7B29
 
 | |
5OX1
 
 | | Glycogen Phosphorylase in complex with JLH270 | | Descriptor: | (2~{R},3~{S},4~{R},5~{R},6~{S})-2-(hydroxymethyl)-6-[5-(4-methoxyphenyl)-1~{H}-1,2,4-triazol-3-yl]oxane-3,4,5-triol, DIMETHYL SULFOXIDE, Glycogen phosphorylase, ... | | Authors: | Kyriakis, E, Stravodimos, G.A, Kantsadi, A.L, Chatzileontiadou, D.S.M, Leonidas, D.D. | | Deposit date: | 2017-09-05 | | Release date: | 2018-02-28 | | Method: | X-RAY DIFFRACTION (1.85 Å) | | Cite: | Probing the beta-pocket of the active site of human liver glycogen phosphorylase with 3-(C-beta-d-glucopyranosyl)-5-(4-substituted-phenyl)-1, 2, 4-triazole inhibitors.
Bioorg. Chem., 77, 2018
|
|
7B28
 | |
7B2A
 
 | |
7B2D
 
 | |
7B26
 
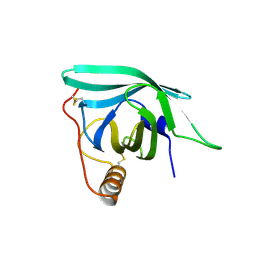
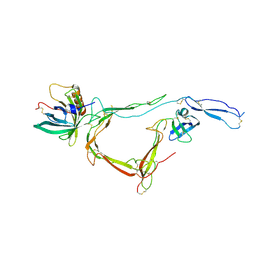 | | CirpA1 in complex with pseudo-monomeric Properdin lacking TSR2-3 | | Descriptor: | CirpA1, Properdin, alpha-D-mannopyranose, ... | | Authors: | Lea, S.M, Johnson, S, Braunger, K. | | Deposit date: | 2020-11-26 | | Release date: | 2021-12-08 | | Last modified: | 2024-07-24 | | Method: | X-RAY DIFFRACTION (3.4 Å) | | Cite: | Structure and function of a family of tick-derived complement inhibitors targeting properdin.
Nat Commun, 13, 2022
|
|
5OMB
 
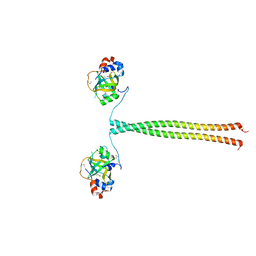 | | Crystal structure of K. lactis Ddc2 N-terminus in complex with S. cerevisiae Rfa1 N-OB domain | | Descriptor: | 1,2-ETHANEDIOL, CHLORIDE ION, DNA damage checkpoint protein LCD1, ... | | Authors: | Deshpande, I, Seeber, A, Shimada, K, Keusch, J.J, Gut, H, Gasser, S.M. | | Deposit date: | 2017-07-28 | | Release date: | 2017-10-25 | | Last modified: | 2017-11-01 | | Method: | X-RAY DIFFRACTION (1.94 Å) | | Cite: | Structural Basis of Mec1-Ddc2-RPA Assembly and Activation on Single-Stranded DNA at Sites of Damage.
Mol. Cell, 68, 2017
|
|
1Z7Y
 
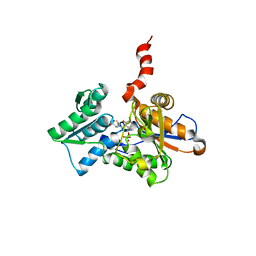 | | Crystal Structure of the Arabidopsis thaliana O-Acetylserine Sulfhydrylase K46A mutant | | Descriptor: | Cysteine synthase, N-[(3-HYDROXY-2-METHYL-5-{[(TRIHYDROXYPHOSPHORANYL)OXY]METHYL}PYRIDIN-4-YL)METHYLENE]METHIONINE | | Authors: | Bonner, E.R, Cahoon, R.E, Knapke, S.M, Jez, J.M. | | Deposit date: | 2005-03-28 | | Release date: | 2005-09-20 | | Last modified: | 2023-08-23 | | Method: | X-RAY DIFFRACTION (2.7 Å) | | Cite: | Molecular Basis of Cysteine Biosynthesis in Plants: STRUCTURAL AND FUNCTIONAL ANALYSIS OF O-ACETYLSERINE SULFHYDRYLASE FROM ARABIDOPSIS THALIANA.
J.Biol.Chem., 280, 2005
|
|
1PLX
 
 | |
1ZMB
 
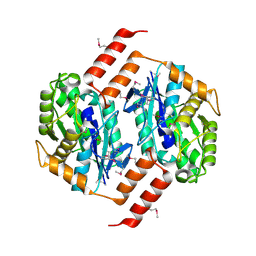 | | Crystal Structure of the Putative Acetylxylan Esterase from Clostridium acetobutylicum, Northeast Structural Genomics Target CaR6 | | Descriptor: | Acetylxylan esterase related enzyme | | Authors: | Forouhar, F, Vorobiev, S.M, Abashidze, M, Ciao, M, Acton, T.B, Montelione, G.T, Hunt, J.F, Tong, L, Northeast Structural Genomics Consortium (NESG) | | Deposit date: | 2005-05-10 | | Release date: | 2005-05-17 | | Last modified: | 2011-07-13 | | Method: | X-RAY DIFFRACTION (2.61 Å) | | Cite: | Crystal Structure of the Putative Acetylxylan Esterase from Clostridium acetobutylicum, Northeast Structural Genomics Target CaR6
To be Published
|
|
5OXH
 
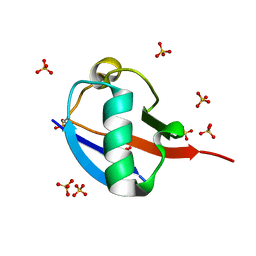 | | C-terminally retracted ubiquitin T66V/L67N mutant | | Descriptor: | SULFATE ION, Ubiquitin T66V/L67N mutant | | Authors: | Gladkova, C, Schubert, A.F, Wagstaff, J.L, Pruneda, J.P, Freund, S.M.V, Komander, D. | | Deposit date: | 2017-09-06 | | Release date: | 2017-11-22 | | Last modified: | 2024-01-17 | | Method: | X-RAY DIFFRACTION (1.601 Å) | | Cite: | An invisible ubiquitin conformation is required for efficient phosphorylation by PINK1.
EMBO J., 36, 2017
|
|
1ZN6
 
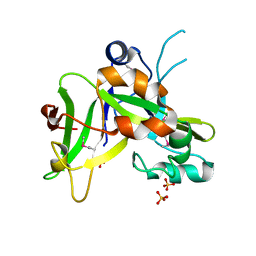 | | X-ray Crystal Structure of Protein Q7WLM8 from Bordetella bronchiseptica. Northeast Structural Genomics Consortium Target BoR19. | | Descriptor: | SULFATE ION, phage-related conserved hypothetical protein | | Authors: | Kuzin, A.P, Yong, W, Vorobiev, S.M, Xiao, R, Ma, L.-C, Acton, T, Montelione, G.T, Hunt, J.F, Tong, L, Northeast Structural Genomics Consortium (NESG) | | Deposit date: | 2005-05-11 | | Release date: | 2005-05-17 | | Last modified: | 2011-07-13 | | Method: | X-RAY DIFFRACTION (1.8 Å) | | Cite: | Novel structure of the hypothetical protein Q7WLM8 from Bordetella bronchiseptica. Northeast Structural Genomics Consortium target BoR19.
To be Published
|
|
5OX4
 
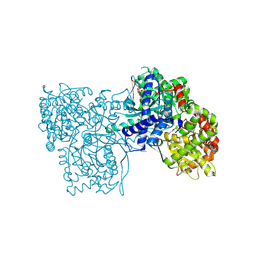 | | Glycogen Phosphorylase in complex with CK900 | | Descriptor: | (2~{S},3~{R},4~{R},5~{S},6~{R})-2-[5-(4-aminophenyl)-4~{H}-1,2,4-triazol-3-yl]-6-(hydroxymethyl)oxane-3,4,5-triol, Glycogen phosphorylase, muscle form, ... | | Authors: | Kyriakis, E, Stravodimos, G.A, Kantsadi, A.L, Chatzileontiadou, D.S.M, Leonidas, D.D. | | Deposit date: | 2017-09-05 | | Release date: | 2018-02-28 | | Method: | X-RAY DIFFRACTION (1.8 Å) | | Cite: | Probing the beta-pocket of the active site of human liver glycogen phosphorylase with 3-(C-beta-d-glucopyranosyl)-5-(4-substituted-phenyl)-1, 2, 4-triazole inhibitors.
Bioorg. Chem., 77, 2018
|
|
1ZX9
 
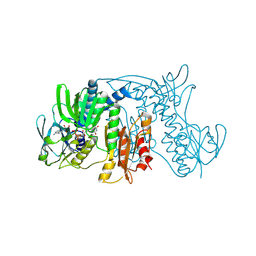 | | Crystal Structure of Tn501 MerA | | Descriptor: | FLAVIN-ADENINE DINUCLEOTIDE, Mercuric reductase | | Authors: | Dong, A, Ledwidge, R, Patel, B, Fiedler, D, Falkowski, M, Zelikova, J, Summers, A.O, Pai, E.F, Miller, S.M. | | Deposit date: | 2005-06-07 | | Release date: | 2005-07-05 | | Last modified: | 2023-08-23 | | Method: | X-RAY DIFFRACTION (1.9 Å) | | Cite: | NmerA, the Metal Binding Domain of Mercuric Ion Reductase, Removes Hg(2+) from Proteins, Delivers It to the Catalytic Core, and Protects Cells under Glutathione-Depleted Conditions
Biochemistry, 44, 2005
|
|
2AFC
 
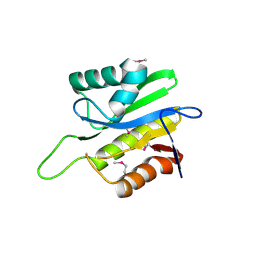 | | X-Ray Crystal Structure of Protein Q8A8B0 from Bacteroides thetaiotaomicron. Northeast Structural Genomics Consortium Target BtR9. | | Descriptor: | conserved hypothetical protein | | Authors: | Kuzin, A.P, Abashidze, M, Vorobiev, S.M, Acton, T, Xiao, R, Conover, K, Ma, L.-C, Cunninggham, K.E, Montelione, G.T, Hunt, J.F, Tong, L, Northeast Structural Genomics Consortium (NESG) | | Deposit date: | 2005-07-25 | | Release date: | 2005-08-02 | | Last modified: | 2011-07-13 | | Method: | X-RAY DIFFRACTION (2.5 Å) | | Cite: | X-ray structure of Q8A8B0 hypothetical protein from Bacteroides thetaiotaomicron
To be Published
|
|
5OMD
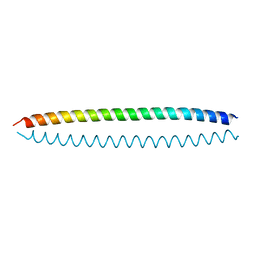 
 | | Crystal structure of S. cerevisiae Ddc2 N-terminal coiled-coil domain | | Descriptor: | DNA damage checkpoint protein LCD1 | | Authors: | Deshpande, I, Seeber, A, Shimada, K, Keusch, J.J, Gut, H, Gasser, S.M. | | Deposit date: | 2017-07-28 | | Release date: | 2017-10-25 | | Last modified: | 2024-01-17 | | Method: | X-RAY DIFFRACTION (2.1 Å) | | Cite: | Structural Basis of Mec1-Ddc2-RPA Assembly and Activation on Single-Stranded DNA at Sites of Damage.
Mol. Cell, 68, 2017
|
|
7BUA
 
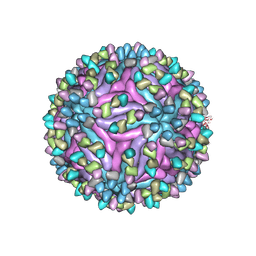 | | Cryo-EM structure of zika virus complexed with Fab SIgN-3C at pH 8.0 | | Descriptor: | 2-acetamido-2-deoxy-beta-D-glucopyranose, Genome polyprotein, SIgN-3C Fab heavy chain, ... | | Authors: | Zhang, S, Chew, S.V, Lim, X.N, Ng, T.S, Kostyuchenko, V.A, Lok, S.M. | | Deposit date: | 2020-04-06 | | Release date: | 2020-05-13 | | Last modified: | 2020-07-29 | | Method: | ELECTRON MICROSCOPY (4.8 Å) | | Cite: | A Human Antibody Neutralizes Different Flaviviruses by Using Different Mechanisms.
Cell Rep, 31, 2020
|
|
6YSL
 | |
1Q7C
 
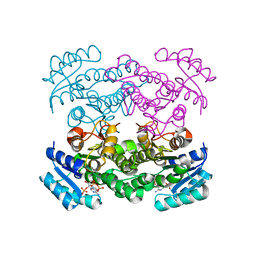 | | The structure of betaketoacyl-[ACP] reductase Y151F mutant in complex with NADPH fragment | | Descriptor: | 3-oxoacyl-[acyl-carrier protein] reductase, NADPH DIHYDRO-NICOTINAMIDE-ADENINE-DINUCLEOTIDE PHOSPHATE | | Authors: | Price, A.C, Zhang, Y.-M, Rock, C.O, White, S.M. | | Deposit date: | 2003-08-17 | | Release date: | 2004-02-17 | | Last modified: | 2024-02-21 | | Method: | X-RAY DIFFRACTION (2.5 Å) | | Cite: | Cofactor-Induced Conformational Rearrangements Establish a Catalytically Competent Active Site and a
Proton Relay Conduit in FabG
Structure, 12, 2004
|
|
1QFK
 
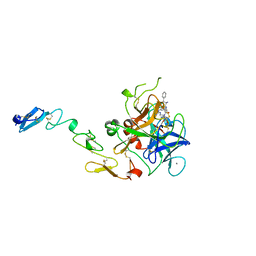 | | STRUCTURE OF HUMAN FACTOR VIIA AND ITS IMPLICATIONS FOR THE TRIGGERING OF BLOOD COAGULATION | | Descriptor: | 2-acetamido-2-deoxy-beta-D-glucopyranose-(1-4)-[beta-L-fucopyranose-(1-6)]2-acetamido-2-deoxy-beta-D-glucopyranose, CALCIUM ION, COAGULATION FACTOR VIIA HEAVY CHAIN, ... | | Authors: | Pike, A.C.W, Brzozowski, A.M, Roberts, S.M, Olsen, O.H, Persson, E. | | Deposit date: | 1999-04-12 | | Release date: | 1999-08-01 | | Last modified: | 2023-08-16 | | Method: | X-RAY DIFFRACTION (2.8 Å) | | Cite: | Structure of human factor VIIa and its implications for the triggering of blood coagulation.
Proc.Natl.Acad.Sci.USA, 96, 1999
|
|
7B9B
 
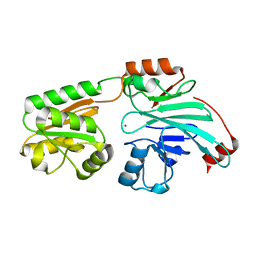 | | Crystal structure of human 5' exonuclease Appollo APO form | | Descriptor: | 5' exonuclease Apollo, NICKEL (II) ION | | Authors: | Newman, J.A, Baddock, H.T, Mukhopadhyay, S.M.M, Burgess-Brown, N.A, von Delft, F, Arrowshmith, C.H, Edwards, A, Bountra, C, Gileadi, O. | | Deposit date: | 2020-12-14 | | Release date: | 2021-01-20 | | Last modified: | 2024-01-31 | | Method: | X-RAY DIFFRACTION (2.8 Å) | | Cite: | A phosphate binding pocket is a key determinant of exo- versus endo-nucleolytic activity in the SNM1 nuclease family.
Nucleic Acids Res., 49, 2021
|
|
7XDP
 
 | |
7X9W
 
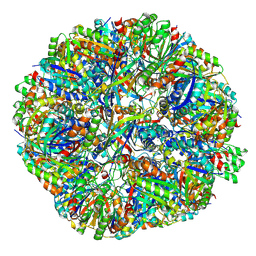 | | Sulfur Oxygenase Reductase from Acidianus ambivalens | | Descriptor: | FE (III) ION, Sulfur oxygenase/reductase | | Authors: | Sobhy, M.A, Hamdan, S.M. | | Deposit date: | 2022-03-16 | | Release date: | 2022-10-12 | | Last modified: | 2022-10-19 | | Method: | ELECTRON MICROSCOPY (2.78 Å) | | Cite: | Cryo-electron structures of the extreme thermostable enzymes Sulfur Oxygenase Reductase and Lumazine Synthase.
Plos One, 17, 2022
|
|
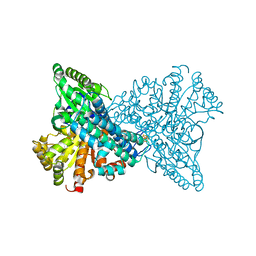
7XDM
 
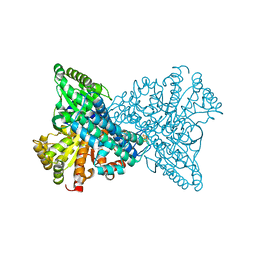 | | ChCODH2 A559W mutant in anaerobic condition | | Descriptor: | Carbon monoxide dehydrogenase 2, FE(4)-NI(1)-S(5) CLUSTER, FE2/S2 (INORGANIC) CLUSTER, ... | | Authors: | Heo, Y.Y, Kim, S.M. | | Deposit date: | 2022-03-28 | | Release date: | 2022-10-19 | | Last modified: | 2023-11-29 | | Method: | X-RAY DIFFRACTION (1.74 Å) | | Cite: | O2-tolerant CO dehydrogenase via tunnel redesign for the removal of CO from industrial flue gas
Nat Catal, 5, 2022
|
|
242D
 
 | | MAD PHASING STRATEGIES EXPLORED WITH A BROMINATED OLIGONUCLEOTIDE CRYSTAL AT 1.65 A RESOLUTION. | | Descriptor: | DNA (5'-D(*CP*GP*CP*GP*(CBR)P*G)-3') | | Authors: | Peterson, M.R, Harrop, S.J, McSweeney, S.M, Leonard, G.A, Thompson, A.W, Hunter, W.N, Helliwell, J.R. | | Deposit date: | 1996-06-20 | | Release date: | 1996-09-19 | | Last modified: | 2024-02-14 | | Method: | X-RAY DIFFRACTION (1.65 Å) | | Cite: | MAD Phasing Strategies Explored with a Brominated Oligonucleotide Crystal at 1.65A Resolution.
J.Synchrotron Radiat., 3, 1996
|
|
