5HOO
 
 | |
4ULL
 
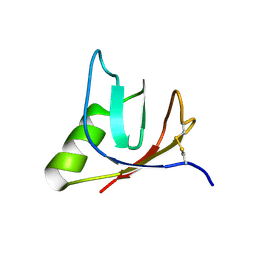 | | SOLUTION NMR STRUCTURE OF VEROTOXIN-1 B-SUBUNIT FROM E. COLI, 5 STRUCTURES | | Descriptor: | Shiga toxin 1B | | Authors: | Richardson, J.M, Evans, P.D, Homans, S.W, Donohue-Rolfe, A. | | Deposit date: | 1996-12-17 | | Release date: | 1997-04-01 | | Last modified: | 2019-12-11 | | Method: | SOLUTION NMR | | Cite: | Solution structure of the carbohydrate-binding B-subunit homopentamer of verotoxin VT-1 from E. coli.
Nat.Struct.Biol., 4, 1997
|
|
5NW9
 
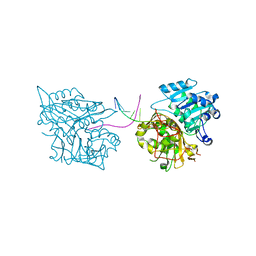 | |
4R79
 
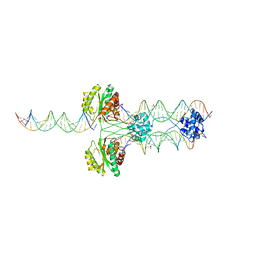 | |
8AF2
 
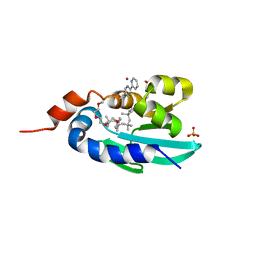 | | Human Sterol Carrier Protein with unnatural amino acid 2,2'-bipyridine alanine incorporated at position 111 | | Descriptor: | COPPER (II) ION, Enoyl-CoA hydratase 2, FRAGMENT OF TRITON X-100, ... | | Authors: | Richardson, J.M, Klemencic, E, Jarvis, A.G. | | Deposit date: | 2022-07-15 | | Release date: | 2023-08-16 | | Last modified: | 2024-04-03 | | Method: | X-RAY DIFFRACTION (2.51 Å) | | Cite: | Using BpyAla to generate copper artificial metalloenzymes: a catalytic and structural study.
Catalysis Science And Technology, 14, 2024
|
|
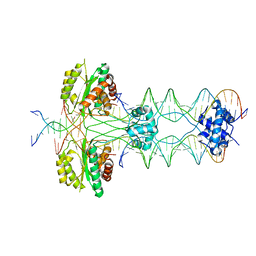
4U7B
 
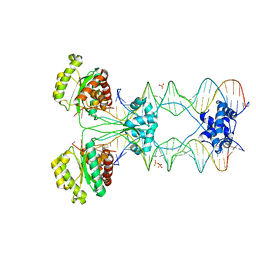 | | Crystal structure of a pre-cleavage Mos1 transpososome | | Descriptor: | DNA (25-MER), DNA (31-MER), Mariner Mos1 transposase, ... | | Authors: | Richardson, J.M. | | Deposit date: | 2014-07-30 | | Release date: | 2015-02-11 | | Last modified: | 2023-12-20 | | Method: | X-RAY DIFFRACTION (3.09 Å) | | Cite: | Structural role of the flanking DNA in mariner transposon excision.
Nucleic Acids Res., 43, 2015
|
|
8AF3
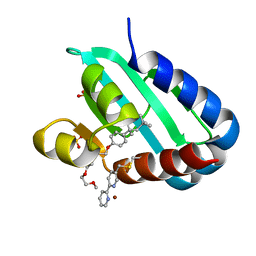 
 | | Sterol carrier protein Artifical metalloenzyme incorporating Q111C mutation coupled to 2,2'-bipyridine | | Descriptor: | COPPER (II) ION, Enoyl-CoA hydratase 2, FRAGMENT OF TRITON X-100, ... | | Authors: | Richardson, J.M, Klemencic, E, Jarvis, A.G. | | Deposit date: | 2022-07-15 | | Release date: | 2023-08-16 | | Last modified: | 2024-07-10 | | Method: | X-RAY DIFFRACTION (1.52 Å) | | Cite: | Using BpyAla to generate copper artificial metalloenzymes: a catalytic and structural study.
Catalysis Science And Technology, 14, 2024
|
|
3FWU
 
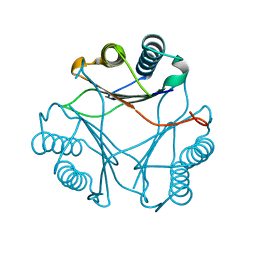 | | Crystal structure of Leishmania major MIF1 | | Descriptor: | Macrophage migration inhibitory factor-like protein | | Authors: | RIchardson, J.M, Walkinshaw, M.D. | | Deposit date: | 2009-01-19 | | Release date: | 2009-03-24 | | Last modified: | 2023-09-06 | | Method: | X-RAY DIFFRACTION (1.8 Å) | | Cite: | Structures of Leishmania major orthologues of macrophage migration inhibitory factor
Biochem.Biophys.Res.Commun., 380, 2009
|
|
3HOT
 
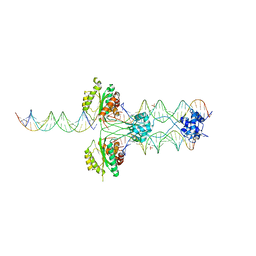 | |
3HOS
 
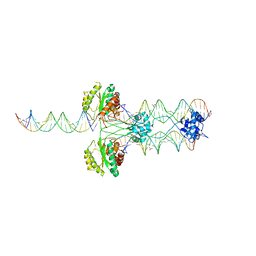 | |
5NWA
 
 | |
4DH4
 
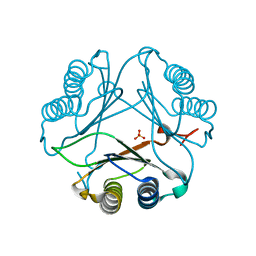 | |
2F7T
 
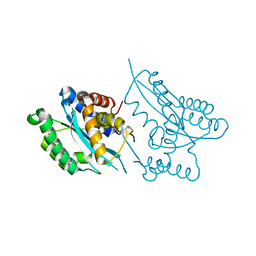 | | Crystal structure of the catalytic domain of Mos1 mariner transposase | | Descriptor: | MAGNESIUM ION, Mos1 transposase | | Authors: | Richardson, J.M, Dawson, A, Taylor, P, Finnegan, D.J, Walkinshaw, M.D. | | Deposit date: | 2005-12-01 | | Release date: | 2006-03-28 | | Last modified: | 2024-02-14 | | Method: | X-RAY DIFFRACTION (2.25 Å) | | Cite: | Mechanism of Mos1 transposition: insights from structural analysis
Embo J., 25, 2006
|
|
1E0F
 
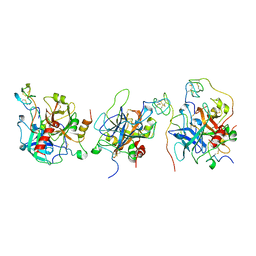 | | Crystal structure of the human alpha-thrombin-haemadin complex: an exosite II-binding inhibitor | | Descriptor: | HAEMADIN, THROMBIN | | Authors: | Richardson, J.L, Kroeger, B, Hoefken, W, Pereira, P, Huber, R, Bode, W, Fuentes-Prior, P. | | Deposit date: | 2000-03-27 | | Release date: | 2000-11-03 | | Last modified: | 2023-12-06 | | Method: | X-RAY DIFFRACTION (3.1 Å) | | Cite: | Crystal Structure of the Human Alpha-Thrombin-Haemadin Complex: An Exosite II-Binding Inhibitor
Embo J., 19, 2000
|
|
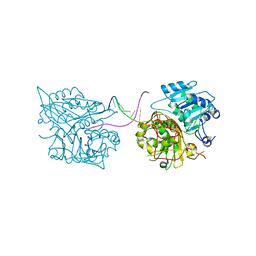
4MDB
 
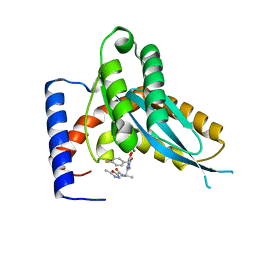 | | Structure of Mos1 transposase catalytic domain and Raltegravir with Mg | | Descriptor: | MAGNESIUM ION, Mariner Mos1 transposase, N-(4-fluorobenzyl)-5-hydroxy-1-methyl-2-(1-methyl-1-{[(5-methyl-1,3,4-oxadiazol-2-yl)carbonyl]amino}ethyl)-6-oxo-1,6-di hydropyrimidine-4-carboxamide | | Authors: | Richardson, J.M. | | Deposit date: | 2013-08-22 | | Release date: | 2014-01-22 | | Last modified: | 2023-09-20 | | Method: | X-RAY DIFFRACTION (1.7 Å) | | Cite: | Structural Basis of Mos1 Transposase Inhibition by the Anti-retroviral Drug Raltegravir.
Acs Chem.Biol., 9, 2014
|
|
4MDA
 
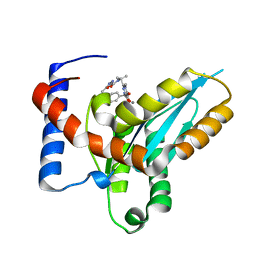 | | Structure of Mos1 transposase catalytic domain and Raltegravir with Mn | | Descriptor: | MANGANESE (II) ION, Mariner Mos1 transposase, N-(4-fluorobenzyl)-5-hydroxy-1-methyl-2-(1-methyl-1-{[(5-methyl-1,3,4-oxadiazol-2-yl)carbonyl]amino}ethyl)-6-oxo-1,6-di hydropyrimidine-4-carboxamide | | Authors: | Richardson, J.M. | | Deposit date: | 2013-08-22 | | Release date: | 2014-01-22 | | Last modified: | 2023-09-20 | | Method: | X-RAY DIFFRACTION (1.7 Å) | | Cite: | Structural Basis of Mos1 Transposase Inhibition by the Anti-retroviral Drug Raltegravir.
Acs Chem.Biol., 9, 2014
|
|
1I53
 
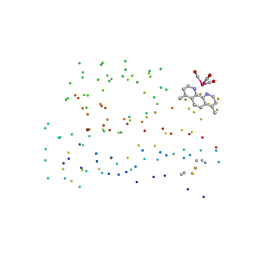 | | RE(I)-TRICARBONYL DIIMINE (Q107H)) AZURIN | | Descriptor: | 4,7-DIMETHYL-[1,10]PHENANTHROLINE, AZURIN, COPPER (II) ION, ... | | Authors: | Di Bilio, A.J, Crane, B.R, Wehbi, W.A, Kiser, C.N, Abu-Omar, M.M, Carlos, R.M, Richards, J.H, Winkler, J.R, Gray, H.B. | | Deposit date: | 2001-02-24 | | Release date: | 2001-10-17 | | Last modified: | 2023-08-09 | | Method: | X-RAY DIFFRACTION (1.8 Å) | | Cite: | Properties of photogenerated tryptophan and tyrosyl radicals in structurally characterized proteins containing rhenium(I) tricarbonyl diimines.
J.Am.Chem.Soc., 123, 2001
|
|
4C83
 
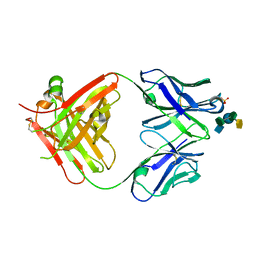 | | Crystal Structure of the IgG2a LPT3 in complex with an 8-sugar inner core analogue of Neisseria meningitidis | | Descriptor: | 2-acetamido-2-deoxy-alpha-D-glucopyranose-(1-2)-L-glycero-alpha-D-manno-heptopyranose-(1-3)-[beta-D-glucopyranose-(1-4)]L-glycero-alpha-D-manno-heptopyranose-(1-5)-3-deoxy-alpha-D-manno-oct-2-ulopyranosonic acid, LPT3 HEAVY CHAIN, LPT3 LIGHT CHAIN, ... | | Authors: | Parker, M.J, Gomery, K, Richard, G, Mackenzie, C.R, Cox, A.D, Richards, J.C, Evans, S.V. | | Deposit date: | 2013-09-29 | | Release date: | 2014-02-12 | | Last modified: | 2023-12-20 | | Method: | X-RAY DIFFRACTION (2.69 Å) | | Cite: | Structural Basis for Selective Cross-Reactivity in a Bactericidal Antibody Against Inner Core Lipooligosaccharide from Neisseria Meningitidis.
Glycobiology, 24, 2014
|
|
2VA8
 
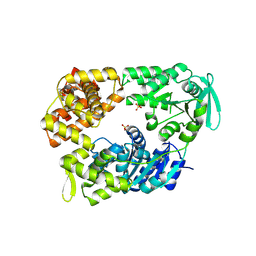 | | DNA Repair Helicase Hel308 | | Descriptor: | SKI2-TYPE HELICASE, SULFATE ION | | Authors: | Johnson, K.A, Richards, J, Liu, H, McMahon, S, Oke, M, Carter, L, Naismith, J.H, White, M.F. | | Deposit date: | 2007-08-30 | | Release date: | 2008-01-15 | | Last modified: | 2024-05-01 | | Method: | X-RAY DIFFRACTION (2.3 Å) | | Cite: | Structure of the DNA Repair Helicase Hel308 Reveals DNA Binding and Autoinhibitory Domains.
J.Biol.Chem., 283, 2008
|
|
5I3I
 
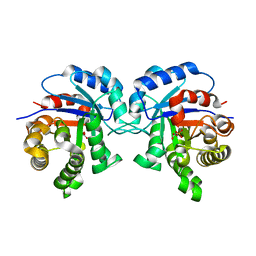 | | Structure-Function Studies on Role of Hydrophobic Clamping of a Basic Glutamate in Catalysis by Triosephosphate Isomerase | | Descriptor: | 2-PHOSPHOGLYCOLIC ACID, Triosephosphate isomerase, glycosomal | | Authors: | Drake, E.J, Gulick, A.M, Richard, J.P, Zhai, X, Kim, K, Reinhardt, C.J. | | Deposit date: | 2016-02-10 | | Release date: | 2016-05-18 | | Last modified: | 2023-09-27 | | Method: | X-RAY DIFFRACTION (2.2 Å) | | Cite: | Structure-Function Studies of Hydrophobic Residues That Clamp a Basic Glutamate Side Chain during Catalysis by Triosephosphate Isomerase.
Biochemistry, 55, 2016
|
|
5I3H
 
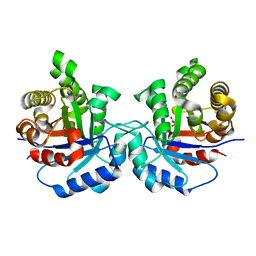 | | Structure-Function Studies on Role of Hydrophobic Clamping of a Basic Glutamate in Catalysis by Triosephosphate Isomerase | | Descriptor: | 2-PHOSPHOGLYCOLIC ACID, POTASSIUM ION, Triosephosphate isomerase, ... | | Authors: | Drake, E.J, Gulick, A.M, Richard, J.P, Zhai, X, Kim, K, Reinhardt, C.J. | | Deposit date: | 2016-02-10 | | Release date: | 2016-05-18 | | Last modified: | 2023-09-27 | | Method: | X-RAY DIFFRACTION (2.25 Å) | | Cite: | Structure-Function Studies of Hydrophobic Residues That Clamp a Basic Glutamate Side Chain during Catalysis by Triosephosphate Isomerase.
Biochemistry, 55, 2016
|
|
5I3F
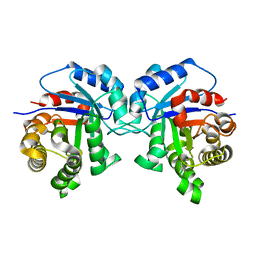 
 | | Structure-Function Studies on Role of Hydrophobic Clamping of a Basic Glutamate in Catalysis by Triosephosphate Isomerase | | Descriptor: | Triosephosphate isomerase, glycosomal | | Authors: | Drake, E.J, Gulick, A.M, Richard, J.P, Zhai, X, Kim, K, Reinhardt, C.J. | | Deposit date: | 2016-02-10 | | Release date: | 2016-05-18 | | Last modified: | 2023-09-27 | | Method: | X-RAY DIFFRACTION (1.72 Å) | | Cite: | Structure-Function Studies of Hydrophobic Residues That Clamp a Basic Glutamate Side Chain during Catalysis by Triosephosphate Isomerase.
Biochemistry, 55, 2016
|
|
5I3J
 
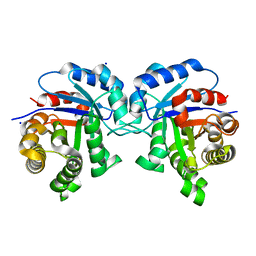 | | Structure-Function Studies on Role of Hydrophobic Clamping of a Basic Glutamate in Catalysis by Triosephosphate Isomerase | | Descriptor: | SODIUM ION, Triosephosphate isomerase, glycosomal | | Authors: | Drake, E.J, Gulick, A.M, Richard, J.P, Zhai, X, Kim, K, Reinhardt, C.J. | | Deposit date: | 2016-02-10 | | Release date: | 2016-05-18 | | Last modified: | 2023-09-27 | | Method: | X-RAY DIFFRACTION (1.8 Å) | | Cite: | Structure-Function Studies of Hydrophobic Residues That Clamp a Basic Glutamate Side Chain during Catalysis by Triosephosphate Isomerase.
Biochemistry, 55, 2016
|
|
4PR6
 
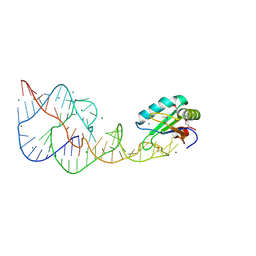 | | A Second Look at the HDV Ribozyme Structure and Dynamics. | | Descriptor: | HDV RIBOZYME SELF-CLEAVED, MAGNESIUM ION, U1 small nuclear ribonucleoprotein A | | Authors: | Kapral, G.J, Jain, S, Noeske, J, Doudna, J.A, Richardson, D.C, Richardson, J.S. | | Deposit date: | 2014-03-05 | | Release date: | 2014-10-29 | | Last modified: | 2014-11-26 | | Method: | X-RAY DIFFRACTION (2.3 Å) | | Cite: | New tools provide a second look at HDV ribozyme structure, dynamics and cleavage.
Nucleic Acids Res., 42, 2014
|
|
4PRF
 
 | | A Second Look at the HDV Ribozyme Structure and Dynamics. | | Descriptor: | Hepatitis Delta virus ribozyme, STRONTIUM ION, U1 small nuclear ribonucleoprotein A | | Authors: | Kapral, G.J, Jain, S, Noeske, J, Doudna, J.A, Richardson, D.C, Richardson, J.S. | | Deposit date: | 2014-03-05 | | Release date: | 2014-10-29 | | Last modified: | 2024-02-28 | | Method: | X-RAY DIFFRACTION (2.395 Å) | | Cite: | New tools provide a second look at HDV ribozyme structure, dynamics and cleavage.
Nucleic Acids Res., 42, 2014
|
|
