3OR7
 
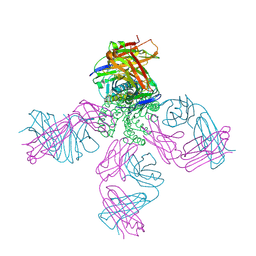 | | On the structural basis of modal gating behavior in K+channels - E71I | | Descriptor: | POTASSIUM ION, Voltage-gated potassium channel, antibody fab fragment heavy chain, ... | | Authors: | Chakrapani, S, Cordero-Morales, J.F, Jogini, V, Pan, A.C, Cortes, D.M, Roux, B, Perozo, E. | | Deposit date: | 2010-09-06 | | Release date: | 2011-01-05 | | Last modified: | 2018-08-29 | | Method: | X-RAY DIFFRACTION (2.3 Å) | | Cite: | On the structural basis of modal gating behavior in K(+) channels.
Nat.Struct.Mol.Biol., 18, 2011
|
|
5V23
 
 | |
5KUA
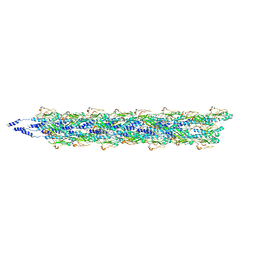 
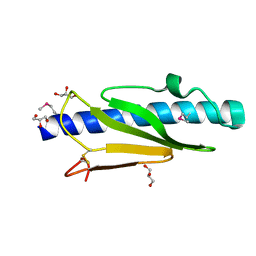
 | | Cryo-EM reconstruction of Neisseria meningitidis Type IV pilus | | Descriptor: | pilin | | Authors: | Kolappan, S, Coureuil, M, Yu, X, Nassif, X, Craig, L, Egelman, E.H. | | Deposit date: | 2016-07-13 | | Release date: | 2016-10-12 | | Last modified: | 2016-11-30 | | Method: | ELECTRON MICROSCOPY (6 Å) | | Cite: | Structure of the Neisseria meningitidis Type IV pilus.
Nat Commun, 7, 2016
|
|
6VTJ
 
 | |
6VTZ
 
 | |
1JSC
 
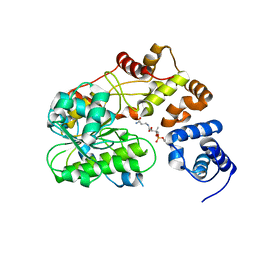
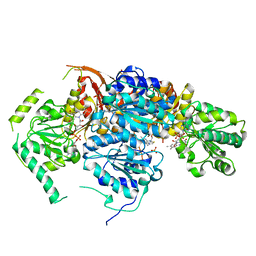 | | Crystal Structure of the Catalytic Subunit of Yeast Acetohydroxyacid Synthase: A target for Herbicidal Inhibitors | | Descriptor: | ACETOHYDROXY-ACID SYNTHASE, DIHYDROGENPHOSPHATE ION, FLAVIN-ADENINE DINUCLEOTIDE, ... | | Authors: | Pang, S.S, Duggleby, R.G, Guddat, L.W. | | Deposit date: | 2001-08-17 | | Release date: | 2002-01-16 | | Last modified: | 2023-08-16 | | Method: | X-RAY DIFFRACTION (2.6 Å) | | Cite: | Crystal structure of yeast acetohydroxyacid synthase: a target for herbicidal inhibitors.
J.Mol.Biol., 317, 2002
|
|
4QT0
 
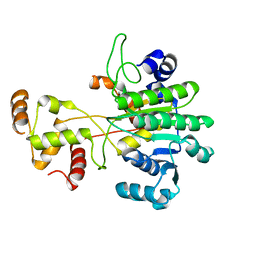
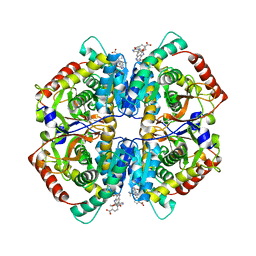 | | Crystal structure of human muscle L-lactate dehydrogenase in complex with inhibitor 1, 3-{[3-CARBAMOYL-7-(2,4-DIMETHOXYPYRIMIDIN-5-YL)QUINOLIN-4-YL]AMINO}BENZOIC ACID | | Descriptor: | 3-{[3-carbamoyl-7-(2,4-dimethoxypyrimidin-5-yl)quinolin-4-yl]amino}benzoic acid, L-lactate dehydrogenase A chain | | Authors: | Kolappan, S, Craig, L. | | Deposit date: | 2014-07-06 | | Release date: | 2014-12-03 | | Last modified: | 2024-02-28 | | Method: | X-RAY DIFFRACTION (3.2 Å) | | Cite: | Structures of lactate dehydrogenase A (LDHA) in apo, ternary and inhibitor-bound forms.
Acta Crystallogr.,Sect.D, 71, 2015
|
|
4QS4
 
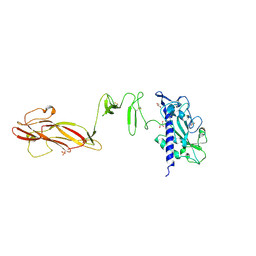 | |
4QSM
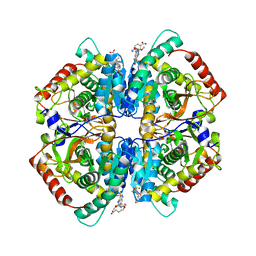 
 | | Crystal structure of human muscle L-lactate dehydrogenase in complex with inhibitor 2, 3-{[7-(2,4-dimethoxypyrimidin-5-yl)-3-sulfamoylquinolin-4-yl]amino}benzoic acid | | Descriptor: | 3-{[7-(2,4-dimethoxypyrimidin-5-yl)-3-sulfamoylquinolin-4-yl]amino}benzoic acid, L-lactate dehydrogenase A chain | | Authors: | Kolappan, S, Craig, L. | | Deposit date: | 2014-07-06 | | Release date: | 2014-12-03 | | Last modified: | 2024-02-28 | | Method: | X-RAY DIFFRACTION (3 Å) | | Cite: | Structures of lactate dehydrogenase A (LDHA) in apo, ternary and inhibitor-bound forms.
Acta Crystallogr.,Sect.D, 71, 2015
|
|
9FHP
 
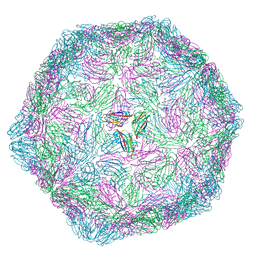 | |
4RZJ
 
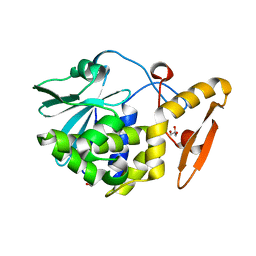 | | Structure of the complex of type 1 ribosome inactivating protein from Momordica balsamina with N-acetylglucosamine at 1.98 Angstrom resolution using crystals grown in different conditions | | Descriptor: | 2-acetamido-2-deoxy-beta-D-glucopyranose, GLYCEROL, Ribosome inactivating protein | | Authors: | Pandey, S, Kushwaha, G.S, Kaur, P, Sharma, S, Singh, T.P. | | Deposit date: | 2014-12-22 | | Release date: | 2015-01-14 | | Last modified: | 2023-09-20 | | Method: | X-RAY DIFFRACTION (1.98 Å) | | Cite: | Structure of the complex of type 1 ribosome inactivating protein from Momordica balsamina with N-acetylglucosamine at 1.98 Angstrom resolution using crystals grown in different conditions
TO BE PUBLISHED
|
|
6P5G
 
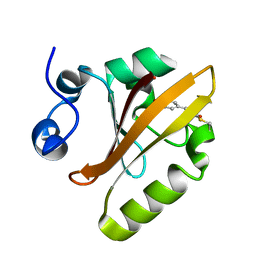 | | Photoactive Yellow Protein PYP Dark Full | | Descriptor: | Photoactive yellow protein | | Authors: | Pandey, S, Schmidt, M. | | Deposit date: | 2019-05-30 | | Release date: | 2019-09-18 | | Last modified: | 2023-11-15 | | Method: | X-RAY DIFFRACTION (1.6 Å) | | Cite: | Time-resolved serial femtosecond crystallography at the European XFEL.
Nat.Methods, 17, 2020
|
|
6P4I
 
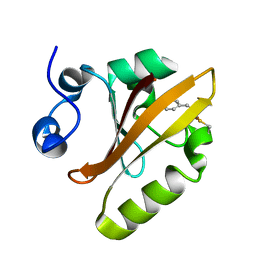 | | Photoactive Yellow Protein PYP 10ps | | Descriptor: | Photoactive yellow protein | | Authors: | Pandey, S, Schmidt, M. | | Deposit date: | 2019-05-27 | | Release date: | 2019-09-18 | | Last modified: | 2023-11-15 | | Method: | X-RAY DIFFRACTION (1.6 Å) | | Cite: | Time-resolved serial femtosecond crystallography at the European XFEL.
Nat.Methods, 17, 2020
|
|
6P5F
 
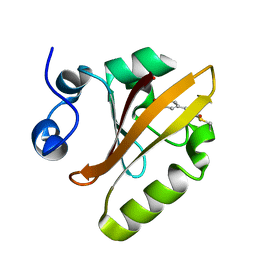 | | Photoactive Yellow Protein PYP Pure Dark | | Descriptor: | Photoactive yellow protein | | Authors: | Pandey, S, Schmidt, M. | | Deposit date: | 2019-05-30 | | Release date: | 2019-09-18 | | Last modified: | 2023-11-15 | | Method: | X-RAY DIFFRACTION (1.7 Å) | | Cite: | Time-resolved serial femtosecond crystallography at the European XFEL.
Nat.Methods, 17, 2020
|
|
6P5E
 
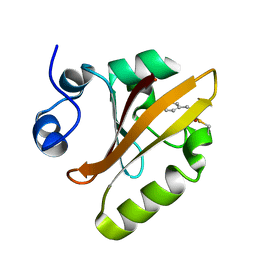 | | Photoactive Yellow Protein PYP 80ps | | Descriptor: | Photoactive yellow protein | | Authors: | Pandey, S, Schmidt, M. | | Deposit date: | 2019-05-30 | | Release date: | 2019-09-18 | | Last modified: | 2023-11-15 | | Method: | X-RAY DIFFRACTION (1.6 Å) | | Cite: | Time-resolved serial femtosecond crystallography at the European XFEL.
Nat.Methods, 17, 2020
|
|
6P5D
 
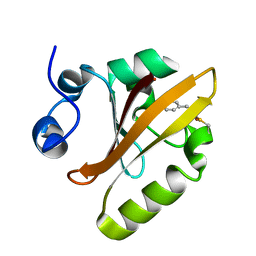 | | Photoactive Yellow Protein PYP 30ps | | Descriptor: | Photoactive yellow protein | | Authors: | Pandey, S, Schmidt, M. | | Deposit date: | 2019-05-30 | | Release date: | 2019-09-18 | | Last modified: | 2023-11-15 | | Method: | X-RAY DIFFRACTION (1.6 Å) | | Cite: | Time-resolved serial femtosecond crystallography at the European XFEL.
Nat.Methods, 17, 2020
|
|
6Y09
 
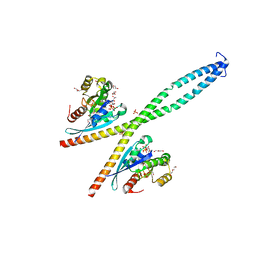 | |
6ZAY
 
 | | Crystal structure of Atg16L in complex with GDP-bound Rab33B | | Descriptor: | 1,2-ETHANEDIOL, Autophagy-related protein 16-1, DI(HYDROXYETHYL)ETHER, ... | | Authors: | Pantoom, S, Wu, Y.W. | | Deposit date: | 2020-06-06 | | Release date: | 2020-09-30 | | Last modified: | 2024-01-24 | | Method: | X-RAY DIFFRACTION (2.4 Å) | | Cite: | RAB33B recruits the ATG16L1 complex to the phagophore via a noncanonical RAB binding protein.
Autophagy, 17, 2021
|
|
6NQ6
 
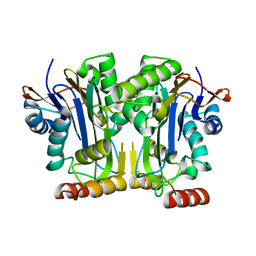 | | Structure & function of a new Aspartylglucosaminuria variant | | Descriptor: | N(4)-(Beta-N-acetylglucosaminyl)-L-asparaginase | | Authors: | Pande, S, Guo, H.C. | | Deposit date: | 2019-01-19 | | Release date: | 2019-04-17 | | Last modified: | 2023-10-11 | | Method: | X-RAY DIFFRACTION (1.5 Å) | | Cite: | The T99K variant of glycosylasparaginase shows a new structural mechanism of the genetic disease aspartylglucosaminuria.
Protein Sci., 28, 2019
|
|
8SLE
 
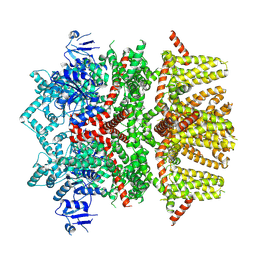 | | Cryo-EM structure of the rat TRPM5 channel in trace calcium, trace-3 | | Descriptor: | Transient receptor potential cation channel subfamily M member 5 | | Authors: | Karuppan, S, Schrag, L.G, Jara-Oseguera, A, Zubcevic, L. | | Deposit date: | 2023-04-21 | | Release date: | 2024-07-03 | | Last modified: | 2024-07-10 | | Method: | ELECTRON MICROSCOPY (4.3 Å) | | Cite: | Structural dynamics at cytosolic interprotomer interfaces control gating of a mammalian TRPM5 channel.
Proc.Natl.Acad.Sci.USA, 121, 2024
|
|
8SLI
 
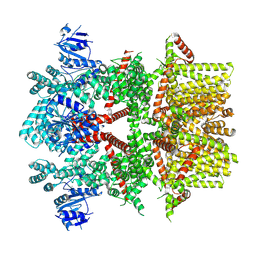 | | Cryo-EM structure of the rat TRPM5 channel in 2mM calcium, high-1 | | Descriptor: | CALCIUM ION, Transient receptor potential cation channel subfamily M member 5 | | Authors: | Karuppan, S, Schrag, L.G, Jara-Oseguera, A, Zubcevic, L. | | Deposit date: | 2023-04-21 | | Release date: | 2024-07-03 | | Last modified: | 2024-07-10 | | Method: | ELECTRON MICROSCOPY (4.9 Å) | | Cite: | Structural dynamics at cytosolic interprotomer interfaces control gating of a mammalian TRPM5 channel.
Proc.Natl.Acad.Sci.USA, 121, 2024
|
|
8SLQ
 
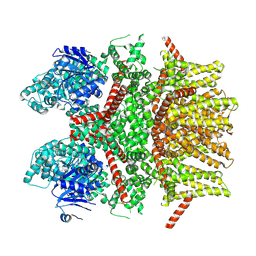 | | Cryo-EM structure of the rat TRPM5 channel in 2mM calcium, high-3 | | Descriptor: | Transient receptor potential cation channel subfamily M member 5 | | Authors: | Karuppan, S, Schrag, L.G, Jara-Oseguera, A, Zubcevic, L. | | Deposit date: | 2023-04-24 | | Release date: | 2024-07-03 | | Last modified: | 2024-07-10 | | Method: | ELECTRON MICROSCOPY (4.5 Å) | | Cite: | Structural dynamics at cytosolic interprotomer interfaces control gating of a mammalian TRPM5 channel.
Proc.Natl.Acad.Sci.USA, 121, 2024
|
|
8SLW
 
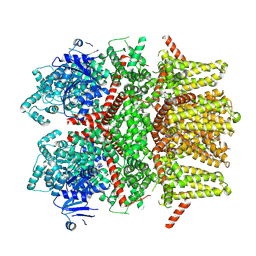 | | Cryo-EM structure of the rat TRPM5 channel in 2mM calcium, high-4 | | Descriptor: | CALCIUM ION, Transient receptor potential cation channel subfamily M member 5 | | Authors: | Karuppan, S, Schrag, L.G, Jara-Oseguera, A, Zubcevic, L. | | Deposit date: | 2023-04-24 | | Release date: | 2024-07-03 | | Last modified: | 2024-07-10 | | Method: | ELECTRON MICROSCOPY (4.2 Å) | | Cite: | Structural dynamics at cytosolic interprotomer interfaces control gating of a mammalian TRPM5 channel.
Proc.Natl.Acad.Sci.USA, 121, 2024
|
|
8SLP
 
 | | Cryo-EM structure of the rat TRPM5 channel in 2mM calcium, high-2 | | Descriptor: | CALCIUM ION, Transient receptor potential cation channel subfamily M member 5 | | Authors: | Karuppan, S, Schrag, L.G, Jara-Oseguera, A, Zubcevic, L. | | Deposit date: | 2023-04-24 | | Release date: | 2024-07-03 | | Last modified: | 2024-07-10 | | Method: | ELECTRON MICROSCOPY (3.8 Å) | | Cite: | Structural dynamics at cytosolic interprotomer interfaces control gating of a mammalian TRPM5 channel.
Proc.Natl.Acad.Sci.USA, 121, 2024
|
|
8SL8
 
 | | Cryo-EM structure of the rat TRPM5 channel in trace calcium, trace-1 | | Descriptor: | Transient receptor potential cation channel subfamily M member 5 | | Authors: | Karuppan, S, Schrag, L.G, Jara-Oseguera, A, Zubcevic, L. | | Deposit date: | 2023-04-21 | | Release date: | 2024-07-03 | | Last modified: | 2024-07-10 | | Method: | ELECTRON MICROSCOPY (4.2 Å) | | Cite: | Structural dynamics at cytosolic interprotomer interfaces control gating of a mammalian TRPM5 channel.
Proc.Natl.Acad.Sci.USA, 121, 2024
|
|
