date: | 2022-12-21 | | Last modified: | 2023-11-29 | | Method: | X-RAY DIFFRACTION (2.782 Å) | | Cite: | Structural characterization of glutamyl-tRNA synthetase (GluRS) from Plasmodium falciparum.
Mol.Biochem.Parasitol., 253, 2022
|
|
7WAL
 
 | | Glutamyl-tRNA synthetase from Plasmodium falciparum (PfERS) in complex with Co | | Descriptor: | BETA-MERCAPTOETHANOL, CHLORIDE ION, COBALT (II) ION, ... | | Authors: | Sharma, V, Manickam, Y, Babbar, P, Sharma, A. | | Deposit date: | 2021-12-14 | | Release date: | 2022-12-21 | | Last modified: | 2024-04-03 | | Method: | X-RAY DIFFRACTION (2.29 Å) | | Cite: | Structural characterization of glutamyl-tRNA synthetase (GluRS) from Plasmodium falciparum.
Mol.Biochem.Parasitol., 253, 2022
|
|
7WAJ
 
 | | Glutamyl-tRNA synthetase from Plasmodium falciparum (PfERS) complexed with ATP and Co | | Descriptor: | ADENOSINE-5'-TRIPHOSPHATE, BETA-MERCAPTOETHANOL, CHLORIDE ION, ... | | Authors: | Sharma, V, Manickam, Y, Babbar, P, Sharma, A. | | Deposit date: | 2021-12-14 | | Release date: | 2022-12-21 | | Last modified: | 2023-11-29 | | Method: | X-RAY DIFFRACTION (2.254 Å) | | Cite: | Structural characterization of glutamyl-tRNA synthetase (GluRS) from Plasmodium falciparum.
Mol.Biochem.Parasitol., 253, 2022
|
|
7WAI
 
 | | Glutamyl-tRNA synthetase from Plasmodium falciparum (PfERS) | | Descriptor: | CHLORIDE ION, Glutamyl-tRNA synthetase | | Authors: | Sharma, V, Manickam, Y, Harlos, K, Sharma, A. | | Deposit date: | 2021-12-14 | | Release date: | 2022-12-21 | | Last modified: | 2023-11-29 | | Method: | X-RAY DIFFRACTION (2.105 Å) | | Cite: | Structural characterization of glutamyl-tRNA synthetase (GluRS) from Plasmodium falciparum.
Mol.Biochem.Parasitol., 253, 2022
|
|
7WAO
 
 | | Glutamyl-tRNA synthetase from Plasmodium falciparum (PfERS) in complex with Mn | | Descriptor: | CHLORIDE ION, Glutamyl-tRNA synthetase, MANGANESE (II) ION | | Authors: | Sharma, V, Manickam, Y, Babbar, P, Sharma, A. | | Deposit date: | 2021-12-14 | | Release date: | 2022-12-21 | | Last modified: | 2023-11-29 | | Method: | X-RAY DIFFRACTION (2.582 Å) | | Cite: | Structural characterization of glutamyl-tRNA synthetase (GluRS) from Plasmodium falciparum.
Mol.Biochem.Parasitol., 253, 2022
|
|
5IO9
 
 | | X-RAY STRUCTURE OF THE N-TERMINAL DOMAIN OF HUMAN DOUBLECORTIN | | Descriptor: | Neuronal migration protein doublecortin | | Authors: | Ruf, A, Benz, J, Burger, D, D'Arcy, B, Debulpaep, M, Di Lello, P, Fry, D, Huber, W, Kremer, T, Laeremans, T, Matile, H, Ross, A, Rudolph, M.G, Rufer, A.C, Sharma, A, Steinmetz, M.O, Steyaert, J, Schoch, G, Stihle, M, Thoma, R. | | Deposit date: | 2016-03-08 | | Release date: | 2016-03-23 | | Last modified: | 2024-01-10 | | Method: | X-RAY DIFFRACTION (1.3 Å) | | Cite: | Crystal Structures of the Human Doublecortin C- and N-terminal Domains in Complex with Specific Antibodies.
J.Biol.Chem., 291, 2016
|
|
5IP4
 
 | | X-RAY STRUCTURE OF THE C-TERMINAL DOMAIN OF HUMAN DOUBLECORTIN | | Descriptor: | Neuronal migration protein doublecortin, XA4551 NANOBODY AGAINST C-DCX | | Authors: | Ruf, A, Benz, J, Burger, D, D'Arcy, B, Debulpaep, M, Di Lello, P, Fry, D, Huber, W, Kremer, T, Laeremans, T, Matile, H, Ross, A, Rudolph, M.G, Rufer, A.C, Sharma, A, Steinmetz, M.O, Steyaert, J, Schoch, G, Stihle, M, Thoma, R. | | Deposit date: | 2016-03-09 | | Release date: | 2016-05-18 | | Last modified: | 2024-01-10 | | Method: | X-RAY DIFFRACTION (1.81 Å) | | Cite: | Crystal Structures of the Human Doublecortin C- and N-terminal Domains in Complex with Specific Antibodies.
J.Biol.Chem., 291, 2016
|
|
5TN9
 
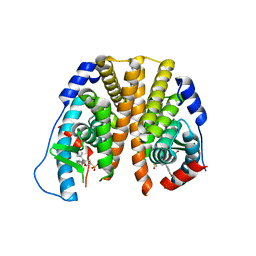 | | Crystal Structure of the ER-alpha Ligand-binding Domain (L372S,L536S) in Complex with the OBHS-BSC, 4-bromophenyl (1R,2R,4S)-5-(4-hydroxyphenyl)-6-(4-(2-(piperidin-1-yl)ethoxy)phenyl)-7-oxabicyclo[2.2.1]hept-5-ene-2-sulfonate | | Descriptor: | 4-bromophenyl (1S,2R,4S)-5-(4-hydroxyphenyl)-6-{4-[2-(piperidin-1-yl)ethoxy]phenyl}-7-oxabicyclo[2.2.1]hept-5-ene-2-sulfonate, Estrogen receptor | | Authors: | Nwachukwu, J.C, Sharma, N, Carlson, K.E, Srinivasan, S, Sharma, A, Katzenellenbogen, J.A, Nettles, K.W. | | Deposit date: | 2016-10-13 | | Release date: | 2017-02-15 | | Last modified: | 2024-03-06 | | Method: | X-RAY DIFFRACTION (2.253 Å) | | Cite: | Exploring the Structural Compliancy versus Specificity of the Estrogen Receptor Using Isomeric Three-Dimensional Ligands.
ACS Chem. Biol., 12, 2017
|
|
5TNB
 
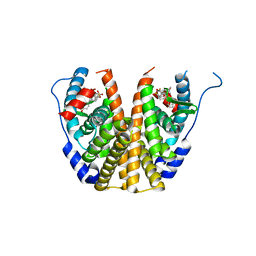 | | Crystal Structure of the ER-alpha Ligand-binding Domain (L372S,L536S) in Complex with the OBHS-BSC, 4-bromophenyl (1R,2R,4S)-6-(4-(2-(dimethylamino)ethoxy)phenyl)-5-(4-hydroxyphenyl)-7-oxabicyclo[2.2.1]hept-5-ene-2-sulfonate | | Descriptor: | 4-bromophenyl (1S,2R,4S)-6-{4-[2-(dimethylamino)ethoxy]phenyl}-5-(4-hydroxyphenyl)-7-oxabicyclo[2.2.1]hept-5-ene-2-sulfonate, Estrogen receptor | | Authors: | Nwachukwu, J.C, Sharma, N, Carlson, K.E, Srinivasan, S, Sharma, A, Katzenellenbogen, J.A, Nettles, K.W. | | Deposit date: | 2016-10-13 | | Release date: | 2017-02-15 | | Last modified: | 2024-03-06 | | Method: | X-RAY DIFFRACTION (2.08 Å) | | Cite: | Exploring the Structural Compliancy versus Specificity of the Estrogen Receptor Using Isomeric Three-Dimensional Ligands.
ACS Chem. Biol., 12, 2017
|
|
7EQU
 | | Crystal structure of the C-lobe of lactoferrin produced by limited proteolysis using pepsin at 2.74A resolution | | Descriptor: | 2-acetamido-2-deoxy-beta-D-glucopyranose-(1-4)-2-acetamido-2-deoxy-beta-D-glucopyranose, BICARBONATE ION, FE (III) ION, ... | | Authors: | Maurya, A, Singh, J, Sharma, A, Sharma, P, Sharma, S, Singh, T.P. | | Deposit date: | 2021-05-04 | | Release date: | 2021-05-19 | | Last modified: | 2023-11-29 | | Method: | X-RAY DIFFRACTION (2.743 Å) | | Cite: | Crystal structure of the C-lobe of lactoferrin produced by limited proteolysis using pepsin at 2.74A resolution
To Be Published
|
|
6AGT
 
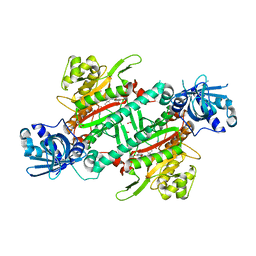 | | Crystal structure of PfKRS complexed with chromone inhibitor | | Descriptor: | COBALT (II) ION, FORMIC ACID, LYSINE, ... | | Authors: | Yogavel, M, Sharma, A, Sharma, A, Baragana, B, Walpole, C. | | Deposit date: | 2018-08-14 | | Release date: | 2019-03-13 | | Last modified: | 2023-11-22 | | Method: | X-RAY DIFFRACTION (1.953 Å) | | Cite: | Lysyl-tRNA synthetase as a drug target in malaria and cryptosporidiosis.
Proc. Natl. Acad. Sci. U.S.A., 116, 2019
|
|
4V5K
 
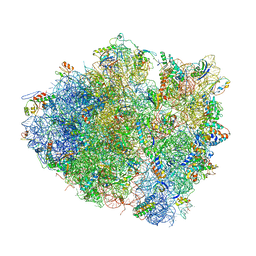 | | Structure of cytotoxic domain of colicin E3 bound to the 70S ribosome | | Descriptor: | 16S ribosomal RNA, 23S ribosomal RNA, 30S RIBOSOMAL PROTEIN S10, ... | | Authors: | Ng, C.L, Lang, K, Meenan, N.A.G, Sharma, A, Kelley, A.C, Kleanthous, C, Ramakrishnan, V. | | Deposit date: | 2010-05-29 | | Release date: | 2014-07-09 | | Last modified: | 2019-07-31 | | Method: | X-RAY DIFFRACTION (3.2 Å) | | Cite: | Structural Basis for 16S Ribosomal RNA Cleavage by the Cytotoxic Domain of Colicin E3.
Nat.Struct.Mol.Biol., 17, 2010
|
|
5ZH4
 
 | | CRYSTAL STRUCTURE OF PfKRS WITH INHIBITOR CLADO-7 | | Descriptor: | (3R)-6,8-dihydroxy-3-{[(2S,6R)-6-methyloxan-2-yl]methyl}-3,4-dihydro-1H-2-benzopyran-1-one, CHLORIDE ION, LYSINE, ... | | Authors: | Babbar, P, Malhotra, N, Sharma, M, Harlos, K, Reddy, D.S, Manickam, Y, Sharma, A. | | Deposit date: | 2018-03-11 | | Release date: | 2018-06-27 | | Last modified: | 2023-11-22 | | Method: | X-RAY DIFFRACTION (2.6 Å) | | Cite: | Specific Stereoisomeric Conformations Determine the Drug Potency of Cladosporin Scaffold against Malarial Parasite
J. Med. Chem., 61, 2018
|
|
5ZKG
 
 | |
5ZH3
 
 | | CRYSTAL STRUCTURE OF PfKRS WITH INHIBITOR CLADO-6 | | Descriptor: | (3S)-6,8-dihydroxy-3-{[(2R,6R)-6-methyloxan-2-yl]methyl}-3,4-dihydro-1H-2-benzopyran-1-one, LYSINE, Lysine-tRNA ligase | | Authors: | Babbar, P, Malhotra, N, Sharma, M, Harlos, K, Reddy, D.S, Manickam, Y, Sharma, A. | | Deposit date: | 2018-03-11 | | Release date: | 2018-06-27 | | Last modified: | 2023-11-22 | | Method: | X-RAY DIFFRACTION (2.86 Å) | | Cite: | Specific Stereoisomeric Conformations Determine the Drug Potency of Cladosporin Scaffold against Malarial Parasite
J. Med. Chem., 61, 2018
|
|
5ZH5
 
 | | CRYSTAL STRUCTURE OF PfKRS WITH INHIBITOR CLADO-2 | | Descriptor: | (3S)-6,8-dihydroxy-3-{[(2R,6S)-6-methyloxan-2-yl]methyl}-3,4-dihydro-1H-2-benzopyran-1-one, CHLORIDE ION, LYSINE, ... | | Authors: | Babbar, P, Malhotra, N, Sharma, M, Harlos, K, Reddy, D.S, Manickam, Y, Sharma, A. | | Deposit date: | 2018-03-11 | | Release date: | 2018-06-27 | | Last modified: | 2023-11-22 | | Method: | X-RAY DIFFRACTION (3.08 Å) | | Cite: | Specific Stereoisomeric Conformations Determine the Drug Potency of Cladosporin Scaffold against Malarial Parasite
J. Med. Chem., 61, 2018
|
|
