1CLC
 
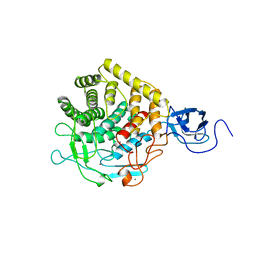 | |
6GDN
 
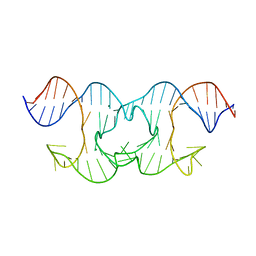 | | Holliday Junctions formed from Telomeric DNA | | Descriptor: | MAGNESIUM ION, Telomere DNA (42-MER) | | Authors: | Parkinson, G.N, Haider, S, Li, P, Khiali, S, Munnur, D, Ramanathan, A. | | Deposit date: | 2018-04-24 | | Release date: | 2018-11-07 | | Last modified: | 2024-01-17 | | Method: | X-RAY DIFFRACTION (3 Å) | | Cite: | Holliday Junctions Formed from Human Telomeric DNA.
J. Am. Chem. Soc., 140, 2018
|
|
1YEB
 | |
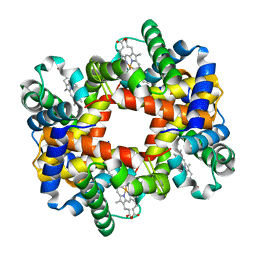
1Y7D
 
 | |
6GE9
 
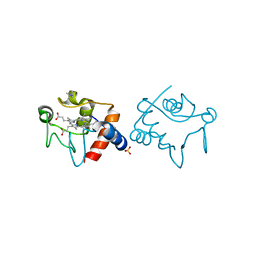
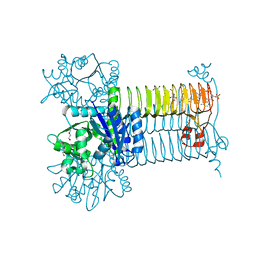 | | Structure of Mycobacterium tuberculosis GlmU bound to Glc-1P and Ac-CoA | | Descriptor: | 1,2-ETHANEDIOL, 1-O-phosphono-alpha-D-glucopyranose, ACETYL COENZYME *A, ... | | Authors: | Craggs, P.D, Mouilleron, S, Rejzek, M, de Chiara, C, Young, R.J, Field, R.A, Argyrou, A, de Carvalho, L.P.S. | | Deposit date: | 2018-04-25 | | Release date: | 2018-05-02 | | Last modified: | 2024-01-17 | | Method: | X-RAY DIFFRACTION (2.26 Å) | | Cite: | The Mechanism of Acetyl Transfer Catalyzed by Mycobacterium tuberculosis GlmU.
Biochemistry, 57, 2018
|
|
6H9H
 
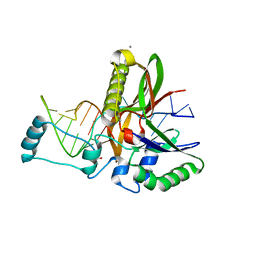 | | Csf5, CRISPR-Cas type IV Cas6 crRNA endonuclease | | Descriptor: | Csf5, GLYCEROL, L(+)-TARTARIC ACID, ... | | Authors: | Pausch, P, Bange, G. | | Deposit date: | 2018-08-04 | | Release date: | 2018-09-26 | | Last modified: | 2024-05-15 | | Method: | X-RAY DIFFRACTION (1.75 Å) | | Cite: | Type IV CRISPR RNA processing and effector complex formation in Aromatoleum aromaticum.
Nat Microbiol, 4, 2019
|
|
8OJF
 
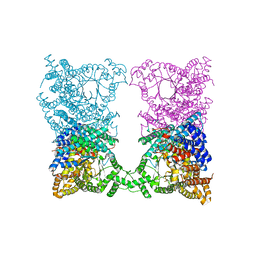 | |
6G4U
 
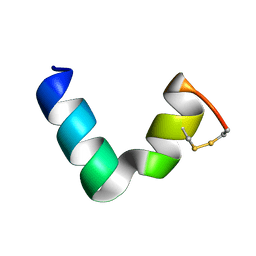 | |
1UTG
 
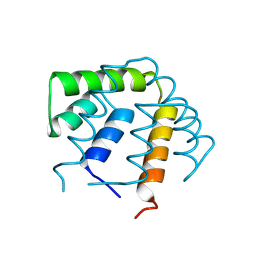 | | REFINEMENT OF THE C2221 CRYSTAL FORM OF OXIDIZED UTEROGLOBIN AT 1.34 ANGSTROMS RESOLUTION | | Descriptor: | UTEROGLOBIN | | Authors: | Morize, I, Surcouf, E, Vaney, M.C, Buehner, M, Mornon, J.P. | | Deposit date: | 1989-04-03 | | Release date: | 1989-10-15 | | Last modified: | 2017-11-29 | | Method: | X-RAY DIFFRACTION (1.34 Å) | | Cite: | Refinement of the C222(1) crystal form of oxidized uteroglobin at 1.34 A resolution.
J.Mol.Biol., 194, 1987
|
|
6G4I
 
 | |
2VEU
 
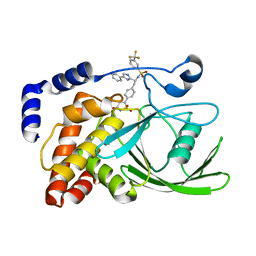 | | Crystal structure of protein tyrosine phosphatase 1B in complex with an isothiazolidinone-containing inhibitor | | Descriptor: | N-[(1S)-2-{4-[(5S)-1,1-dioxido-3-oxoisothiazolidin-5-yl]phenyl}-1-(4-phenyl-1H-imidazol-2-yl)ethyl]-3-(trifluoromethyl)benzenesulfonamide, TYROSINE-PROTEIN PHOSPHATASE NON-RECEPTOR TYPE 1 | | Authors: | Douty, B, Wayland, B, Ala, P.J, Bower, M.J, Pruitt, J, Bostrom, L, Wei, M, Klabe, R, Gonneville, L, Wynn, R, Burn, T.C, Liu, P.C.C, Combs, A.P, Yue, E.W. | | Deposit date: | 2007-10-27 | | Release date: | 2007-11-06 | | Last modified: | 2024-05-08 | | Method: | X-RAY DIFFRACTION (2.4 Å) | | Cite: | Isothiazolidinone Inhibitors of Ptp1B Containing Imidazoles and Imidazolines
Bioorg.Med.Chem.Lett., 18, 2008
|
|
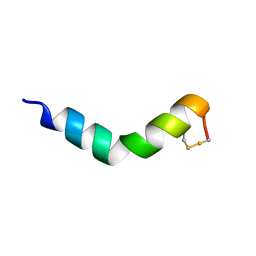
6G4V
 
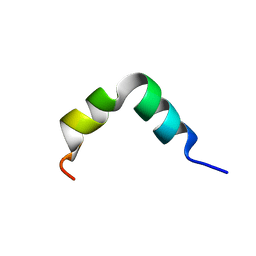 | | The solution NMR structure of [C18S,C24S]brevinin-1BYa in 33% trifluoroethanol | | Descriptor: | [C18S,C24S]brevinin-1BYa | | Authors: | Timmons, P.B, O'Flynn, D.P, Conlon, J.M, Hewage, C.M. | | Deposit date: | 2018-03-28 | | Release date: | 2019-10-16 | | Last modified: | 2024-06-19 | | Method: | SOLUTION NMR | | Cite: | Insights into conformation and membrane interactions of the acyclic and dicarba-bridged brevinin-1BYa antimicrobial peptides.
Eur.Biophys.J., 48, 2019
|
|
4ORG
 
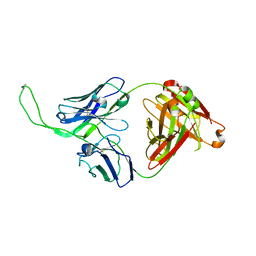 | | Crystal structure of human Fab CAP256-VRC26.04, a potent V1V2-directed HIV-1 neutralizing antibody | | Descriptor: | CAP256-VRC26.04 heavy chain, CAP256-VRC26.04 light chain | | Authors: | Gorman, J, Doria-Rose, N.A, Schramm, C.A, Moore, P.L, Mascola, J.R, Shapiro, L, Morris, L, Kwong, P.D. | | Deposit date: | 2014-02-11 | | Release date: | 2014-02-26 | | Last modified: | 2023-12-06 | | Method: | X-RAY DIFFRACTION (3.121 Å) | | Cite: | Developmental pathway for potent V1V2-directed HIV-neutralizing antibodies.
Nature, 509, 2014
|
|
6G4X
 
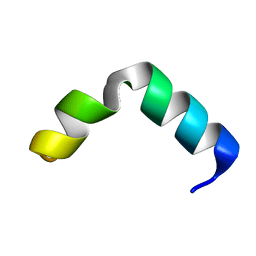 | | The solution NMR structure of [C18S,C24S]brevinin-1BYa in sodium dodecyl sulphate micelles | | Descriptor: | [C18S,C24S]brevinin-1BYa | | Authors: | Timmons, P.B, O'Flynn, D.P, Conlon, J.M, Hewage, C.M. | | Deposit date: | 2018-03-28 | | Release date: | 2019-10-16 | | Last modified: | 2024-06-19 | | Method: | SOLUTION NMR | | Cite: | Insights into conformation and membrane interactions of the acyclic and dicarba-bridged brevinin-1BYa antimicrobial peptides.
Eur.Biophys.J., 48, 2019
|
|
2VAZ
 
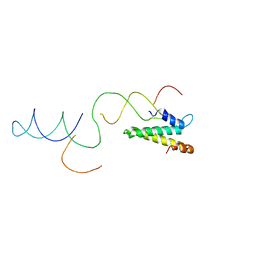 | | Model of the S15-mRNA complex fitted into the cryo-EM map of the 70S entrapment complex. | | Descriptor: | 30S RIBOSOMAL PROTEIN S15, RPSO MRNA OPERATOR | | Authors: | Marzi, S, Myasnikov, A.G, Serganov, A, Ehresmann, C, Romby, P, Yusupov, M, Klaholz, B.P. | | Deposit date: | 2007-09-05 | | Release date: | 2007-10-02 | | Last modified: | 2024-05-08 | | Method: | ELECTRON MICROSCOPY (10 Å) | | Cite: | Structured Mrnas Regulate Translation Initiation by Binding to the Platform of the Ribosome.
Cell(Cambridge,Mass.), 130, 2007
|
|
2V8U
 
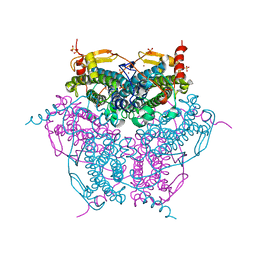 | | Atomic resolution structure of Mn catalase from Thermus Thermophilus | | Descriptor: | LITHIUM ION, MANGANESE (II) ION, MANGANESE-CONTAINING PSEUDOCATALASE, ... | | Authors: | Barynin, V.V, Antonyuk, S.V, Vaguine, A.A, Melik-Adamyan, W.R, Popov, A.N, Lamsin, V.S, Harrison, P.M, Artymiuk, P.J. | | Deposit date: | 2007-08-14 | | Release date: | 2007-09-25 | | Last modified: | 2024-05-08 | | Method: | X-RAY DIFFRACTION (1.05 Å) | | Cite: | Three-Dimentional Structure of the Enzyme Dimanganese Catalase from Thermus Thermophilus at 1 Angstrom Resolution
Crystallogr.Rep.(Transl. Kristallografiya), 45, 2000
|
|
1Y31
 
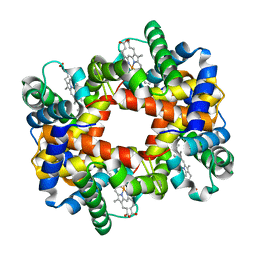 | |
5UVF
 
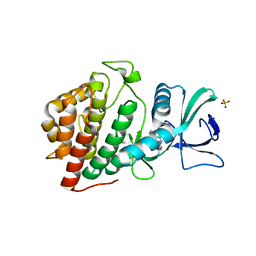 | | Crystal Structure of the Human vaccinia-related kinase bound to BI-D1870 | | Descriptor: | (7S)-2-[(3,5-difluoro-4-hydroxyphenyl)amino]-5,7-dimethyl-8-(3-methylbutyl)-7,8-dihydropteridin-6(5H)-one, CHLORIDE ION, DI(HYDROXYETHYL)ETHER, ... | | Authors: | Counago, R.M, Bountra, C, Arruda, P, Edwards, A.M, Gileadi, O, Structural Genomics Consortium (SGC) | | Deposit date: | 2017-02-20 | | Release date: | 2017-03-22 | | Last modified: | 2023-10-04 | | Method: | X-RAY DIFFRACTION (2 Å) | | Cite: | Structural characterization of human Vaccinia-Related Kinases (VRK) bound to small-molecule inhibitors identifies different P-loop conformations.
Sci Rep, 7, 2017
|
|
6GKO
 
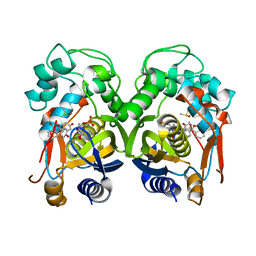 | | Mouse thymidylate synthase cocrystallized with dUMP and soaked in phenolphthalein | | Descriptor: | 2'-DEOXYURIDINE 5'-MONOPHOSPHATE, 3,3-bis(4-hydroxyphenyl)-2-benzofuran-1-one, DIMETHYL SULFOXIDE, ... | | Authors: | Maj, P, Wilk, P, Jarmula, A, Weiss, M.S, Rode, W. | | Deposit date: | 2018-05-21 | | Release date: | 2019-06-05 | | Last modified: | 2024-01-17 | | Method: | X-RAY DIFFRACTION (1.839 Å) | | Cite: | Thymidylate synthase fragment screening
To Be Published
|
|
2WZL
 
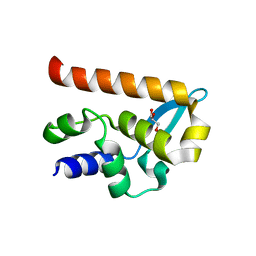 | | The Structure of the N-RNA Binding Domain of the Mokola virus Phosphoprotein | | Descriptor: | GLYCEROL, PHOSPHOPROTEIN | | Authors: | Assenberg, R, Delmas, O, Ren, J, Vidalain, P, Verma, A, Larrous, F, Graham, S, Tangy, F, Grimes, J, Bourhy, H. | | Deposit date: | 2009-11-30 | | Release date: | 2009-12-15 | | Last modified: | 2023-12-20 | | Method: | X-RAY DIFFRACTION (2.1 Å) | | Cite: | The Structure of the N-RNA Binding Domain of the Mokola Virus Phosphoprotein
J.Virol., 84, 2010
|
|
1ZUF
 
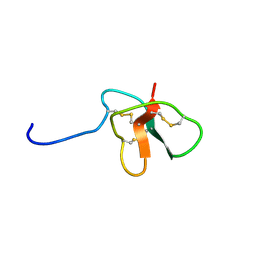 | | Solution Structure of DLP-4 | | Descriptor: | Defensin-like peptide 2/4 | | Authors: | Torres, A.M, Tsampazi, C, Geraghty, D.P, Bansal, P.S, Alewood, P.F, Kuchel, P.W. | | Deposit date: | 2005-05-31 | | Release date: | 2005-08-02 | | Last modified: | 2022-03-02 | | Method: | SOLUTION NMR | | Cite: | D-amino acid residue in a defensin-like peptide from platypus venom: effect on structure and chromatographic properties.
Biochem.J., 391, 2005
|
|
5GT3
 
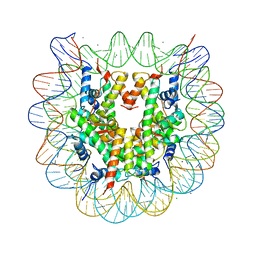 | | Crystal structure of nucleosome particle in the presence of human testis-specific histone variant, hTh2b | | Descriptor: | CHLORIDE ION, DNA (146-MER), Histone H2A type 1-D, ... | | Authors: | Kumarevel, T, Sivaraman, P. | | Deposit date: | 2016-08-18 | | Release date: | 2017-02-15 | | Last modified: | 2023-11-08 | | Method: | X-RAY DIFFRACTION (2.91 Å) | | Cite: | Structural analyses of the nucleosome complexes with human testis-specific histone variants, hTh2a and hTh2b
Biophys. Chem., 221, 2017
|
|
1ZUE
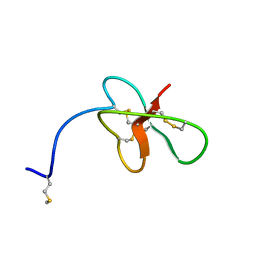 
 | | Revised Solution Structure of DLP-2 | | Descriptor: | Defensin-like peptide 2/4 | | Authors: | Torres, A.M, Tsampazi, C, Geraghty, D.P, Bansal, P.S, Alewood, P.F, Kuchel, P.W. | | Deposit date: | 2005-05-30 | | Release date: | 2005-08-02 | | Last modified: | 2022-03-02 | | Method: | SOLUTION NMR | | Cite: | D-amino acid residue in a defensin-like peptide from platypus venom: effect on structure and chromatographic properties.
Biochem.J., 391, 2005
|
|
6GTJ
 
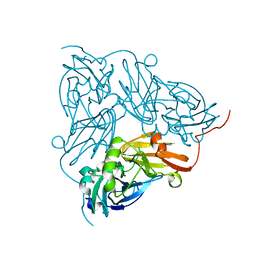 | | Neutron crystal structure for copper nitrite reductase from Achromobacter Cycloclastes at 1.8 A resolution | | Descriptor: | COPPER (II) ION, Copper-containing nitrite reductase | | Authors: | Antonyuk, S.V, Blakeley, M.P, Halsted, T.P, Eady, R.R, Hasnain, S.S. | | Deposit date: | 2018-06-18 | | Release date: | 2019-07-03 | | Last modified: | 2024-01-17 | | Method: | NEUTRON DIFFRACTION (1.801 Å) | | Cite: | Catalytically important damage-free structures of a copper nitrite reductase obtained by femtosecond X-ray laser and room-temperature neutron crystallography.
Iucrj, 6, 2019
|
|
1COW
 
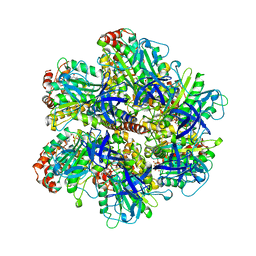 | | BOVINE MITOCHONDRIAL F1-ATPASE COMPLEXED WITH AUROVERTIN B | | Descriptor: | ADENOSINE-5'-DIPHOSPHATE, AUROVERTIN B, BOVINE MITOCHONDRIAL F1-ATPASE, ... | | Authors: | van Raaij, M.J, Abrahams, J.P, Leslie, A.G.W, Walker, J.E. | | Deposit date: | 1996-05-08 | | Release date: | 1996-08-17 | | Last modified: | 2024-02-07 | | Method: | X-RAY DIFFRACTION (3.1 Å) | | Cite: | The structure of bovine F1-ATPase complexed with the antibiotic inhibitor aurovertin B.
Proc.Natl.Acad.Sci.USA, 93, 1996
|
|
