1PXH
 
 | | Crystal structure of protein tyrosine phosphatase 1B with potent and selective bidentate inhibitor compound 2 | | Descriptor: | ACETIC ACID, MAGNESIUM ION, N-{1-[5-(1-CARBAMOYL-2-MERCAPTO-ETHYLCARBAMOYL)-PENTYLCARBAMOYL]-2-[4-(DIFLUORO-PHOSPHONO-METHYL)-PHENYL]-ETHYL}-3-{2-[4-(DIFLUORO-PHOSPHONO-METHYL)-PHENYL]-ACETYLAMINO}-SUCCINAMIC ACID, ... | | Authors: | Sun, J.P, Fedorov, A, Lee, S.Y, Guo, X.L, Shen, K, Lawrence, D.S, Almo, S.C, Zhang, Z.Y. | | Deposit date: | 2003-07-04 | | Release date: | 2003-08-12 | | Last modified: | 2023-08-16 | | Method: | X-RAY DIFFRACTION (2.15 Å) | | Cite: | Crystal structure of PTP1B complexed with a potent and selective bidentate inhibitor.
J.Biol.Chem., 278, 2003
|
|
2F1G
 
 | | Cathepsin S in complex with non-covalent 2-(Benzoxazol-2-ylamino)-acetamide | | Descriptor: | Cathepsin S, GLYCEROL, N~2~-1,3-BENZOXAZOL-2-YL-3-CYCLOHEXYL-N-{2-[(4-METHOXYPHENYL)AMINO]ETHYL}-L-ALANINAMIDE | | Authors: | Spraggon, G, Hornsby, M, Lesley, S.A, Tully, D.C, Harris, J.L, Karenewsky, D.S, Kulathila, R, Clark, K. | | Deposit date: | 2005-11-14 | | Release date: | 2006-04-04 | | Last modified: | 2011-07-13 | | Method: | X-RAY DIFFRACTION (1.9 Å) | | Cite: | Synthesis and evaluation of arylaminoethyl amides as noncovalent inhibitors of cathepsin S. Part 3: Heterocyclic P3.
Bioorg.Med.Chem.Lett., 16, 2006
|
|
1K9L
 
 | |
1K9H
 
 | | NMR structure of DNA TGTGAGCGCTCACA | | Descriptor: | 5'-D(*TP*GP*TP*GP*AP*GP*CP*GP*CP*TP*CP*AP*CP*A)-3' | | Authors: | Kaluarachchi, K, Gorenstein, D.G, Luxon, B.A. | | Deposit date: | 2001-10-29 | | Release date: | 2001-11-07 | | Last modified: | 2024-05-22 | | Method: | SOLUTION NMR | | Cite: | How Do Proteins Recognize DNA? Solution Structure and Local Conformational Dynamics of Lac Operators by 2D NMR
J.Biomol.Struct.Dyn., Conversation 11, 2000
|
|
2JZI
 
 | |
1CXR
 
 | |
1R4H
 
 | | NMR Solution structure of the IIIc domain of GB Virus B IRES Element | | Descriptor: | 5'-R(*GP*GP*GP*CP*AP*AP*GP*CP*CP*C)-3' | | Authors: | Kaluarachchi, K, Thiviyanathan, V, Rijinbrand, R, Lemon, S.M, Gorenstein, D.G. | | Deposit date: | 2003-10-06 | | Release date: | 2004-10-19 | | Last modified: | 2024-05-22 | | Method: | SOLUTION NMR | | Cite: | Mutational and structural analysis of stem-loop IIIC of the hepatitis C virus and GB virus B internal ribosome entry sites.
J.Mol.Biol., 343, 2004
|
|
1E41
 
 | |
2K11
 
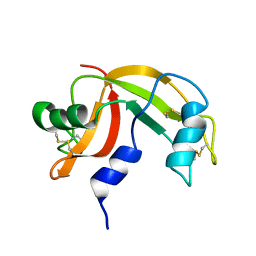 | | Solution structure of human pancreatic ribonuclease | | Descriptor: | Pancreatic Ribonuclease | | Authors: | Kover, K.E, Bruix, M, Santoro, J, Batta, G, Laurents, D.V, Rico, M. | | Deposit date: | 2008-02-20 | | Release date: | 2008-06-03 | | Last modified: | 2022-03-16 | | Method: | SOLUTION NMR | | Cite: | The solution structure and dynamics of human pancreatic ribonuclease determined by NMR spectroscopy provide insight into its remarkable biological activities and inhibition.
J.Mol.Biol., 379, 2008
|
|
1K8S
 
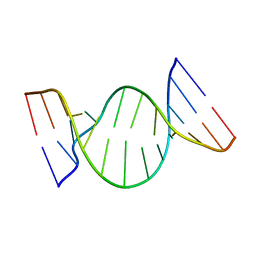 | | BULGED ADENOSINE IN AN RNA DUPLEX | | Descriptor: | 5'-R(*GP*CP*GP*GP*CP*AP*CP*CP*UP*GP*CP*C)-3', 5'-R(*GP*GP*CP*AP*GP*AP*GP*UP*GP*CP*CP*GP*C)-3' | | Authors: | Thiviyanathan, V, Guliaev, A.B, Leontis, N.B, Gorenstein, D.G. | | Deposit date: | 2001-10-25 | | Release date: | 2001-11-14 | | Last modified: | 2024-05-22 | | Method: | SOLUTION NMR | | Cite: | Solution conformation of a bulged adenosine base in an RNA duplex by relaxation matrix refinement.
J.Mol.Biol., 300, 2000
|
|
1T28
 
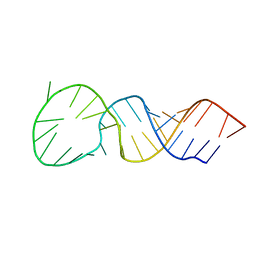 | | High resolution structure of a picornaviral internal cis-acting replication element | | Descriptor: | 34-MER | | Authors: | Thiviyanathan, V, Yang, Y, Kaluarachchi, K, Reynbrand, R, Gorenstein, D.G, Lemon, S.M. | | Deposit date: | 2004-04-20 | | Release date: | 2004-08-10 | | Last modified: | 2024-05-22 | | Method: | SOLUTION NMR | | Cite: | High resolution structure of a picornaviral internal cis-acting replication element(cre).
Proc.Natl.Acad.Sci.USA, 101, 2004
|
|
1IDV
 
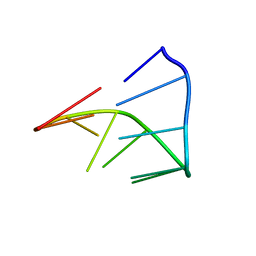 | |
1E3Y
 
 | |
1J3G
 
 | | Solution structure of Citrobacter Freundii AmpD | | Descriptor: | AmpD protein, ZINC ION | | Authors: | Liepinsh, E, Genereux, C, Dehareng, D, Joris, B, Otting, G. | | Deposit date: | 2003-01-31 | | Release date: | 2003-02-18 | | Last modified: | 2023-12-27 | | Method: | SOLUTION NMR | | Cite: | NMR Structure of Citrobacter freundii AmpD, Comparison with Bacteriophage T7 Lysozyme
and Homology with PGRP Domains
J.Mol.Biol., 327, 2003
|
|
2KD8
 
 | |
2KB5
 
 | | Solution NMR Structure of Eosinophil Cationic Protein/RNase 3 | | Descriptor: | Eosinophil cationic protein | | Authors: | Rico, M, Bruix, M, Laurents, D.V, Santoro, J, Jimenez, M, Boix, E, Moussaoui, M, Nogues, M. | | Deposit date: | 2008-11-20 | | Release date: | 2009-06-23 | | Last modified: | 2021-10-20 | | Method: | SOLUTION NMR | | Cite: | The (1)H, (13)C, (15)N resonance assignment, solution structure, and residue level stability of eosinophil cationic protein/RNase 3 determined by NMR spectroscopy
Biopolymers, 91, 2009
|
|
